
•REVIEW• November 2022 Vol.65 No.11: 2162–2190
https://doi.org/10.1007/s11427-022-2120-1
Mechanisms of chromatin-based epigenetic inheritance
Wenlong Du
1†
, Guojun Shi
2†
, Chun-Min Shan
3†
, Zhiming Li
4†
, Bing Zhu
1,5*
, Songtao Jia
6*
,
Qing Li
2*
& Zhiguo Zhang
4*
1
National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of
Sciences, Beijing 100101, China;
2
State Key Laboratory of Protein and Plant Gene Research, School of Life Sciences and Peking-Tsinghua Center for Life Sciences,
Peking University, Beijing 100871, China;
3
State Key Laboratory of Plant Genomics, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China;
4
Institutes of Cancer Genetics, Herbert Irving Comprehensive Cancer Center, Columbia University Medical Center, New York, NY 10032, USA;
5
College of Life Sciences, University of Chinese Academy of Sciences, Beijing 100049, China;
6
Department of Biological Sciences, Columbia University, New York, NY 10027, USA
Received February 9, 2022; accepted April 27, 2022; published online June 30, 2022
Multi-cellular organisms such as humans contain hundreds of cell types that share the same genetic information (DNA se-
quences), and yet have different cellular traits and functions. While how genetic information is passed through generations has
been extensively characterized, it remains largely obscure how epigenetic information encoded by chromatin regulates the
passage of certain traits, gene expression states and cell identity during mitotic cell divisions, and even through meiosis. In this
review, we will summarize the recent advances on molecular mechanisms of epigenetic inheritance, discuss the potential impacts
of epigenetic inheritance during normal development and in some disease conditions, and outline future research directions for
this challenging, but exciting field.
epigenetic inheritance, histone modification, DNA methylation, histone deposition, DNA replication
Citation: Du, W., Shi, G., Shan, C.M., Li, Z., Zhu, B., Jia, S., Li, Q., and Zhang, Z. (2022). Mechanisms of chromatin-based epigenetic inheritance. Sci China
Life Sci 65, 2162–2190. https://doi.org/10.1007/s11427-022-2120-1
Introduction
Epigenetics was coined by Waddington in 1942 as a frame-
work for the generation of distinct phenotypes in multi-cel-
lular organisms (Waddington, 1942). At the time, DNA was
not discovered as the carrier of genetic information that
governs the transmission of genetic traits from generation to
generation. Since then, it has been increasingly clear that
epigenetic regulation plays a critical role in the development
of multicellular organisms including human, and mis-reg-
ulations of the epigenetic network are the drivers for many
forms of diseases including cancer and aging (Margueron
and Reinberg, 2010; Benayoun et al., 2015; Allis and Jenu-
wein, 2016; Jones et al., 2016). In this review, we will focus
on discussing the molecular mechanisms underlying how
epigenetic information is inherited into daughter cells during
mitotic cell divisions. While we will mention several ex-
amples on the trans-generational epigenetic inheritance, we
will concentrate our discussion on epigenetic inheritance
during mitosis, and refer the readers to other reviews dis-
cussing the mechanisms and the impacts of trans-genera-
tional epigenetic inheritance (Heard and Martienssen, 2014;
Horsthemke, 2018).
In early days of epigenetic research, scientists described
© Science China Press and Springer-Verlag GmbH Germany, part of Springer Nature 2022 life.scichina.com link.springer.com
SCIENCE CHINA
Life Sciences
†Contributed equally to this work
and studied biological phenomena that cannot be explained
by genetic information alone. These examples include po-
sition effect variegation observed in Drosophila, X chro-
mosome inactivation in female mammals, genome
imprinting in mammals, and para-mutations observed in
plants. Position effect variegation is a phenomenon in which
the white gene in Drosophila eye is expressed in some cells
but silenced in others when the white gene translocates closer
to heterochromatin region, a highly condensed chromatin
domain that is transcriptionally silent (Tartof et al., 1984).
That the expression of a gene was based on its location on the
chromosome, but not the gene itself, was also observed in
budding yeast when a gene was inserted closer to telomeres
(telomere position effects) (Gottschling et al., 1990). X-
chromosome inactivation in female mammals is a mechan-
ism whereby one of two X-chromosomes is inactivated in
female mammals during early embryogenesis to balance the
expression of genes on X-chromosomes between male and
female. Moreover, once silenced, the inactivated X-chro-
mosome remains silent during subsequent cell divisions
(Plath et al., 2002). Genome imprinting is a phenomenon in
which the maternal or paternal allele of a gene is expressed,
while the other allele is silenced (Ferguson-Smith and
Bourc’his, 2018). These examples remain the best to illus-
trate the modern definition of epigenetics, heritable changes
in gene expression/phenotypes without alterations at the
underlying DNA sequences (Allis et al., 2007; Margueron
and Reinberg, 2010). While not all inheritable epigenetic
information is encoded by the chromatin, such as prions, in
this review, we will focus on discussion of inheritance of
epigenetic information encoded by chromatin in eukaryotes.
In eukaryotic cells, the genetic material forms a highly
ordered structure, chromatin, consisting of proteins, DNA
and RNA. The basic repeat unit of chromatin is the nucleo-
some, consisting of 147 bp of DNA wrapped around a his-
tone octamer composed of one H3-H4 tetramer and two
H2A-H2B dimers (Zhou et al., 2019; Talbert and Henikoff,
2021). Chromatin is further organized into distinct domains
such as heterochromatin and euchromatin, which tradition-
ally represent chromatin regions with inactive and active
gene transcription, respectively. For in-depth discussion,
please see recent reviews on insights of three-dimensional
chromatin structures (Dekker and Mirny, 2016; Yu and Ren,
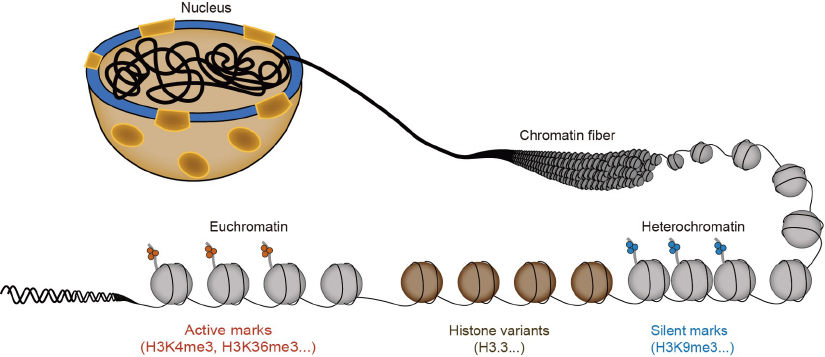
2017; Li et al., 2020a). Furthermore, heterochromatin and
euchromatin are marked by different posttranslational mod-
ifications on histones (Figure 1). For instance, di- and tri-
methylation of histone H3 lysine 9 (H3K9me2/me3) mark
constitutive heterochromatic regions, such as repetitive DNA
sequences including endogenous retroviral elements (ERVs),
pericentric heterochromatin regions and telomeric hetero-
chromatin (Grewal and Moazed, 2003). On the other hand,
tri-methylation of histone H3 lysine 27 (H3K27me3) plays
an important role in the repression of gene transcription
during development (Margueron and Reinberg, 2011). Be-
sides these repressive marks, other histone modifications are
associated with active gene transcription. Tri-methylation of
H3 lysine 4 (H3K4me3) is highly enriched at promoters of
actively transcribed genes (Shilatifard, 2012), whereas
H3K36me3 marks the gene bodies of actively transcribed
genes (Wagner and Carpenter, 2012). In addition to histone
modifications, histone variants, a group of proteins that
adopt similar fold as core histones, reside in specific chro-
matin regions and are also important for the establishment
and maintenance of chromatin states (Loyola and Almouzni,
2007; Talbert and Henikoff, 2010). For instance, histone H3
variant CenH3 proteins occupy centromeric heterochromatin
regions and are critical for the establishment of a functional
kinetochore for chromosome segregation during mitosis.
Histone variant H3.3, which differs from canonical H3.1/
H3.2 by 4 or 5 amino acids, marks actively transcribed re-
gions, whereas canonical H3.1/H3.2 are enriched at hetero-
chromatin. Moreover, DNA cytosine can be methylated
Figure 1 A cartoon depicts representative chromatin states.
2163
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
(5mC) or hydroxymethylated (5hmC), which are distributed
on chromatin differently. At constitutive heterochromatin
regions, 5mC co-localizes with H3K9me2/me3 (see detailed
discussion below). In contrast, 5hmC in general is found at
promoters and enhancers of actively transcribed genes. Fi-
nally, non-coding RNAs also play a role in forming distinct
chromatin states (Zaratiegui et al., 2007). In summary,
chromatin is demarcated by histone modifications, histone
variants, DNA methylation and non-coding RNA (not dis-
cussed in this review). Together, they play an important role
in the establishment and maintenance of chromatin struc-
tures, gene expression and cell identity.
During DNA replication, chromatin structures are tran-
siently disassembled to allow DNA replication machinery
to access replicating DNA. Following DNA replication,
distinct chromatin states, marked by different histone
modifications, histone variants, DNA methylation and non-
coding RNA must be restored to maintain chromatin
structures and gene expression states (Moazed, 2011; Ma-
cAlpine and Almouzni, 2013; Serra-Cardona and Zhang,
2017). How distinct chromatin states are inherited follow-
ing DNA replication lies in the heart of epigenetics. In this
review, we will first discuss how nucleosomes, the basic
repeat units of chromatin, are assembled following DNA
replication, and outline the general principles in the passage
of histone modifications into daughter cells. As an example,
we will discuss in depth on how H3K9 methylation in S.
pombe is inherited during mitotic cell division. Further-
more, we will discuss how DNA methylation is inherited,
and highlight the potential interplay between DNA me-
thylation and histone modifications to maintain chromatin
states. Finally, we will discuss the potential impact of
dysregulation of epigenetic inheritance in development and
human diseases and outline future research directions for
this challenging, but exciting field.
DNA replication-coupled nucleosome assembly
A brief overview of DNA replication in eukaryotic cells
During S phase of the cell cycle, DNA sequence must be
faithfully replicated to maintain genome integrity. DNA re-
plication initiates stochastically from DNA replication ori-
gins (MacAlpine, 2021). While replication origins are well-
defined and contain consensus sequence motifs in Sacchar-
omyces cerevisiae, DNA replication origins in higher eu-
karyotic cells are specified and influenced by local chromatin
structures (Hu et al., 2020; Long et al., 2020). The initial step
in the initiation of DNA replication is the assembly of pre-
replication complex (pre-RC) at a replication origin. During
this process, a group of proteins are orderly assembled into a
large complex at G1 phase at replication origins (Bell and
Dutta, 2002; Burgers and Kunkel, 2017). First, origin re-
cognition complex (ORC), which is composed of six sub-
units (Orc1–6), recognizes replication origins (Bell and
Stillman, 1992), and together with CDC6 and CDC10-de-
pendent transcript 1 (CDT1), loads the hexameric mini-
chromosome maintenance (MCM) complex, consisting of
MCM2–7, at replication origins to form the pre-RC complex
(Donovan et al., 1997; Tanaka et al., 1997). The loaded
MCM complexes at this stage are head-to-head inactive
double hexamers and encircle double-stranded (ds) DNA.
Phosphorylation of the MCM complex by DDK (DBF4-de-
pendent kinase) and CDKs and subsequent binding of
CDC45 and the DNA replication complex GINS (go-ichi-ni-
san) lead to formation of two active replicative helicases, the
CMG helicase (Cdc45-MCM-GINS) (Ilves et al., 2010). The
CMG complex unwinds dsDNA into ssDNA, which is
coated with ssDNA binding protein, replication protein A
(RPA). Two short RNA-DNA primers are then synthesized
by the primase-DNA polymerase alpha (Polα) complex,
which are used by DNA polymerase epsilon (Polε) to syn-
thesize the leading strands continuously and DNA poly-
merase delta (Polδ) to synthesize the lagging strands as
Okazaki fragments. Finally, Ctf4 (AND1 in mammalian
cells) connects the CMG helicase with Polα primase, which
likely coordinates leading and lagging DNA synthesis as
well as nucleosome assembly of parental histones (See
Discussion below). Together, the multi-component protein
machinery, namely the replisome, replicates DNA in a highly
regulated manner.
An overview of DNA replication-coupled nucleosome
assembly
In general, nucleosomes limit the accessibility of protein
machinery involved in various DNA transactions such as
DNA replication, repair and gene transcription to the nu-
cleosomal DNA. Therefore, during DNA replication, 1-2
nucleosomes ahead of DNA replication forks are temporarily
disassembled to allow the replisome to access DNA. Fol-
lowing the passage of DNA replication forks, replicated
DNA is reassembled into nucleosomes using both parental
histones and newly synthesized histones in a process tightly
coupled to on-going DNA replication (DNA replication-
coupled nucleosome assembly) (McKnight and Miller, 1977;
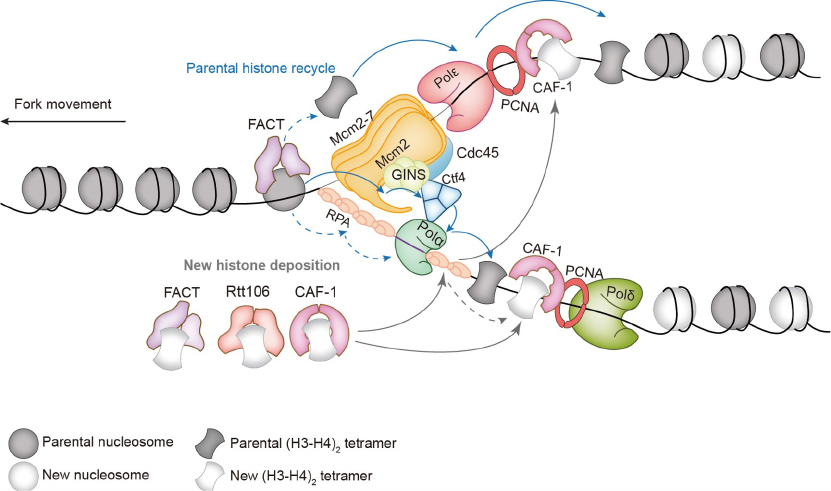
Stillman, 1986; Li et al., 2013) (Figure 2). Moreover, par-
ental (H3.1-H4)
2
tetramers remain intact and generally do
not split during DNA replication (Xu et al., 2010). Mean-
while, newly synthesized H3.1-H4 are deposited onto re-
plicating DNA in tetramer forms mediated by histone
chaperones (Fazly et al., 2012; Liu et al., 2012b; Su et al.,
2012). Therefore, parental and newly synthesized (H3.1-
H4)
2
tetramers form distinct nucleosomes following DNA
replication. On the contrary, newly synthesized H2A-H2B
could be found in nucleosomes containing parental H3-H4
2164
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
tetramers in one cell cycle, consistent with the idea that
nucleosomal H2A-H2B can exchange relatively freely with
parental H2A-H2B following DNA replication. Further-
more, deposition of H3-H4 tetramers is the rate-limiting step
of nucleosome formation (Smith and Stillman, 1991).
Therefore, we will focus on the discussion of replication-
coupled nucleosome assembly into three parts, dis-assembly
of preexisting nucleosomes (or parental nucleosomes) lo-
cated ahead of the replication fork, recycling of parental
histone H3-H4 tetramers, and deposition of newly synthe-
sized H3-H4 tetramers to form nucleosomes de novo.
Disassembly of parental nucleosomes
Previous studies reveal that approximately 300 bp of naked
DNA resides ahead of the replication forks, suggesting that
1-2 nucleosomes ahead of DNA replication forks are tem-
porarily disrupted (Lucchini et al., 2001). In Xenopus egg
extracts, using single-molecule imaging, it was reported that
nucleosome ahead of the replication fork is evicted and
parental histones are recycled (Gruszka et al., 2020). To-
gether, these studies support the idea that nucleosomes ahead
of DNA replication forks are disassembled temporarily.
Several factors are likely involved in the disassembly of
nucleosomes ahead of DNA replication forks. First, ATP-
dependent chromatin remodeling complexes, which utilize
the energy of ATP hydrolysis to alter the position of nu-
cleosomes along the DNA and to evict nucleosomal histones,
are likely involved in this process. Supporting this idea,
several chromatin remodeling complexes including INO80,
SWR1, ISW1 and ISW2 in budding yeast and their mam-
malian counterparts also participate in the DNA replication
process (Papamichos-Chronakis and Peterson, 2008; Vincent
et al., 2008; Morrison and Shen, 2009; Kurat et al., 2017).
However, to what extent that these chromatin remodeling
complexes remodel parental nucleosomes ahead of DNA
replication forks remains elusive. Second, the FACT (facil-
itates chromatin transactions) complex has been implicated
in remodeling nucleosomes ahead of DNA replication forks.
FACT, consisting of two subunits, Spt16 and Pob3 (SSRP1
in mammals), is a histone chaperone that binds to both H3-
H4 tetramers and H2A-H2B dimers (Belotserkovskaya and
Reinberg, 2004; Formosa and Winston, 2020). It has been
shown that FACT is essential for transcription on chromatin
template in vitro proposedly through removing H2A-H2B
from nucleosomes (LeRoy et al., 1998; Orphanides et al.,
1998). Recent studies using purified proteins in reconstituted
DNA replication system indicate that FACT is also essential
for DNA replication through chromatin template (Kurat et
al., 2017). Thus, FACT plays an important role in both DNA
replication and gene transcription through chromatin. In vi-
tro, FACT can alter the contacts between histones and DNA
without ATP hydrolysis. However, FACT itself could not
disassemble nucleosomes in vitro (Chen et al., 2018; Wang et
al., 2018b). Based on the Cryo-EM structures, FACT re-
cognizes partially unwrapped nucleosome structures (Liu et
al., 2020). In cells, FACT co-purifies with MCM2-7 complex
in both yeast and mammalian cells. FACT can also promote
DNA unwinding by MCMs in vitro (Gambus et al., 2006;
Tan et al., 2006). Together, these studies suggest that after
nucleosome disassembly, FACT may work with MCM he-
licase complex to facilitate nucleosome reassembly during
DNA replication (Figure 2). However, whether and how
FACT functions in parental nucleosome disassembly and
subsequent transfer of parental histones onto replicated DNA
remain unclear. Finally, Asf1, another histone chaperone
proposed to be involved in parental nucleosome disassembly,
is best known for its role in shuttling newly synthesized H3-
H4 in the process of de novo nucleosome assembly. It has
been shown that Asf1 co-purifies with MCM2–7 complex in
mammalian cells, and this interaction is bridged by histone
H3-H4 in the nucleus (Groth et al., 2007). A mutation on
Asf1-V94R, which disrupts Asf1 binding to H3-H4, also
compromise the Asf1-MCM interactions. Structure analysis
of the Asf1-H3-H4-MCM2 complex indicates that MCM2
N-terminus can bind to the H3-H4 tetramer and hijack the H3
interface involved in tetramer formation (Clément and Al-
mouzni, 2015; Huang et al., 2015). In cells, it has been
shown that histone chaperone Asf1 can facilitate nucleosome
disassembly at promoter region or gene body during tran-
scription (Adkins et al., 2004; Adkins and Tyler, 2004; Gao
et al., 2018). However, Asf1 cannot disassemble nucleo-
somes in vitro, indicating that other factors collaborate with
Asf1 to accomplish parental histone eviction in vivo (Don-
ham et al., 2011). Together, these studies suggest that mul-
tiple factors including chromatin remodeling complexes and
histone chaperones are likely involved in the disassembly of
nucleosomes ahead of DNA replication forks. However, to
what extent these factors function in nucleosome dis-
assembly and subsequent parental histone transfer remains to
be determined.
Parental histone transfer at the replication forks
Once parental nucleosomes ahead of replication forks are
disassembled, parental histones with modifications must be
transferred onto replicating DNA strands for the formation of
nucleosomes. This parental histone transfer and/or recycling
process is critical for the inheritance of histone modifica-
tions, but remains elusive for over 4 decades. For instance,
based on metabolic labeling of DNA and proteins during S
phase, it was proposed that parental histones are randomly
and equally distributed onto replicated DNA strands (Seale,
1976). Recent studies indicate that parental H3-H4 tetramers
likely remember their position along the DNA. These studies
are made possible with the development of novel techniques.
2165
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
For instance, by monitoring parental H3-H4 on a plasmid in
two different in vitro DNA replication systems, it has been
shown that parental H3-H4 are transferred locally in the
Xenopus DNA replication system, but are dispersed in SV40
DNA replication system (Madamba et al., 2017). The major
distinction between these two systems is that different heli-
cases are used in the DNA replication. In Xenopus extract,
CMG is the replicative helicase, whereas large T antigen is
the replicative helicase in the SV40 DNA replication system.
More recently, two studies show that parental nucleosomes
form positional memory following DNA replication (Esco-
bar et al., 2019; Schlissel and Rine, 2019). Both studies
started with labeling parental nucleosomes at a particular
locus covalently with biotin, and then tracked the fate of
labeled nucleosomes through DNA replication. In budding
yeast, it has been shown that labeled histone H3 can re-
member its positions along the DNA following replication
and gene transcription (Schlissel and Rine, 2019). In mouse
embryonic stem (ES) cells, by monitoring parental nucleo-
somal H3.1 that is enriched at silent chromatin regions, it has
been shown that parental H3.1 is transferred locally at re-
pressive regions, but is dispersed at actively transcribed re-
gions (Escobar et al., 2019). Of note, H3.3, but not H3.1, is
enriched at actively transcribed regions (Loyola and Al-
mouzni, 2007; Talbert and Henikoff, 2010). Therefore, it
would be interesting to determine whether parental H3.3 is
also transferred locally or dispersed at actively transcribed
regions.
Recent studies have discovered specific protein factors
involved in the transfer of parental H3-H4 onto replicating
DNA (Figure 2). First, it has been shown in both yeast and
mouse ES cells, mutations at the histone binding motif of
MCM2, a subunit of the CMG helicase, result in defects in
the transfer of parental H3-H4 to lagging strands of DNA
replication forks (Gan et al., 2018; Petryk et al., 2018). Early
studies indicate that human MCM complex binds to H3 and
H4 in HeLa cell extracts, and the N-terminus of mouse
MCM2 is required for the histone binding activity (Ishimi et
al., 1998; Ishimi et al., 2001). Similarly, the N-terminal
histone binding motif (HBM) of yeast Mcm2 was reported to
interact with all four histones released from chromatin
(Foltman et al., 2013). Interestingly, mouse MCM2 can bind
to H3-H4 and assemble a nucleosome-like structure in vitro,
supporting the idea that MCM2 histone binding domain also
possesses histone chaperone activity. Using the eSPAN
(enrichment and sequencing of protein-associated nascent
DNA) that measures the relative amount of parental and
newly synthesized histones at the leading and lagging strands
of DNA replication forks, it has been shown that parental H3
marked with H3K4me3 are transferred almost equally to
leading and lagging strands, with a slight preference for
lagging strands (Yu et al., 2018a). In contrast, new histones
marked by H3K56ac (acetylation on H3 lysine 56) showed
an opposite pattern. In cells with mcm2-3A mutation that
disrupts the interaction between Mcm2 and H3-H4, parental
H3K4me3 are enriched at leading strands due to defects in
the transfer of parental histones to lagging strands (Gan et al.,
2018). Similarly, using SCAR-seq (sister chromatids after
replication by DNA sequencing) in mouse ES cells with
mutations disrupting MCM2 binding to histones, marks on
Figure 2 DNA replication coupled nucleosome assembly pathways with key factors involved in nucleosome assembly indicated.
2166
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
parental histone show asymmetric distribution (Petryk et al.,
2018). These results show that the histone binding ability of
MCM2 is critical for parental histone transfer to lagging
strands of DNA replication forks.
The CMG helicase interacts with leading strand poly-
merase Polε and travels along with leading strand template
(Fu et al., 2011; Burgers and Kunkel, 2017). How does
MCM2, traveling along the leading strands, facilitates the
transfer of parental histones to the lagging strands of DNA
replication forks? To answer this question, it should be noted
that the CMG helicase interacts with Ctf4, which forms a
trimer that also interacts with Pol1, the catalytic subunit of
Polα primase enriched at lagging strands (Simon et al.,
2014). Studies from budding yeast show that mutations at
Ctf4 that cannot bridge the CMG-Pol1 interaction or Pol1
mutants that cannot bind to Ctf4 display similar defects in
parental histone transfer to lagging strands (Gan et al., 2018).
Finally, like Mcm2, Pol1 also contains a conserved histone
binding motif (Evrin et al., 2018). Both yeast and mouse Pol1
bind to H3-H4 preferentially over H2A-H2B. Mutations at
the histone binding motif of Pol1 also result in defects in
parental histone transfer in a manner similar to Mcm2 mutant
defective in histone binding (Li et al., 2020b). Together,
these studies indicate that Mcm2-Ctf4-Polα axis regulates
the transfer of parental histone H3-H4 to lagging strands of
DNA replication forks.
In budding yeast and mouse ES cells, using eSPAN ana-
lysis, it has been shown that deletion of Dpb3 (POLE4 in
mammals) or Dpb4 (POLE3 in mammals) leads to the dra-
matic reduction of the transfer of parental histones to leading
strands of DNA replication forks (Yu et al., 2018a). Dpb3
and Dpb4 are two subunits of leading strand DNA poly-
merase, Polε. However, Dpb3 and Dpb4 are not required for
enzymatic activity of Polε. Dpb3 and Dpb4 in fission yeast
form a dimer with the structure similar to H2A-H2B (He et
al., 2017). Moreover, Dpb3-Dpb4 co-purify with all four
core histones (Tackett et al., 2005) and interact with H3-H4
preferentially over H2A-H2B in vitro (Yu et al., 2018a).
Similarly, POLE3-POLE4 formed a stable dimer and could
bind to histone H3-H4 directly but not H2A-H2B (Bellelli et
al., 2018). Together, these studies indicate that Dpb3 and
Dpb4 serve as histone chaperones to promote the transfer of
parental histones to leading strands of DNA replication
forks.
Budding yeast cells with mcm2-3A mutation showed mild
defects in the loss of transcriptional silencing at hetero-
chromatin loci. Similar effects were also observed for cells
lacking Dpb3 and Dpb4 in both budding and fission yeast
(He et al., 2017; Yu et al., 2018a). Moreover, mcm2-3A dpb3
double mutant cells show defects in memory of nucleosome
positions following DNA replication (Schlissel and Rine,
2019). In mouse ES cells, the MCM2 and Polα mutants with
impaired parental histone transfer show defects in the re-
pression of ERVs (Li et al., 2020b). Together, these studies
indicate that the precise transfer of parental H3-H4 to re-
plicating DNA strands is important to maintain hetero-
chromatin states. Of note, both yeast and mouse ES cells
lacking these factors involved in parental histone transfer
have largely normal growth, suggesting that additional fac-
tors participate in the transfer of parental histones.
Deposition of newly synthesized histone H3-H4
After DNA duplication, parental histones contribute to only
half of the total histones required for the assembly of re-
plicating DNA into nucleosomes. Therefore, newly synthe-
sized histones are needed to complete the nucleosome
assembly of replicated DNA. Compared with the transfer of
parental histones, de novo deposition of new H3-H4 is re-
latively well studied (Serra-Cardona and Zhang, 2017). As
detailed below, de novo deposition of new H3-H4 requires a
group of histone chaperones that mediate histone folding,
import and deposition onto replicating DNA. Moreover,
modifications on newly synthesized H3-H4 also regulate the
interactions between histones and histone chaperones. Fi-
nally, these histone chaperones interact with components of
replisomes to facilitate the deposition of new H3-H4 onto
replicating DNA strands (Figure 2).
Histone chaperones form a coordination network for
deposition of new H3-H4
Histone chaperones are essential for de novo histone de-
position. These histone chaperones form a coordination
network for the deposition of newly synthesized H3-H4,
which first form a heterodimer. With the aid of other protein
chaperones involved in protein folding, new H3-H4 form a
complex with histone chaperone Asf1, which does not show
nucleosome assembly activity in vitro, indicating that Asf1
may not participate in the assembly event directly (Tyler et
al., 1999; English et al., 2005; English et al., 2006). Con-
sistent with this observation, the structure of Asf1-H3-H4
complex reveals that Asf1 binds to the H3-H4 dimer through
the H3 interface involved in the formation of H3-H4 tetra-
mers, and thus Asf1 blocks the H3-H4 tetramer formation
(English et al., 2006). Therefore, once associated with Asf1,
H3-H4 must be transferred to downstream chaperones in-
cluding chromatin assembly factor 1 (CAF-1) for deposition
onto replicating DNA.
CAF-1 was the first histone chaperone discovered in-
volved in replication coupled nucleosome assembly (Still-
man, 1986; Verreault et al., 1996; Kaufman et al., 1997).
CAF-1 consists of three subunits, Cac1, Cac2 and Cac3 in
yeast, corresponding to p150, p60 and p48 in mammalian
cells. One CAF-1 molecule binds to one H3-H4 dimer and
the dimerization of two CAF-1 complexes triggers the for-
mation of a H3-H4 tetramer (Liu et al., 2016; Mattiroli et al.,
2167
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
2017). Asf1 binds to the Cac2 subunit of histone chaperone
CAF-1 and the conformational changes allow the delivery of
H3-H4 dimer from Asf1 to CAF-1, thus providing direct
evidence for coordination between histone chaperones (Tyler
et al., 2001; Mello et al., 2002). In addition to direct inter-
action between Asf1 and CAF-1, previous studies suggest
that ubiquitination of H3K122 will destabilize the interaction
between Asf1 and H3-H4 complex, which in turn facilitates
the transfer of H3-H4 from Asf1 to CAF-1 (Han et al., 2013).
In yeast, yeast cells lacking CAF-1 are viable (Kaufman et
al., 1997), suggesting that other histone chaperones likely
promote deposition of new H3-H4 onto replicating DNA.
Indeed, it has been shown that Rtt106 (Regulator of Ty1
transposon 106) functions in parallel with CAF-1 in de-
position of new H3-H4 (Huang et al., 2005; Huang et al.,
2007). In addition to CAF-1 and Rtt106, using a separation
of functional mutant alleles, FACT has also been shown to
function in the deposition of newly synthesized H3-H4
during replication (Yang et al., 2016). FACT contains mul-
tiple PH (pleckstrin homology) domains and can bind to H3-
H4 with newly synthesized histone marks. Thus, multiple
chaperones function in the deposition of new H3-H4 onto
replicating DNA. Furthermore, in cells, these chaperones co-
purify with each other. For instance, FACT can co-purify
with CAF-1 and Rtt106, and the interaction between them is
bridged by H3K56Ac and peaks during S phase (Yang et al.,
2016). In addition, CAF-1 also co-purifies with Rtt106
(Huang et al., 2005). These physical interactions indicate that
these chaperones form a coordination network for de novo
histone deposition during S phase.
Histone modifications and variant amino acids on histone
proteins regulate the interaction between newly synthesized
histones and histone chaperones
Newly synthesized histones are also modified post-trans-
lationally and most of these modifications are distinct from
modifications on parental histones. For instance, acetyla-
tion of histone H4 lysine 5 and 12 (H4K5,12) by HAT1-
RbAp46 acetyltransferase and acetylation at some lysine
residues on H3 tails (H3K4,9,14,23,27) are marks on newly
synthesized histones across almost all species (Sobel et al.,
1995; Verreault et al., 1996). In fungal species, H3K56ac is
a mark on new H3 (Masumoto et al., 2005; Zhou et al.,
2006). H3K56ac is catalyzed by the Rtt109-Vps75 com-
plex and histone chaperone Asf1 is essential for H3K56
acetylation (Han et al., 2007a; Han et al., 2007b). The
structure of Rtt109 in complex with Asf1-H3-H4 indicates
that while Asf1 has little contact with Rtt109, it positions
H3 lysine 56 for acetylation by Rtt109 (Zhang et al., 2018).
In addition to histone acetylation, mono-methylation of
histone H3K9 (H3K9me1) by SETDB1 is also found on
H3.1 prior to deposition in mammalian cells (Loyola et al.,
2006).
Several functions have been uncovered for the modifica-
tions on newly synthesized H3-H4. First, the acetylation of
H4K5,12 occurs in cytoplasm and promotes the nuclear
import of histone H3-H4 mediated by histone chaperone
Asf1 and the Importin complex (Zhang et al., 2012; An et al.,
2017). Importin Kap123 contains two lysine binding pock-
ets, and acetylation at lysine residues on histone H3 and H4
weakens the interaction of H3-H4 with importin (An et al.,
2017). Second, H3K56 acetylation regulates the interactions
between H3-H4 and CAF-1 and Rtt106 (Chen et al., 2008; Li
et al., 2008). Moreover, acetylation at both H3 and H4 tails
also significantly increases the interaction of CAF-1 and
Rtt106 with new H3-H4 and promotes replication-coupled
nucleosome assembly (Burgess et al., 2010). Rtt106 contains
two tandem PH domains that likely bind to H3K56 acety-
lated H3-H4 (Su et al., 2012). However, how CAF-1 re-
cognizes H3K56ac and acetylates H3 and H4 tails remains to
be determined. Furthermore, it remains unclear whether
H3K56ac, which is present at low abundance in metazoans,
also has a role in replication-coupled nucleosome assembly.
Finally, it has been proposed that H3K9me1 helps the re-
storation of H3K9me2/me3 by serving as a substrate for
H3K9 methyl-transferases that catalyze di- and tri-methyla-
tion (Loyola et al., 2006). For a detailed description of his-
tone modifications’ role in replication-coupled nucleosome
assembly we refer readers to other reviews like “All roads
lead to chromatin”(Li et al., 2013).
In addition to histone modifications, variant amino acids
found on histone H3.1/H3.2 and H3.3 play a key role in
regulating the interaction between H3-H4 and the corre-
sponding histone chaperones. Histone H3.1/H3.2 differ from
H3 variant H3.3 by four or five amino acids, with the three
variant amino acids located at residues 87 to 90 (SAVM in
H3.1/H3.2 vs. AAIG in H3.3). H3.1/H3.2 bind to histone
chaperone CAF-1 and is deposited during S phase of cell
cycle in the replication-coupled nucleosome assembly
pathway (Ahmad and Henikoff, 2002a, 2022b). In contrast,
H3.3 associates with histone chaperones HIRA and DAXX,
and it can be deposited both during and outside of S phase
(Tagami et al., 2004; Drané et al., 2010; Goldberg et al.,
2010). Mutating the three variant amino acids between H3.1/
H3.2 and H3.3 can alter their interactions with CAF-1 and/or
HIRA/DAXX and subsequent deposition onto DNA (Lewis
et al., 2010; Elsässer et al., 2012; Liu et al., 2012a). Finally, it
has been shown that phosphorylation of H4 serine 47 inhibits
the interaction between CAF-1 and H3-H4 and promotes the
interaction between HIRA and H3-H4 (Kang et al., 2011).
Together, these studies indicate that modifications on newly
synthesized H3-H4 and variant amino acids on histone pro-
teins regulate the dynamic interactions between histone and
histone chaperones, thereby providing the supply of other
half of histones for the assembly of newly replicated DNA
into nucleosomes.
2168
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Histone chaperones connect to replication forks via inter-
actions with replisome components
How do histone chaperones deposit newly synthesized H3-
H4 specifically at replicated DNA? The answer to this
question lies at least partially in the physical interactions
between histone chaperones and replisome components.
Early studies showed that CAF-1 interacts with PCNA
(proliferating cell nuclear antigen) (Shibahara and Stillman,
1999). PCNA forms a homotrimer (Pol30 subunits in bud-
ding yeast) and functions as a sliding clamp for both Polδ
and Polε involved in lagging and leading strand DNA
synthesis, respectively (Choe and Moldovan, 2017). De-
pletion of PCNA inhibits CAF-1 mediated chromatin as-
sembly in vitro (Shibahara and Stillman, 1999).
Furthermore, site-specific PCNA mutations that disrupt the
CAF-1-PCNA interaction in budding yeast, while showing
minor effects on cell growth, result in defects in transcrip-
tional silencing, in the same pathway as cells lacking CAF-1
(Zhang et al., 2000). A recent discovery found that in-
troduction of the same PCNA mutations in mouse ES cells
led to defects in differentiation in vitro, and embryonic
lethality during mouse early development (Cheng et al.,
2019). Together, these studies suggest that the PCNA-CAF-
1 interaction is important for the deposition of new H3-H4
and embryonic development.
In addition to PCNA, RPA, the single-stranded DNA
binding protein at the replication forks, can also interact with
multiple histone chaperones. RPA contains three subunits
named as Rfa1, Rfa2 and Rfa3 in budding yeast or RPA70,
RPA32 and RPA14 in humans, respectively. RPA interacts
with histone chaperones FACT, CAF-1 and Rtt106, but not
Asf1 (Liu et al., 2017). Genetic analysis suggests the po-
tential coordination between FACT and RPA during nu-
cleosome assembly (VanDemark et al., 2006). Besides
histone chaperones, RPA also binds to free histone H3-H4
directly but not intact nucleosomes or H2A-H2B (Liu et al.,
2017). Furthermore, histone H3-H4 promotes the interaction
of RPA with those histone chaperones. Moreover, RPA can
also deposit H3-H4 onto adjacent double strand DNA when
bound to ssDNA, indicating a role of RPA in histone de-
position mediated by multiple histone chaperones (Liu et al.,
2017). Finally, it has been shown that FACT co-purifies with
MCM helicases in both yeast and mammalian cells (Gambus
et al., 2006; Tan et al., 2006). Together, these studies indicate
that histone chaperones involved in de novo deposition of
new H3-H4 interact with multiple components of replisomes,
which likely mediate the ability of these histone chaperones
to deposit H3-H4 in the DNA replication-coupled process.
However, the functional significance of several aforemen-
tioned interactions between histone chaperones and repli-
some components in replication-coupled nucleosome
assembly remains to be determined.
General principles for the restoration of histone
modifications following DNA replication
Early studies on X-chromosome inactivation, position effect
variegation in Drosophila, genome imprinting, silent chro-
matin at mating type locus in both budding and fission yeast
strongly support the idea that heterochromatin domains can
be inherited through mitotic cell divisions. These studies
were performed before the discoveries that distinct histone
modifications mark active and repressive chromatin domains
(Grewal and Jia, 2007).
It is well accepted that DNA methylation is heritable,
however, it is clear that not all histone modifications are
heritable for various reasons (Zhu and Reinberg, 2011;
Ptashne, 2013; Reinberg and Vales, 2018). Currently, it is
estimated that over 80–100 posttranslational modifications
on four histone proteins can be identified (Zhao and Garcia,
2015). Some of these histone modifications such as acet-
ylation are quite labile with a half-life less than one cell cycle
(Zee et al., 2010). Therefore, it is unlikely that those labile
histone modifications can be used as templates for the re-
storation of the modification following DNA replication
without the aid of other factors. In addition, it is known that
most nucleosomal H2A-H2B proteins exchange relatively
freely with newly synthesized H2A-H2B within one cell
cycle (Xu et al., 2010). Therefore, it is likely that most
modifications on H2A-H2B might not be heritable. Of note,
it has been recently shown that H2AK119 ubiquitination
located at repressive heterochromatin can be inherited (Zhao
et al., 2020a), suggesting that some H2A-H2B modifications
are heritable. Compared with H2A-H2B, H3-H4 tetramers,
once assembled into nucleosomes, are relatively stable and
do not exchange freely with newly synthesized H3-H4. In-
deed, methylation of H3 and H4, including H3K27me3 and
H3K9me2/m3, are widely accepted as inheritable epigenetic
marks (Margueron et al., 2009; Liu et al., 2015; Coleman and
Struhl, 2017; Laprell et al., 2017) and have a half-life over
one cell cycle. These findings suggest that histone mod-
ifications with longer half-life are more likely to be trans-
mitted following DNA replication. Moreover, it is known
that histone H3.1 at active or repressive chromatin regions
shows distinct patterns following DNA replication (Escobar
et al., 2019), suggesting that the heritability of histone
modifications likely also depends on local chromatin en-
vironment. Therefore, future studies are warranted to explore
the regulatory network that governs the inheritance of dif-
ferent epigenetic modifications.
An early insight into the inheritance of histone modifica-
tions came from studies on H3K27me3, which reported that
EED, a subunit of the PRC2 complex catalyzing H3K27me3
(Margueron and Reinberg, 2011; Holoch and Margueron,
2017), has a chromodomain that recognizes H3K27me3. In
vitro studies indicate that binding of H3K27me3 by EED
2169
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
stimulates the enzymatic activity of PRC2 to methylate
neighboring nucleosomes without this modification (Mar-
gueron et al., 2009). In mouse ES cells, mutations at EED
chromodomain impairing its binding to H3K27me3 result in
defects in the spreading of H3K27me3 (Oksuz et al., 2018).
Similarly, G9a/GLP, the methyltransferases for H3K9me2,
harbor ankyrin repeat domains, and the association of G9a/
GLP with H3K9me2 also stimulates their enzymatic activ-
ities. Mice with mutations at GLP ankyrin repeat show de-
fects in growth ossification and postnatal lethality (Liu et al.,
2015). Moreover, Suv39h1/h2, the enzymes catalyzing
H3K9me3 in mammalian cells, contain a chromodomain that
recognizes H3K9me3, although the functional significance
of this domain is not well explored. In fission yeast, the
recognition of H3K9me2/me3 by the chromodomain of Clr4,
the sole H3K9me3 writer, is important for inheritance of this
mark (Ragunathan et al., 2015; Wang and Moazed, 2017).
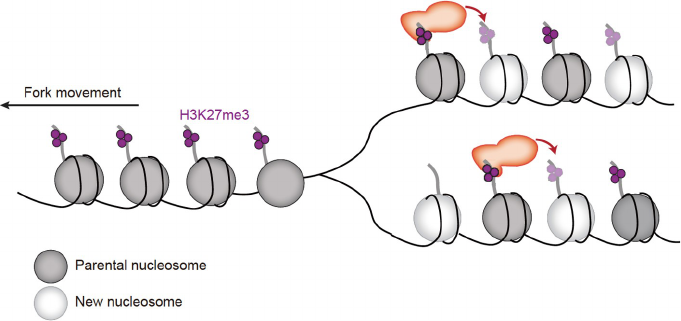
Together, these studies support a positive feedback model
whereby H3K9 or H3K27 enzymes first recognize (read)
their cognate modifications on nucleosomes from parental
histones and then modify (write) nucleosomes containing
newly synthesized histones without this mark following
DNA replication (Figure 3).
It was proposed that repressive marks including H3K9me3
and H3K27me3 are inheritable (see discussion below),
whereas active marks such as H3K4me3 are not (Reinberg
and Vales, 2018). However, in the literature, there are ex-
amples that active chromatin domains are also heritable. For
instance, it has been shown that the active gene state can
persist through 24 cell divisions in the absence of gene
transcription in nuclear transfer experiments and this epi-
genetic memory depends on the incorporation of H3.3, a
histone H3 variant marking actively transcribed genes, as
well as on H3.3 lysine 4 (Ng and Gurdon, 2008; Hörman-
seder et al., 2017). In C. elegans, mutations at H3K4me3
methyltransferases result in increased life span, and this in-
crease can be transmitted into descendants up to three gen-
erations, suggesting that certain chromatin loci marked by
H3K4me3 can be maintained trans-generationally (Greer et
al., 2011). In mouse mature oocytes, a non-canonical form of
H3K4me3 that contains broad H3K4me3 peaks at the pro-
moters and distal loci was discovered. These broad
H3K4me3 domains can be inherited in post-fertilization
embryos, before being erased at two cell embryo stages
(Dahl et al., 2016; Zhang et al., 2016). These studies strongly
suggest that active marks such as H3K4me3 may also be
inherited under certain conditions. Supporting this idea, the
Spp1 (CFP1 in humans), a subunit of the COMPASS com-
plex that catalyzes H3K4me3, also contains a PHD domain
that binds to H3K4me3 (He et al., 2019). Indeed, it has been
recently shown that both gene transcription machinery and
the read of H3K4me3 by Spp1 help recruit the COMPASS
complex for the restoration of H3K4me3 following DNA
replication (Serra-Cardona et al., 2022).
The read-write mechanism is just one part of the puzzles
for the restoration of histone modifications following DNA
replication. In fact, the inheritance of histone modifications
is much more complicated. For instance, it has been shown
that different histone modifications are restored on newly
synthesized histones at different rates following DNA re-
plication. Moreover, while restoration of histone modifica-
tions may start at S phase of the cell cycle, it takes until next
G1 for cells to fully restore most histone modifications (Xu
et al., 2011; Alabert et al., 2015). Furthermore, the cis-reg-
ulatory element called PRE involved in the establishment of
H3K27 methylation in early embryo is needed for the stable
maintenance of this mark, most likely through recruiting
PRC2 along with the read-write mechanism, to methylate
H3K27 in nucleosomes formed with newly synthesized H3-
H4 following DNA replication (Coleman and Struhl, 2017;
Laprell et al., 2017). Moreover, when the PRE is removed,
there is still considerable, residual capacity for copying the
mark, likely due to the function of the read-write mechanism
(Coleman and Struhl, 2017). In S. pombe and as described in
detail in the next section, both cis-regulatory elements and
RNAi machinery play important roles in the inheritance of
H3K9 methylation.
Several factors likely contribute to the complex nature for
the stable inheritance of histone modifications. First, com-
pared to DNA sequences, histone modifications are re-
versible due to the presence of eraser proteins, providing a
balance and competition between writers and erasers for a
particular histone modification. Therefore, in principle, cells
need to increase the local concentration of writers and/or
reduce the concentration of the erasers for the histone
modifications in order to faithfully maintain them during cell
division. Second, there are cross-talks among histone mod-
ifications at different chromatin regions. For instance,
H3K36 methylation, an active mark, can counterbalance
H3K27 methylation, a silent chromatin mark (Yuan et al.,
2011). Therefore, an increase in the concentration of writers/
erasers for H3K36 methylation can in principle influence the
dynamics of H3K27 methylation, or vice versa. Third, some
histone modifications such as H3K4me3 are deposited co-
transcriptionally (Soares et al., 2017; Bae et al., 2020).
Moreover, ongoing transcription can promote active histone
turnovers, i.e., exchange between parental histones and
newly synthesized histones. Therefore, it is proposed that
factors inhibiting histone turnover/exchange likely play an
important role in epigenetic inheritance (Aygün et al., 2013).
Finally, there are cross-talks between histone modifications
and DNA methylation (see detailed discussion below). Be-
cause of these complications, we propose that the restoration
of histone modifications following DNA replication requires
the interplay of histone modifications, cis-regulatory DNA
elements, non-coding RNA and DNA methylation.
2170
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Below, we use the inheritance of H3K9 methylation in S.
pome as the model to discuss these ideas for the following
reasons. First, key factors involved in H3K9 methylation and
heterochromatin assembly are highly conserved in higher
organisms. Second, the genetic power of yeast system allows
precise genetic manipulations. Third, heterochromatin pro-
teins are not essential for cell viability, allowing greater
flexibility for genetic analyses. Fourth, there is usually a
single gene encoding H3K9 heterochromatin regulators,
avoiding complications from multiple proteins with partially
overlapping functions. Finally, fission yeast does not have
DNA methylation. Together, this system makes it possible to
dissect the molecular mechanisms underlying the inheritance
of heterochromatin marked by H3K9 methylation.
Inheritance of H3K9 methylation, a lesson learned
from S. pombe
In fission yeast, large heterochromatin domains are present at
the pericentric region, silent mating-type region, and sub-
telomeres (Grewal and Jia, 2007). These regions all contain
repetitive DNA sequences, and the formation of hetero-
chromatin is critical for suppressing recombination between
repeats to maintain genome stability. Heterochromatin also
silences the transcription of genes within and near it in a
sequence-independent manner to regulate gene expression
programs.
Nucleosomes within these heterochromatic regions are
methylated at histone H3 lysine 9 (H3K9me). H3K9me re-
cruits heterochromatin protein 1 (HP1) family proteins Swi6
and Chp2, which in turn recruit diverse proteins to regulate
different biological processes (Grewal and Jia, 2007). Clr4 is
the sole histone H3K9 methyltransferase critical for hetero-
chromatin formation (Rea et al., 2000; Nakayama et al.,
2001), which contains a SET domain that catalyzes
H3K9me, and a chromodomain that recognizes H3K9me3
(Zhang et al., 2008). Mutations of the chromodomain that
affect Clr4 interaction with H3K9me3 reduced binding of
Clr4 to its target sites, and H3K9me3 domains are no longer
properly inherited. These results support the idea that Clr4
not only “writes” H3K9me3 but also “reads” it, forming a
positive feedback loop. The coupling of “read” and “write”
activities is also critical for restoring H3K9me3 domain after
DNA replication, where parental histones containing
H3K9me3 serve as seeds for the recruitment of Clr4 to
modify newly synthesized histones.
Early studies of heterochromatin at the silent mating-type
locus established that heterochromatin can be epigenetically
inherited, even before the role of histone H3K9 methylation
in heterochromatin assembly has been discovered. Fission
yeast has two different mating types: P (plus) and M (minus).
The mating type of a cell is determined by the gene content
within the mat1 locus, which is actively transcribed (Figure
4). Cells can also switch their mating types using one of the
two donor sequences, mat2P or mat3M, located more than 10
kilobases away from mat1. The donors, as well as the se-
quences between them, are silenced by heterochromatin.
Among the sequences between donors is cenH (centromere
homology), which is homologous to pericentric repeats.
Replacing cenH with a ura4
+
reporter gene leads to cells with
one of two stably maintained states: “ura4-on” (the reporter
is expressed) and “ura4-off” (the reporter is repressed)
(Grewal and Klar, 1996) (Figure 4A). Due to the low
switching rate from ura4-on to ura4-off, heterochromatin of
ura4-off cells is presumed to be maintained in the absence of
de novo establishment. Genetic analyses demonstrate that
these epigenetic states are inherited through both mitosis and
meiosis, behaving similarly to gene alleles (Grewal and Klar,
1996). Later it was demonstrated that the two epigenetic
alleles are different in their chromatin environments, such as
H3K9 methylation and Swi6 protein levels (Nakayama et al.,
2000; Hall et al., 2002).
The cenH sequence as well as pericentric repeats are later
Figure 3 The read and write mechanism contributes to the restoration of key histone modifications following DNA replication. Please note that the
restoration of histone modifications starts during S phase and may last till G1 phase of the cell cycle.
2171
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
found to initiate heterochromatin formation through the
RNA interference (RNAi) pathway (Hall et al., 2002; Volpe
et al., 2002). The DNA repeats are transcribed, generating
double-stranded RNAs (dsRNAs) (Volpe et al., 2002), which
are processed by the ribonuclease Dicer (Dcr1) into small
interfering RNAs (siRNAs). The Argonaute protein (Ago1)
within the RNA-induced transcriptional silencing complex
(RITS) binds to siRNAs and directs RITS to nascent RNA
transcripts originated from repeat regions (Motamedi et al.,
2004; Verdel et al., 2004). RITS then recruits the CLRC
complex, which contains the H3K9 methyltransferase Clr4,
to initiate H3K9me3 (Zhang et al., 2008; Bayne et al., 2010;
Gerace et al., 2010). Therefore, cenH is critical for the initial
RNAi-mediated targeting of Clr4 to the mating-type region
to establish heterochromatin. But once formed, this hetero-
chromatin is efficiently inherited by subsequent generations
even in the absence of cenH and RNAi.
However, such a simplified explanation is complicated by
later findings that transcription factors Atf1/Pcr1 recognize
target sequences within the silent mating-type region and
cooperate with RNAi to recruit Clr4 to establish hetero-
chromatin (Jia et al., 2004; Kim et al., 2004; Wang and
Moazed, 2017; Wang et al., 2021). Although Atf1/Pcr1
binding sites cannot independently initiate heterochromatin
formation, it still raises concern that removal of cenH does
not completely abolish heterochromatin establishment. To
precisely measure heterochromatin inheritance in the ab-
sence of de novo establishment, ectopic heterochromatin is
established by recruiting a TetR and Clr4-SET domain
(TetR-Clr4-SET) fusion protein to tetO binding sites, leading
to the silencing of adjacent report genes (Audergon et al.,
2015; Ragunathan et al., 2015) (Figure 4B). Releasing TetR-
Clr4-SET from tetO binding sites by the addition of tetra-
cycline allows the examination of heterochromatin main-
tenance through the self-templated restoration of H3K9me3
by endogenous Clr4. This artificial heterochromatin can
persist after TetR-Clr4-SET release, although only after re-
moving an anti-silencing protein Epe1. Moreover, the in-
heritance of such chromatin structure is dependent on the
ability of the Clr4 chromodomain to recognize H3K9me3
(Audergon et al., 2015; Ragunathan et al., 2015). These re-
sults clearly demonstrate that cells can indeed mediate epi-
genetic inheritance of H3K9me3 marked chromatin by
coupling the “reading” and “writing” of H3K9me3. How-
ever, they also indicate that this mechanism alone is not
sufficient to maintain heterochromatin states because of
other mechanisms that counter the inheritance of H3K9
methylation, such as Epe1.
Epe1 contains a JmjC domain, which typically catalyzes
histone demethylation (Tsukada et al., 2006). However, no
demethylase activity of Epe1 has been demonstrated in vitro,
and the commonly used mutations expected to abolish Epe1
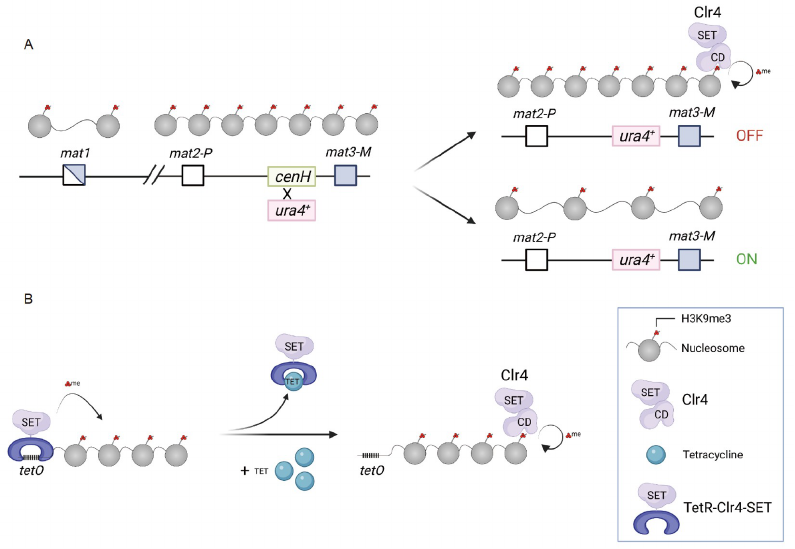
Figure 4 Epigenetic inheritance of H3K9me3 in S. Pombe. A, At the silent mating-type region, the replacement of cenH with a ura4
+
reporter results in two
metastable epigenetic states: ura4-on and ura4-off. The ura4-off state can be maintained during mitosis and meiosis through the coupling of “read-write”
activities of Clr4. B, The tetO sites recruit TetR-Clr4-SET to establish ectopic heterochromatin. The addition of tetracycline release TetR-Clr4-SET and
heterochromatin is maintained by endogenous Clr4.
2172
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
demethylase activity actually influence protein-protein in-
teractions (Raiymbek et al., 2020). Moreover, Epe1 is known
to exert its function on heterochromatin independent of its
JmjC domain (Wang et al., 2013; Bao et al., 2019; Sorida et
al., 2019). Therefore, the mechanisms whereby Epe1 coun-
teracts histone-based heterochromatin maintenance remain
unclear.
In addition to Epe1, other mechanisms that counteract
heterochromatin inheritance have also been uncovered. At
pericentric repeats, RNAi is the major pathway to establish
heterochromatin. However, heterochromatin is not properly
maintained in RNAi mutants, consistent with the existence of
mechanisms that counteract heterochromatin inheritance.
Interestingly, mutations of the Mst2 histone acetyltransferase
complex, INO80 chromatin remodeling complex, the Paf1C
complex associated with transcription, or Epe1 bypass RNAi
for heterochromatin inheritance (Trewick et al., 2007; Reddy
et al., 2011; Ragunathan et al., 2015; Sadeghi et al., 2015;
Shan et al., 2020). A common theme is that all of them
promote histone turnover at heterochromatin (Aygün et al.,
2013; Sadeghi et al., 2015; Wang et al., 2015; Shan et al.,
2020). A higher histone turnover rate leads to the loss of the
parental histones containing H3K9me3 at the original loca-
tion after DNA replication, therefore breaking the read-write
cycle for chromatin-based epigenetic inheritance (Shan et al.,
2021). As a result, RNAi is constantly needed to maintain a
high concentration of Clr4 to counteract the loss of parental
histones by histone turnover. Supporting this idea, the cou-
pling of siRNA production and H3K9me positive feedback
loops also promotes the inheritance of ectopic hetero-
chromatin induced by siRNAs (Yu et al., 2018b). Therefore,
faithful inheritance of H3K9me3 marked chromatin in fis-
sion yeast adopts multiple approaches including the read-
write mechanism (Wang et al., 2018a), inhibition of histone
turnover, and an increase in the local concentration of H3K9
methyltransferase via the RNAi machinery and DNA se-
quence-specific binding proteins.
DNA methylation inheritance including de novo
deposition and maintenance of DNA
methyltransferases
In mammals, DNA methylation primarily occurs on the fifth
position of cytosine (5-methylcytosine) in the palindromic
CpG context, and DNA methylation is one of the well-stu-
died epigenetic modifications. DNA methylation plays im-
portant roles in stably silencing the inactivated X
chromosome, repetitive elements, imprinting genes and de-
velopmental genes. Long-term transcription repression effect
of DNA methylation is mediated by recruiting methyl-CpG-
binding protein 2 (MECP2) in complex with histone deace-
tylases (HDACs), which reduce chromatin accessibility and
cause local condensation (Jones et al., 1998; Nan et al., 1998;
Muotri et al., 2010). Alternative silencing strategy of DNA
methylation is to prevent methylation-sensitive transcription
factors (TFs) from binding to their cognate sequences (Tate
and Bird, 1993). Besides, DNA methylation could stably
repress gene expression during mitosis and confer plasticity
upon stimulation, suggesting that DNA methylation can also
serve as an epigenetic marker for regulation of epigenetic
transcriptional memory described below.
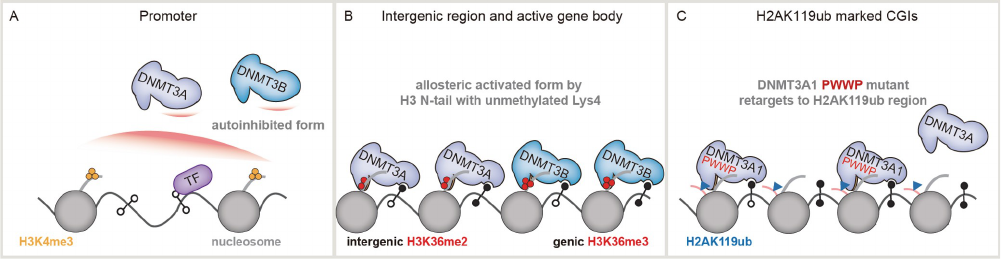
DNA methylation is catalyzed by DNA methyltransferases
(DNMTs) in mammalian cells, and cytosines in CpG palin-
drome can be unmethylated, hemi-methylated or fully-me-
thylated at various genomic regions. Based on the preference
of the methylated state of cytosine substrates, DNMTs are
classified into two groups: maintenance DNA methyl-
transferase (DNMT1) that shows selective activity toward
hemi-methylated CpG substrates, and de novo DNA me-
thyltransferases (DNMT3A, DNMT3B and rodent specific
DNMT3C) that exhibit comparable activity on both un-
methylated and hemi-methylated substrates (Figure 5).
In the early studies, DNMT1 was defined as maintenance
DNA methyltransferase based on its preferential activity on
hemi-methylated CpGs (Bestor et al., 1988; Pradhan et al.,
1999). However, recent studies challenged this simple clas-
sification model of DNMTs. Firstly, biochemical results
identified considerable de novo methyl-transfer activity of
DNMT1 on unmethylated substrates (Bestor and Ingram,
1983; Jeltsch and Jurkowska, 2014). Further in vivo studies
confirmed the notable de novo activity of DNMT1 enzyme.
In oocytes depleted of Stella, which is a maternal factor
essential for early development, DNMT1 was aberrantly
accumulated at vast chromatin regions, with significant de
novo methyl-transfer activities (Li et al., 2018b). This ac-
tivity was also observed in “Dnmt1” only oocytes (i.e., oo-
cytes with naturally silenced Dnmt3b and genetically
depleted Dnmt3a) (Li et al., 2018b), further confirming the
de novo activity of DNMT1. Furthermore, a well-designed
hairpin-bisulfite sequencing study identified about 0–5% de
novo activity of DNMT1 during replication-coupled phase
(Ming et al., 2021a). Moreover, although DNMT3s mainly
work as de novo methyltransferase, they lack selectivity to-
ward unmethylated and hemi-methylated CpG dinucleotide
substrates. Hairpin-bisulfite sequencing found significant
fully-methylated CpG sites in Dnmt1 depleted ESCs (Arand
et al., 2012), indicating DNMT3s also contribute to DNA
methylation maintenance in vivo. Finally, DNMT3 enzymes
could fill gaps caused by inefficiency of DNMT1 and
counteract demethylation mediated by ten-eleven transloca-
tion (TET) enzymes (Ramsahoye et al., 2000; Liang et al.,
2002; Jackson et al., 2004). Thus, DNMT1 and DNMT3
enzymes are both responsible for considerable de novo de-
position and maintenance of DNA methylation, at least in
some cellular contexts (Ming et al., 2021b). Consistent with
2173
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
this idea, these enzymes show context dependent functions in
mammalian development. It would be interesting for future
studies to dissect the contribution of de novo deposition and
maintenance activity of different DNMTs at different de-
velopmental stages or pathological contexts. We will focus
on discussing latest findings on mechanisms of DNA me-
thylation inheritance, the crosstalk between DNA methyla-
tion and histone modifications, and the role of DNA
methylation in epigenetic transcriptional memory.
Dynamics of DNA methylation inheritance
Globally, genomic DNA methylation exhibits a bimodal
distribution pattern in mammalian cells. Most genomic CpG
sites are hypermethylated, however, a fraction of CpG sites
residing in CG-dense DNA sequences named CpG islands
(CGIs) are generally hypomethylated (Cooper et al., 1983;
Bird et al., 1985). CGIs predominantly localize at regions
nearby the transcription start sites (TSSs) or promoters of
house-keeping genes and developmental genes. In addition,
many regulatory elements used to control gene expression
are largely resistant to CpG methylation (Hon et al., 2013;
Ziller et al., 2013; Rasmussen and Helin, 2016). The bimodal
landscape of mammalian methylome is a result of the dy-
namic balance between DNA methylation and demethylation
activities. During development, bulk genomic DNA methy-
lation pattern is static upon differentiation, and demethyla-
tion only occurs at specific sites in response to certain
cellular signals. In contrast, global DNA demethylation
happens in primordial germ cell (PGC) specification stage
and pre-implantation embryos to reset the methylome pat-
tern. In general, DNA methylation inheritance during mitotic
cell division is sophisticatedly regulated by multiple me-
chanisms: chromatin targeting and activity control of
DNMTs to counterbalance the effects of imperfect main-
tenance efficiency of DNMT1 and demethylation mediated
by TET family enzymes. Moreover, DNA methylation
maintenance is mainly a regional regulatory event (Wang et
al., 2020; Ming et al., 2021a) as local chromatin environment
including histone modifications and neighboring CpG state
(CpG densities and methylation levels) are important for the
dynamic turnover and inheritance of mammalian DNA me-
thylome.
Maintenance of DNA methylation during mitotic cell
division
Recent studies indicated that maintenance methylation oc-
curred quickly in a replication-coupled manner, with ap-
proximately 50% of the CpG sites methylated within minutes
and 80% of the sites within 30 min after DNA replication.
However, restoration of DNA methylation following DNA
replication also occured outside the S phase (replication-
uncoupled phase) (Figure 6) (Charlton et al., 2018; Xu and
Corces, 2018; Ming et al., 2021a). To achieve these, mam-
mals have evolved multiple mechanisms to ensure the fide-
lity and robustness of DNA methylation maintenance. For
instance, during mitotic cell division, the chromatin targeting
activity and protein stability of DNMT1 are regulated to
safeguard DNA methylation maintenance. Several key co-
factors, such as PCNA, LIG1 (DNA ligase 1), UHRF1
(ubiquitin-like with PHD and RING finger domains 1),
PAF15 (PCNA associated factor 15) and LSH (lymphoid-
specific helicase), are required for the maintenance role of
DNMT1. During replication-coupled phase, the highly effi-
cient maintenance activity of DNMT1 relies on its connec-
tion with DNA replication forks mediated by multiple
protein-protein interactions including PCNA-DNMT1,
UHRF1-LIG1 and ubiquitinated H3-DNMT1 interactions.
DNMT1 interacts with PCNA, a DNA clamp tethering DNA
polymerases to DNA replication forks, through PBD domain
(PCNA binding domain) (Chuang et al., 1997; Egger et al.,
2006). PAF15 contains a N-terminal H3-like sequence that
could be ubiquitinated at Lys 15 and Lys 24 by UHRF1 (Karg
et al., 2017), and ubiquitinated PAF15 binds to the replication
focus targeting sequence (RFTS) of DNMT1, which facil-
itates the association of DNMT1 with replisomes (González-
Magaña et al., 2019; Nishiyama et al., 2020). LIG1 is a
component of the replication machinery and is responsible
for ligating Okazaki fragments, of which the Lys 126 (K126)
Figure 5 Dynamic interplays between histone modifications and DNA methylation.
2174
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
and surrounding residues mimicked histone H3K9 site (Ferry
et al., 2017) that can be methylated by G9a/GLP. UHRF1
could bind to methylated LIG1 K126 to contact with re-
plication fork and promote maintenance of the lagging strand
(Ferry et al., 2017). These interactions together facilitate the
efficient methylation maintenance function of DNMT1 in the
replication-coupled phase (Figure 6A).
UHRF1 participates in DNA methylation maintenance
through both replication-coupled phase and replication-un-
coupled maintenance phase (Figure 6B) (Ming et al., 2021a).
Early studies found that UHRF1 was essential for genomic
DNA methylation inheritance by recruiting DNMT1 to me-
thylated sites (Bostick et al., 2007; Sharif et al., 2007).
UHRF1 contains several chromatin targeting domains,
which function cooperatively to promote proper chromatin
targeting of UHRF1 (Arita et al., 2012; Cheng et al., 2013;
Rothbart et al., 2013; Vaughan et al., 2018). The SET and
RING-associated (SRA) domain shows higher binding se-
lectivity towards hemi-methylated CpG sites (hemi-mCG),
which are established after DNA replication (Avvakumov et
al., 2008; Hashimoto et al., 2008; Qian et al., 2008; Fang et
al., 2016). The tandem tudor (TTD) domain of UHRF1
shows high affinity to H3K9me2/3 modification, and the
plant homeodomain (PHD) domain prefers the H3 N-tail
with unmethylated H3R2 (Rajakumara et al., 2011; Arita et
al., 2012; Cheng et al., 2013). Recognition of H3K9me2/3
modified nucleosome is mediated by cooperative binding of
TTD and PHD (Cheng et al., 2013; Rothbart et al., 2013).
Thus, UHRF1 could be recruited to DNA replication foci at
heterochromatin through the UHRF1-LIG1 interaction, re-
cognition of hemi-mCG by SRA domain and the interaction
between TTD-PHD domain of UHFR1 and H3K9me2/3.
Moreover, the TTD domain of UHRF1 and its recognition of
H3K9me2/3 modification are critical also for replication-
uncoupled DNA maintenance (Ming et al., 2021a). UHRF1
contains a RING-finger E3 ligase domain, which is re-
sponsible for histone H3 ubiquitination (Nishiyama et al.,
2013; Qin et al., 2015), and the hydrophobic patch of the
ubiquitin-like (UBL) domain of UHRF1 is required for ef-
ficient H3 ubiquitination, mainly through stabilizing the E2/
E3/chromatin complex (Foster et al., 2018). Early studies
proposed that UHRF1 facilitates methylation maintenance
activity of DNMT1 through recruiting of DNMT1 to re-
plication forks (Bostick et al., 2007; Sharif et al., 2007).
However, recent works indicated that ubiquitinated histone
by UHRF1 could also promote DNMT1 recruitment and
activate its methyltransferase activity (Nishiyama et al.,
2013; Qin et al., 2015; Ishiyama et al., 2017; Foster et al.,
2018; Li et al., 2018a). After DNA replication, many hemi-
CG sites were hindered by histones and other chromatin
proteins. Research found that chromatin remodeler LSH
might function in remodeling nucleosomal CpG sites to ex-
pose them to DNMT1 (Dennis et al., 2001). LSH could
promote nucleosomal CpG methylation maintenance in re-
plication-uncoupled phase, especially in heterochromatin
regions. Besides, LSH also associates with UHRF1, which
could assist the UHRF1-DNMT1 DNA methylation main-
tenance pathway(Han et al., 2020). Among all co-factors of
DNMT1, UHRF1 appears to be the most important co-factor
for the maintenance of DNA methylation, as Uhrf1 depletion
dramatically damages the pattern and kinetics of the main-
tenance of DNA methylation, comparable to Dnmt1 knock-
out.
Mechanisms for restricting excessive activity of DNMT1
Multiple mechanisms are evolved to restrict the activity of
DNMT1 to avoid accumulation of aberrant DNA methyla-
tion at unmethylated sites through its de novo methylation
activity. First, the protein levels of both DNMT1 and its key
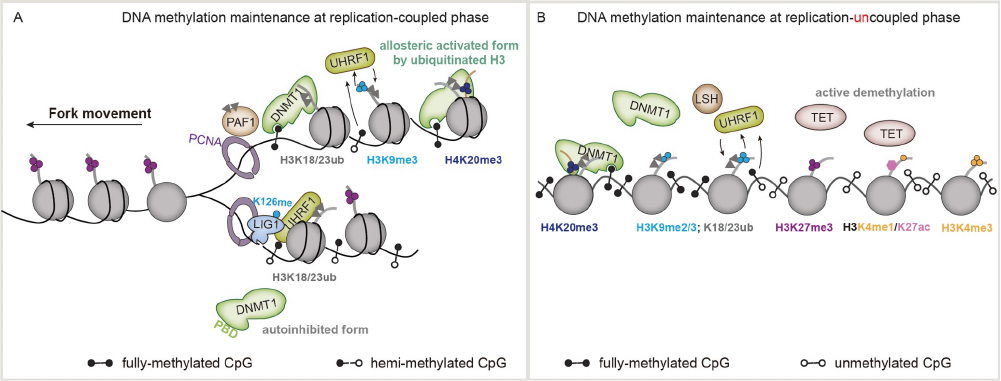
Figure 6 Epigenetic inheritance of DNA methylation.
2175
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
cofactor UHRF1 are cell-cycle regulated. It was reported that
DNMT1 stability is regulated by a series of integrated post-
translational modifications, including methylation (Estève et
al., 2009; Estève et al., 2011), acetylation (Du et al., 2010),
and phosphorylation (Estève et al., 2011), which co-
ordinately determine DNMT1 ubiquitination levels and
protein stability. Methyltransferase SET7/9 methylates
DNMT1 and triggers poly-ubiquitination and degradation of
DNMT1 (Estève et al., 2009; Estève et al., 2011), whereas
lysine-specific demethylase 1 (LSD1) stabilizes DNMT1
proteins, likely through demethylation (Zhang et al., 2019).
Previous studies reported that AKT1 kinase phosphorylates
DNMT1 Ser143 and interferes with lysine 142 methylation
by SET7/9 (Estève et al., 2011). Therefore, the interplay of
these two modifications affects cellular DNMT1 stability.
Another study identified that cell-cycle regulated methyl-
transferase SET8/PR-Set7 could control the stability of both
DNMT1 and UHRF1 through its methylation activity, fol-
lowed by subsequent poly-ubiquitination mediated protein
degradation (Zhang et al., 2019). SET8/PR-Set7 could down-
regulate UHRF1 in G2/M phase, causing repression of the
activity of DNMT1 on post-replicated DNA (Zhang et al.,
2019). Thus, SET8/PR-Set7 and LSD1 compete to regulate
genomic DNA methylation, most likely through regulation
of UHRF1 protein levels. DNMT1 could be acetylated by
acetyltransferase TIP60 for subsequent ubiquitination and
proteasomal degradation (Du et al., 2010). On the contrary,
histone deacetylase 1 (HDAC1) and deubiquitinase could
stabilize cellular DNMT1 levels (Du et al., 2010; Cheng et
al., 2015). The acidic pocket in ubiquitin-specific protease 7
(USP7) interacts with lysine residues within KG linker of
DNMT1, and this interaction is important for USP7 mediated
DNMT1 stabilization (Cheng et al., 2015). Acetylation of
lysine residues in DNMT1 KG linker interferes its binding to
USP7, thus promoting ubiquitination and degradation of
DNMT1 (Cheng et al., 2015). Besides, multiple-interaction
networks among DNMT1, UHRF1, PCNA, LIG1, PAF15,
LSH and histone H3 ubiquitination not only facilitate the
sophisticated contact of DNMT1 with the replication fork in
the replication-coupled phase, but also promote its targeting
to sites for methylation in the replication-uncoupled phase.
For instance, the specific binding between the SRA domain
of UHRF1 and hemi-methylated CpG ensures DNMT1 to
predominantly function as a maintenance methyltransferase.
The interaction between the TTD-PHD module of UHRF1
and H3K9me2/3 helps to confer some degree of targeting
specificity for DNMT1 (Cheng et al., 2013; Rothbart et al.,
2013). Importantly, DNMT1 mediated methylation main-
tenance heavily relies on H3 ubiquitination, which has a fast
turnover rate and is removed by USP7 after DNMT1 re-
cruitment (Nishiyama et al., 2013; Yamaguchi et al., 2017).
Interestingly, the de novo methylation activity of DNMT1
has to be tightly controlled. For example, during oocyte
maturation, Stella is required to prevent aberrant accumula-
tion of DNA methylation mediated by the de novo methy-
lation activity of DNMT1, via disrupting the chromatin
association of UHRF1 (Li et al., 2018b; Du et al., 2019). It is
interesting to investigate whether the de novo methylation
activity of DNMT1 is also under tight control in other post-
mitotic cells that may allow aberrant methylation accumu-
lation by the weak de novo methylation activity of DNMT1,
especially during aging. Finally, DNA methylation main-
tenance efficiency is affected by the methylation levels of
nearby CpG sites, which ensures the robustness in main-
taining a bistable system that allows faithful maintenance of
highly methylated regions and unmethylated regions, but not
intermediately methylated regions (Ming et al., 2021a).
Crosstalk between DNA methylation and histone
modifications
Somatic DNA methylation is set up de novo at early embryo
development and maintained during subsequent mitotic cell
cycles. Recent studies revealed complicated interplay be-
tween DNA methylation and histone modifications. As de-
tailed below, some histone modifications help to recruit
DNMTs to certain genomic regions and boost their methyl-
transfer activities, while others exclude them from chromatin
with certain genomic features and suppress their catalytic
abilities. Moreover, transitions from histone modification to
DNA methylation were also observed in some processes
including X-chromosome inactivation. For instance, during
early stage of X-chromosome inactivation, H3K27 methy-
lation by PRC2 complex silences one of the two X chro-
mosome in female mammals, and this silencing mechanism
is replaced by promoter DNA methylation during later stage
of this process (Avner and Heard, 2001; Csankovszki et al.,
2001; Sado et al., 2004; Disteche and Berletch, 2015; Pinter
et al., 2012). Interestingly, inactivated X-chromosome was
globally hypomethylated due to reduced H3K36 methylation
caused by transcriptional silencing, except for promoter re-
gions.
H3K4 methylation counters DNA methylation
DNA methylation is largely excluded from H3K4 methyla-
tion marked regions, such as active gene promoters and en-
hancers. In recent years, a series of studies revealed the
underlying mechanism whereby H3K4 methylation repels
the deposition of DNA methylation. DNMT3L is an en-
zymatically inactive homolog of DNMT3A and DNMT3B,
and is required for establishing the DNA methylation land-
scape during gametogenesis (Bourc’his et al., 2001). More-
over, DNMT3L was reported to stimulate the activity of
DNMT3A (Chedin et al., 2002). Structure analysis indicates
that DNMT3L forms a functional heterotetramer with
DNMT3A to promote de novo DNA methylation (Jia et al.,
2176
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
2007), and this interaction also prevents the oligomerization
of DNMT3A (Jurkowska et al., 2011). DNMT3L binds to the
N-terminal H3 sequence, and this interaction was specifi-
cally suppressed by H3K4 methylation both in vitro and in
vivo (Figure 5A) (Ooi et al., 2007). The DNMT3A-
DNMT3L structural data indicates that DNMT3L binds to
unmethylated H3 tail and promotes de novo DNA methyla-
tion through either enhancing the recruitment or activation of
DNMT3A (Ooi et al., 2007). Another structural study found
that the ATRX-DNMT3-DNMT3L (ADD) domain of
DNMT3A also specifically binds to H3 N-tail without H3K4
methylation (Otani et al., 2009), suggesting ADD might re-
cognize the unmethylated state of H3K4 and help DNMT3A
to target chromatin properly. Methylation analysis using in
vitro reconstituted chromatin showed that full-length
DNMT3A and full-length DNMT3A/3L complexes methy-
late DNA, preferentially at linker DNA regions, of H3K4-
unmethylated chromatin more efficiently than H3K4me3
marked chromatin (Zhang et al., 2010). The authors con-
cluded that the improved activity of DNMT3A on H3K4-
unmethylated chromatin was due to the selective binding
property of ADD domain to H3K4-unmethylated region.
Supporting this idea, the activity of catalytic domain of
DNMT3A was not affected by H3K4me3 per se (Zhang et
al., 2010). However, it is still unclear whether unmethylated
H3 tail could induce allosteric activation of functional
DNMT3A complex. Strikingly, an independent study
showed that the activity of DNMT3A was stimulated by up to
8-fold by H3K4-unmethylated H3 tail (Li et al., 2011).
However, the underlying molecular mechanism of allosteric
regulation remains elusive. The autoinhibitory DNMT3A-
DNMT3L complex and catalytically active DNMT3A-
DNMT3L-H3 complex helped to clarify this issue. In the
autoinhibitory structure, ADD domain of DNMT3A sup-
presses the methylation activity of DNMT3A by binding to
catalytic domain (CD) of DNMT3A and thereby blocking
DNA binding of CD (Guo et al., 2015). Histone H3 tail with
unmethylated state of H3K4 specifically disrupts the ADD-
CD interaction, and therefore release the autoinhibitory ef-
fects of DNMT3A (Guo et al., 2015). These studies provide a
new insight in understanding the mutually exclusive geno-
mic distribution of DNA methylation and H3K4 methyla-
tion. H3K4 methylation at promoters might be a potential
mechanism for excluding DNA methylation at CGIs, how-
ever, mechanisms that maintain the hypomethylated state of
CGIs are still elusive. It is proposed that transcription factors
(Brandeis et al., 1994; Macleod et al., 1994), active de-
methylation by TET enzymes (Williams et al., 2012; Putiri et
al., 2014; Verma et al., 2018) and skewed GC distribution
nearby TSS (Ginno et al., 2012) all contribute to this process.
In contrast to Dnmt3a or Dnmt3b deficient mice, Dnmt3l
knockout mice are viable (Okano et al., 1999; Bourc’his et
al., 2001). The functional requirement of DNMT3L for de
novo methylation in vivo is still not fully answered.
Although DNMT3A is generally depleted at CGIs, it has
been found that mutations at PWWP domain result in re-
distribution of DNMT3A to genomic regions including CGIs
marked by ubiquitinated H2AK119 (Figure 5C) (Remacha et
al., 2018; Heyn et al., 2019; Weinberg et al., 2021). The
amino terminus of DNMT3A1 interacts with H2AK119ub-
marked nucleosomes, serving as another chromatin targeting
strategy for DNMT3A1 (Weinberg et al., 2021). This novel
interaction explains the aberrant genomic distribution of
DNMT3A and hypermethylation at Polycomb-regulated re-
gions in paragangliomas and microcephalic dwarfism con-
taining mutations at the PWWP domain of DNMT3A. In the
future, it would be interesting to determine whether the N-
terminus of DNMT3B and the tissue-specific or cancer-
specific DNMT3A/3B isoforms (Chen et al., 2002; La Salle
and Trasler, 2006; Gopalakrishnan et al., 2009; Duymich et
al., 2016) also utilize similar strategies for chromatin tar-
geting.
Different methylation state of H3K36 dictates chromatin
targeting of DNMT3A and DNMT3B
For many years, the PWWP domain of DNMT3A/3B has
been linked to chromatin targeting. PWWP domain, which
contains a conserved aromatic cage for recognition of me-
thylated lysine residue, is important for protein chromatin
targeting through synergistic binding of histone and DNA
(Wu et al., 2011; Qin and Min, 2014; Dukatz et al., 2019).
Early study demonstrated PWWP domains are important for
targeting DNMT3A/DNMT3B to major satellite regions
(Chen et al., 2004; Ge et al., 2004). Loss of DNA methyla-
tion at satellite sequences was found in genetic diseases
bearing PWWP mutations, such as ICF (immunodeficiency,
centromere instability and facial anomalies) syndrome
(Shirohzu et al., 2002). Together, these reports link the
chromatin recruitment of DNMT3A/3B through the PWWP
domain. Biochemical and structural studies indicate that
PWWP domain of DNMT3A/3B interacts with H3K36me3
(Dhayalan et al., 2010; Qin and Min, 2014; Rondelet et al.,
2016). Genomic binding profile studies found that DNMT3B
is preferentially recruited to transcribed gene bodies, which
are enriched with active H3K36me3 mark mediated by SET
domain containing 2 (SETD2) (Figure 5B) (Sun et al., 2005;
Edmunds et al., 2008; Baubec et al., 2015). SETD2 and
PWWP domain are both required for proper targeting of
DNMT3B to active gene bodies (Baubec et al., 2015),
highlighting the importance of H3K36me3-PWWP interac-
tion for de novo methylation. However, DNMT3A, different
to DNMT3B, could interact with both di- and tri-methylated
state of histone H3K36, and shows a higher binding affinity
towards H3K36me2 (Weinberg et al., 2019). H3K36me2 is
distributed at both intergenic regions and gene bodies, and is
catalyzed by two NSD histone methyltransferase family
2177
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
enzymes, such as NSD1 and NSD2 (Kuo et al., 2011).
Genome wide analysis demonstrated that DNMT3A is tar-
geted to intergenic regions through DNMT3A-H3K36me2
interaction, which can be mistargeted to H3K36me3 mod-
ified gene bodies in cells depleted of Nsd1 and Nsd2
(Weinberg et al., 2019). Thus, the PWWP domains of
DNMT3A and DNMT3B recognize different methylation
state of H3K36 and target these enzymes for methylation at
different chromatin regions.
Relationship between H3K27me3 and DNA methylation
Mammalian H3K27me3 modification is catalyzed by Poly-
comb repressive complex 2 (PRC2) (Margueron and Rein-
berg, 2011). H3K27me3 is almost exclusively associated
with CGI regions, which is generally hypomethylated in
embryonic stem cells (ESCs) (Cooper et al., 1983; Bird et al.,
1985; Ku et al., 2008), indicating the mutually exclusive
distribution of H3K27me3 and DNA methylation in ESCs.
This might be partially explained by the dependency of DNA
methylation on H3K36me2/3, and the antagonizing dis-
tribution pattern of H3K36 and H3K27 methylation (Papp
and Müller, 2006; Schmitges et al., 2011; Yuan et al., 2011;
Gaydos et al., 2012; Popovic et al., 2014; Lu et al., 2016;
Huang and Zhu, 2018). Further studies focused on the PRC2
accessory proteins uncovered a more direct molecular me-
chanism underlying the mutually exclusive distribution of
H3K27me3 and DNA methylation. PRC2 accessory pro-
teins, such as PHF1, MTF2, JARID2, AEBP2 and PHF19,
are likely involved in the recruitment and/or regulation of
enzymatic activity of PRC2 (Cao et al., 2008; Boulay et al.,
2011; Casanova et al., 2011; Ballaré et al., 2012; Brien et al.,
2012; Hunkapiller et al., 2012; Oksuz et al., 2018; Youmans
et al., 2018; Højfeldt et al., 2019). Structural analysis iden-
tified the N-terminus of PHF1 and MTF2 bind to un-
methylated CpG motif (Li et al., 2017), highlighting a
potential mechanism for the restrictive recruitment of PRC2
to these unmethylated regions. In addition, PRC2 was also
functionally associated with TET1 enzyme (Neri et al.,
2013). Collectively, these studies uncovered the mutually
exclusive distribution of H3K27me3 and DNA methylation,
and the underlying potential molecular mechanisms. How-
ever, H3K27me3 and DNA methylation do overlap at some
genomic regions in certain somatic and cancer cells
(Brinkman et al., 2012; Statham et al., 2012). During dif-
ferentiation and carcinogenesis, H3K27me3-silenced gene
promoters contain DNA methylation in some sites (Ohm et
al., 2007; Schlesinger et al., 2007; Mohn et al., 2008; Rose
and Klose, 2014; Chen et al., 2019; Sendžikaitė et al., 2019).
Moreover, it has been shown that during early phase of X-
chromosome inactivation, expression of Xist RNA recruits
Polycomb complex for gene silencing. Subsequently, the
H3K27me3 silencing mechanism is switched to promoter
DNA methylation for long term silencing (Augui et al., 2011;
Jégu et al., 2017; Galupa and Heard, 2018). However, the
inactivated X-chromosome exhibits a global DNA hypo-
methylated state due to transcription silencing and reduced
H3K36 methylation. Therefore, the relationship and transi-
tion between DNA and H3K27 methylation are complicated,
which requires further mechanistic studies.
Crosstalk between H3K9me2/3 and DNA methylation
There is a direct cross talk between H3K9 methylation and
DNA methylation. H3K9 methylation is required for all
DNA methylation in Neurospora crassa (Tamaru and Selker,
2001; Tamaru et al., 2003). In Arabidopsis thaliana CpNpG
methylation is also dependent on H3K9 methylation (Jack-
son et al., 2002). Albeit lack of a strict link between H3K9me
and DNA methylation deposition in mammals, these two
repressive modifications are co-localized at heterochromatin
regions. Early studies proposed that DNMT3A/3B could
deposit DNA methylation through binding to hetero-
chromatin protein 1 (HP1) that recognizes H3K9me3 (Leh-
nertz et al., 2003). Further, several studies describe direct
interactions between DNMT3A/3B and the H3K9 methyl-
transferases, such as SUV39H1 (Fuks et al., 2003), SETDB1
(Li et al., 2006) and G9a/GLP (Epsztejn-Litman et al., 2008;
Chang et al., 2011). However, the functional significance of
these interactions to promote DNA methylation has not yet
been fully studied.
Compared with de novo DNA methylation, DNA methy-
lation maintenance mediated by DNMT1-UHRF1 machinery
shows a closer connection with H3K9me2/3 modification.
As mentioned above, the TTD and PHD domains of UHRF1
cooperatively bind to H3K9me2/3 modification and show a
preference on trimethylated state of H3K9 (Figure 6A)
(Hashimoto et al., 2009; Rottach et al., 2010). Due to the
technical limitations of knockdown and overexpression ex-
periments, TTD was thought to be essential for UHRF1
chromatin targeting and DNA methylation maintenance in
early studies (Rothbart et al., 2012; Rothbart et al., 2013).
However, genome edited homozygous TTD mutant mice
only shows about 10% reduction of DNA methylation in
various tissues tested (Zhao et al., 2016), indicating limited
role of TTD for DNA methylation maintenance. These works
all focused on the readout of global methylation level, but
little is known about the contribution of TTD in the kinetics
of DNA methylation maintenance. Recent kinetic analysis
demonstrated that TTD and H3K9me2/3 are important for
replication-uncoupled DNA methylation maintenance (Ming
et al., 2021a). Besides, RFTS domain of DNMT1 could di-
rectly recognize H3K9me3 and facilitate DNA methylation
maintenance (Ren et al., 2020). DNMT1 was also reported to
directly interact with G9a to promote its maintenance effi-
ciency during replication (Estève et al., 2006). Together,
these studies highlight the importance of direct crosstalk
between H3K9 methylation and DNA methylation.
2178
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
H3 ubiquitination facilitates the recruitment and activation
of DNMT1
Detailed molecular mechanisms of the UHRF1-DNMT1
maintenance apparatus in DNA methylation inheritance were
carefully illustrated until recent years. Histone H3 was
identified as a ubiquitination target of UHRF1 using Xenopus
egg extracts (Nishiyama et al., 2013); and both the PHD and
RING domains are important for efficient ubiquitination of
H3 (Qin et al., 2015). Mass spectrometry analysis identified
that H3 could be ubiquitinated at Lys14 (Ishiyama et al.,
2017), Lys18 (Qin et al., 2015; Ishiyama et al., 2017) and
Lys23 (Nishiyama et al., 2013; Ishiyama et al., 2017), and
these ubiquitinated sites could be recognized by the ubiquitin
interacting motif (UIM) of RFTS domain in DNMT1 protein
(Qin et al., 2015). Crystal structure identified a novel re-
cognition mode of RFTS which simultaneously binds to
double ubiquitinated H3 at K18 and K23 (Ishiyama et al.,
2017). It was also reported that RFTS domain mediates the
homodimerization (Fellinger et al., 2009) and autoinhibition
(Syeda et al., 2011; Takeshita et al., 2011) of DNMT1.
Strikingly, DNMT1 opens its active site upon binding to
H3K18ub/K23ub by the RFTS domain, indicating that ubi-
quitinated histone by the UHRF1 could allosterically activate
the activity of DNMT1. Therefore, these works together
demonstrated the essential role of the ubiquitin-binding
module of DNMT1 in DNA methylation maintenance (Ish-
iyama et al., 2017).
A role of H4K20 methylation in recruiting DNMT1 to
LINE-1 region
Generally, DNA methylation, H3K9me3 and H4K20me3 co-
exist at many heterochromatin regions. These repressive
marks function cooperatively to silence repetitive DNA se-
quences in mammalian genomes. Compared with the well-
known connections between DNA methylation and
H3K9me3, the crosstalk between DNA methylation and
H4K20me3 is less reported. Disturbance of DNA methyla-
tion and H4K20me3 frequently occurs in cancer cells
(Feinberg and Vogelstein, 1983; Eden et al., 2003; Fraga et
al., 2005). Reactivation of repetitive elements, especially
long interspersed nuclear element-1 (LINE-1), is tightly as-
sociated with genome rearrangements in cancers (Rodriguez-
Martin et al., 2020), highlighting the importance of the re-
pression of repetitive LINE-1. DNMT1 contains two bromo-
adjacent homology (BAH) domains (Yarychkivska et al.,
2018), which have been recently shown to specifically re-
cognize methylated H4K20 with a preference for H4K20me3
(Ren et al., 2021). Furthermore, H4K20me3 bound to BAH1
domain could induce an allosteric stimulation of DNMT1
activity (Ren et al., 2021), and the BAH1-H4K20me3
binding module facilitates DNA methylation maintenance
especially for the LINE-1 elements (Ren et al., 2021). This
work provides a direct crosstalk between DNA methylation
and H4K20me3. Thus, RFTS and BAH1 domains of
DNMT1 bind to H3ub/H3K9me3 and H4K20me3, respec-
tively, highlighting multivalent communications among re-
pressive marks and DNA methylation maintenance.
Nevertheless, it remains unclear whether the de novo activity
of DNMT1 participates in DNA methylation at LINE-1 re-
gions. Furthermore, novel crosstalk between histone mod-
ifications and different domains of DNMTs warrants further
investigation.
Roles of DNA methylation in transcriptional
memory
Epigenetic memory of gene transcription was described as a
heritable change in gene expression or behavior that is in-
duced by an experienced stimulus (D’Urso and Brickner,
2014). Epigenetic memory could be set up and maintained by
various epigenetic players, including DNA methylation,
histone modifications, histone variants and chromatin re-
modelers. Epigenetic memory can be divided into cellular
transcriptional memory and transgenerational memory based
on the time scales of memory maintained. Cellular tran-
scriptional memory refers to the mitotically heritable tran-
scriptional state in response to development cues or
environmental stimuli, while the transgenerational memory
describes meiotically heritable transcriptional profile gen-
erated by experiences of previous generations (D’Urso and
Brickner, 2014). Adaptive immunity, chronic inflammation,
and neuronal memory are ideally suitable contexts for
studying the molecular mechanisms underlying the estab-
lishment and maintenance of transcriptional memory.
Transcriptional memory allows certain genes to respond
more rapidly and robustly toward previously experienced
signals (Bergink et al., 1973). During past years, transcrip-
tional memory of the inducible inositol-1-phosphate syn-
thase (INO1) and galactokinase (GAL) genes in
Saccharomyces cerevisiae system are thoroughly studied. It
has been shown that several factors/players including nuclear
periphery retention, intragenic looping, H2A.Z variant de-
position, H3K4 methylation and chromatin remodeler SWI/
SNF are important for transcriptional memory. In addition to
histone related players, transcriptional memory of Tat gene
upon glucocorticoid induction is associated with DNA de-
methylation event (Thomassin et al., 2001), and similar de-
methylation was also reported at IL2 gene locus after T cell
activation (Murayama et al., 2006). These studies imply the
biological significance of DNA demethylation on transcrip-
tional memory establishment and maintenance. Recent
published works further demonstrated the importance of
DNA methylation in transcriptional memory regulation.
Although short-term treatment of tumor necrosis factor α
(TNF-α) could activate DNA methylation silenced IL32 gene
2179
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
without demethylation step, prolonged TNF-α treatment in-
duces DNA demethylation at both the promoter and CGI
region of IL32 gene which depends on TET and p65 (Zhao et
al., 2019). Strikingly, demethylation-induced transcriptional
activation of IL32 persists for a long time after withdrawing
of TNF-α (Zhao et al., 2019). Moreover, sustained TNF-α
administration uncovers a transcriptional memory induced
by the key proinflammatory cytokine TNF-α (Zhao et al.,
2020b). CALCB gene, the key therapeutic target gene in
migraine, shows the strongest transcriptional memory and
relies on the active demethylation mediated by TET enzymes
(Zhao et al., 2020b). These results suggest that inflammatory
signals and memory consolidation might play a role in the
development of chronic migraine. Collectively, these works
demonstrated that transcriptional memory provoked by TNF-
α is governed by active DNA demethylation by TET en-
zymes. The hypomethylated state of memory gene and re-
lated regulatory region might facilitate the chromatin binding
of subsequent methylation-sensitive transcription factors,
which in turn provoke rapid and robust transcription acti-
vation in the subsequent encounter of inflammatory stimuli.
Notably, it is intriguing whether DNA demethylation medi-
ated transcriptional memory towards certain environmental
and cellular stimuli might also be involved in the develop-
ment of adaptive immunity malfunction, chronic inflamma-
tion, aging and cancer.
Functional impact of epigenetic inheritance
Factors discussed above involved in epigenetic inheritance are
important to maintain chromatin states and cell identity.
Therefore, it is easy to envision the critical roles of epigenetic
inheritance in cell identity during normal development, in
disease evolution and in response to environmental stress/cues.
In reality, it is challenging to directly link a malfunction in
epigenetic inheritance to the occurrence of a particular phe-
notypes/disease at organism levels. Below, we outline several
examples in which alterations of factors involved in epigenetic
inheritance contribute to developmental defects and cancers.
Defects in genomic imprinting
A couple of well-studied examples linking defects in epi-
genetic inheritance to human diseases are the Prader-Willi
syndrome (PWS) and Angelman syndrome (AS). PWS and
AS are distinct human neurological disorders resulting from
defects in genomic imprinting of a gene cluster at 15q11q13
locus. While some genes are only expressed from the ma-
ternal allele, several genes including SNRPN and SnoRNA
are expressed paternally, with the maternal allele silenced
through DNA methylation. PWS is caused by the loss of the
expression of paternally expressed genes, whereas AS is
caused by loss of expression of maternally expressed genes.
It is estimated that 86% patients with PWS and 92% patients
with AS are caused by epimutations without changes at un-
derlying DNA sequence. Of note, about one third of these AS
patients show somatic mosaicism in which cells with im-
printing defects and normal cells co-exist (Horsthemke and
Buiting, 2008). These results indicate that a majority of PWS
and AS cases are caused by sporadic errors during the pro-
cess of establishment, and maintenance of this imprinting
locus. Future studies are needed to understand the molecular
basis for the generation of epimutations at this imprinted
gene cluster.
With the advancement of sequencing-based technologies,
more and more imprinted genes have been identified. Cur-
rently, it is estimated that over 220 genes are imprinted in
human genome (Horsthemke and Buiting, 2008). Moreover,
in addition to DNA methylation based on mechanism of
genomic imprinting, H3K27me3 alone can also imprint
genes during mouse early development (Inoue et al., 2017a;
Inoue et al., 2017b). These advancements will likely provide
additional insights into how alterations in imprinting con-
tribute to human diseases. For more information about
genomic imprinting, we direct readers to two recent reviews
on this topic (Peters, 2014; Monk et al., 2019).
A critical role for CAF-1 in maintaining chromatin states
and cell identity during development and tumorigenesis
CAF-1, the first identified histone chaperone involved in
deposition of new H3-H4 onto replicating DNA, plays an
important role in maintaining chromatin states from yeast to
human. An early study in Arabidopsis found that CAF-1 is
important to maintain cellular and functional organization of
both shoot apical meristem and the root apical meristem
(Kaya et al., 2001), which are responsible for postembryonic
development of plant architecture. Recently, it has been
shown that depletion of CAF-1 in mouse ES cells results in
an increase in 2C-like cells (Ishiuchi et al., 2015). Further-
more, depletion of CAF-1 in mouse embryonic fibroblast
increases in the reprograming efficiency of these cells into
iPSC, likely due to an increase in chromatin accessibility
(Cheloufi et al., 2015). Therefore, CAF-1 is important to
maintain chromatin states and cell identity likely in all cell
types during normal development.
Two recent studies also report that alterations in CAF-1
expression can promote tumorigenesis and drive tumor me-
tastasis. It is known for a long time that CHAF1B, a subunit
of CAF-1, is overexpressed in several solid tumors and acute
megakaryocytic leukemia (AMKL) (Polo et al., 2010; Sy-
karas et al., 2021). However, it was not known whether the
overexpression of CHAF1B has any role in tumorigenesis.
Using mouse models, it has been shown that CAF-1 is es-
sential for normal hematopoiesis. However, overexpression
2180
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
of CHAF1B interferes with the association of transcription
factors such as CEBPA involved in myeloid differentiation,
which in turn promotes leukemia. The effects of CHAF1B
overexpression on leukemia genesis are linked to the role of
CHAF1B in nucleosome assembly of new H3-H4 (Volk et
al., 2018). On newly replicating chromatin, transcription
factor binding sites are temporarily blocked (Ramachandran
and Henikoff, 2016). As parental H3-H4 can memorize their
positions along DNA following DNA replication (Escobar et
al., 2019), it is likely that the block of transcription factors is
caused by the deposition of new H3-H4 by CAF-1. There-
fore, overexpression of CHAF1B likely exacerbates the
blocking effects of CAF-1, thereby inhibiting myeloid dif-
ferentiation. In the future, it would be interesting to de-
termine whether CAF-1 overexpression in solid tumors also
plays a causal role in tumorigenesis.
While CAF-1 overexpression promotes leukemia, a recent
study indicates that reduced CAF-1 expression contributes to
tumor metastasis (Gomes et al., 2019). Tumor metastasis,
referring to cancer cells migrating from the primary organs
through the blood or lymph systems and forming tumors at
new organs, contributes to the largest fraction of cancer-
induced death (Fares et al., 2020). Genome wide analysis of
several tumors and their matched metastatic ones indicate
that epigenetic changes, but not genetic mutations, are likely
the dominant force in the development of tumor metastasis
(McDonald et al., 2017; Chatterjee et al., 2018). Using car-
cinoma models, it has been shown that metastatic signals
suppress the expression of CAF-1, leading to reduced density
of canonical histone H3.1/H3.2, which are assembled into
nucleosomes by CAF-1. This will trigger an increase in
HIRA mediated nucleosome assembly of H3.3 and the ac-
quisition of more aggressive and metastatic characteristics of
cancer. Depletion of HIRA suppresses the metastatic phe-
notypes (Gomes et al., 2019). Together, these studies high-
light the dynamic regulation of CAF-1-mediated nucleosome
assembly of H3.1/H3.2 and HIRA-mediated nucleosome
assembly of H3.3 in changes of cell identity to a more me-
tastatic one. In the future, it would be interesting to de-
termine to what extent other factors involved in epigenetic
inheritance discussed above may play in promoting cell fate
changes and thereby metastasis.
Summary and future directions
In the last several years, we have witnessed major advances
in the understanding of epigenetic inheritance. Specifically,
studies from various systems have established that repressive
histone modifications can be inherited, at least in part,
through the read-and-write mechanism. Because of the dy-
namic and reversible natures of histone modifications, it is
also clear that non-coding RNAs and DNA sequence specific
binding proteins are needed to recruit histone modifying
enzymes locally for the stable inheritance of a histone
modification. More importantly, we have also witnessed
major advances in our understanding of the recycling of
parental histones, which contain epigenetic modifications,
following DNA replication. Finally, we have also begun to
appreciate the importance of maintenance of chromatin
states and cell identity to prevent diseases including tumors.
These advances have laid a solid foundation for dissecting
molecular mechanisms of epigenetic inheritance during
normal development, and in tumorigenesis. However, many
questions still remain to be answered. How is parental his-
tone transfer/recycling regulated? Are there other factors
involved in parental histone transfer? Is there any co-
ordination between parental histone transfer and de novo
deposition of new H3-H4, and if there is, how do these two
pathways coordinate to promote nucleosome formation?
Considering the extensive cross talk between DNA methy-
lation and histone modifications, do protein machineries
involved in the heritance of DNA methylation also con-
tribute to the inheritance of histone modifications, or vice
versa? Do alterations in epigenetic inheritance contribute to
the establishment of alterative chromatin states that specify
disease evolution such as the transition of cancer cells to
metastatic cells? Future studies to address these and other
questions in epigenetic inheritance will advance our under-
standing of epigenetic inheritance and the contribution of
malfunction of this process to human disease.
Compliance and ethics The author(s) declare that they have no conflict
of interest.
Acknowledgements This work was supported by the National Natural
Science Foundation of China (31725015, 31830048 to Q.L. and 32000417
to W.D.), the Beijing Outstanding Young Scientist Program
(BJJWZYJH01201910001005 to Q.L.), the National Key Research and
Development Project of China (2019YFA0508903 to Q.L.), the China
Postdoctoral Science Foundation (2020M670487 to W. D.), the Chinese
Academy of Sciences (XDB 37010100 and QYZDY-SSW-SMC031 to B.Z.),
the K. C. Wong educational foundation (GJTD-2020-06 to B.Z.) and the
National Institutes of Health (R35 GM126910 to S.J. and R35 GM115018 to
Z.Z.). We apologize that we could not cite all references because of space
limitations.
References
Adkins, M.W., Howar, S.R., and Tyler, J.K. (2004). Chromatin disassembly
mediated by the histone chaperone Asf1 is essential for transcriptional
activation of the yeast PHO5 and PHO8 genes. Mol Cell 14, 657–666.
Adkins, M.W., and Tyler, J.K. (2004). The histone chaperone Asf1p
mediates global chromatin disassembly in vivo. J Biol Chem 279,
52069–52074.
Ahmad, K., and Henikoff, S. (2002a). Histone H3 variants specify modes
of chromatin assembly. Proc Natl Acad Sci USA 99, 16477–16484.
Ahmad, K., and Henikoff, S. (2002b). The histone variant H3.3 marks
active chromatin by replication-independent nucleosome assembly. Mol
Cell 9, 1191–1200.
Alabert, C., Barth, T.K., Reverón-Gómez, N., Sidoli, S., Schmidt, A.,
2181
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Jensen, O.N., Imhof, A., and Groth, A. (2015). Two distinct modes for
propagation of histone PTMs across the cell cycle. Genes Dev 29, 585–
590.
Allis, C.D., and Jenuwein, T. (2016). The molecular hallmarks of
epigenetic control. Nat Rev Genet 17, 487–500.
Allis, C.D., Jenuwein, T., and Reinberg, D. (2007). Epigenetics (Cold
Spring Harbor: Cold Spring Harbor Laboratory).
An, S., Yoon, J., Kim, H., Song, J.J., and Cho, U.S. (2017). Structure-based
nuclear import mechanism of histones H3 and H4 mediated by Kap123.
eLife 6, e30244.
Arand, J., Spieler, D., Karius, T., Branco, M.R., Meilinger, D., Meissner,
A., Jenuwein, T., Xu, G., Leonhardt, H., Wolf, V., et al. (2012). In vivo
control of CpG and non-CpG DNA methylation by DNA
methyltransferases. PLoS Genet 8, e1002750.
Arita, K., Isogai, S., Oda, T., Unoki, M., Sugita, K., Sekiyama, N., Kuwata,
K., Hamamoto, R., Tochio, H., Sato, M., et al. (2012). Recognition of
modification status on a histone H3 tail by linked histone reader
modules of the epigenetic regulator UHRF1. Proc Natl Acad Sci USA
109, 12950–12955.
Audergon, P.N.C.B., Catania, S., Kagansky, A., Tong, P., Shukla, M.,
Pidoux, A.L., and Allshire, R.C. (2015). Restricted epigenetic
inheritance of H3K9 methylation. Science 348, 132–135.
Augui, S., Nora, E.P., and Heard, E. (2011). Regulation of X-chromosome
inactivation by the X-inactivation centre. Nat Rev Genet 12, 429–442.
Avner, P., and Heard, E. (2001). X-chromosome inactivation: counting,
choice and initiation. Nat Rev Genet 2, 59–67.
Avvakumov, G.V., Walker, J.R., Xue, S., Li, Y., Duan, S., Bronner, C.,
Arrowsmith, C.H., and Dhe-Paganon, S. (2008). Structural basis for
recognition of hemi-methylated DNA by the SRA domain of human
UHRF1. Nature 455, 822–825.
Aygün, O., Mehta, S., and Grewal, S.I.S. (2013). HDAC-mediated
suppression of histone turnover promotes epigenetic stability of
heterochromatin. Nat Struct Mol Biol 20, 547–554.
Bae, H.J., Dubarry, M., Jeon, J., Soares, L.M., Dargemont, C., Kim, J.,
Geli, V., and Buratowski, S. (2020). The Set1 N-terminal domain and
Swd2 interact with RNA polymerase II CTD to recruit COMPASS. Nat
Commun 11, 2181.
Ballaré, C., Lange, M., Lapinaite, A., Martin, G.M., Morey, L., Pascual, G.,
Liefke, R., Simon, B., Shi, Y., Gozani, O., et al. (2012). Phf19 links
methylated Lys36 of histone H3 to regulation of Polycomb activity. Nat
Struct Mol Biol 19, 1257–1265.
Bao, K., Shan, C.M., Moresco, J., Yates Iii, J., and Jia, S. (2019). Anti-
silencing factor Epe1 associates with SAGA to regulate transcription
within heterochromatin. Genes Dev 33, 116–126.
Baubec, T., Colombo, D.F., Wirbelauer, C., Schmidt, J., Burger, L., Krebs,
A.R., Akalin, A., and Schübeler, D. (2015). Genomic profiling of DNA
methyltransferases reveals a role for DNMT3B in genic methylation.
Nature 520, 243–247.
Bayne, E.H., White, S.A., Kagansky, A., Bijos, D.A., Sanchez-Pulido, L.,
Hoe, K.L., Kim, D.U., Park, H.O., Ponting, C.P., Rappsilber, J., et al.
(2010). Stc1: a critical link between RNAi and chromatin modification
required for heterochromatin integrity. Cell 140, 666–677.
Bell, S.P., and Dutta, A. (2002). DNA replication in eukaryotic cells. Annu
Rev Biochem 71, 333–374.
Bell, S.P., and Stillman, B. (1992). ATP-dependent recognition of
eukaryotic origins of DNA replication by a multiprotein complex.
Nature 357, 128–134.
Bellelli, R., Belan, O., Pye, V.E., Clement, C., Maslen, S.L., Skehel, J.M.,
Cherepanov, P., Almouzni, G., and Boulton, S.J. (2018). POLE3-pole4
is a histone H3-H4 chaperone that maintains chromatin integrity during
DNA replication. Mol Cell 72, 112–126.e5.
Belotserkovskaya, R., and Reinberg, D. (2004). Facts about FACT and
transcript elongation through chromatin. Curr Opin Genet Dev 14, 139–
146.
Benayoun, B.A., Pollina, E.A., and Brunet, A. (2015). Epigenetic
regulation of ageing: linking environmental inputs to genomic
stability. Nat Rev Mol Cell Biol 16, 593–610.
Bergink, E.W., Kloosterboer, H.J., Gruber, M., and Ab, G. (1973).
Estrogen-induced phosphoprotein synthesis in roosters. Biochim
Biophys Acta (BBA)-Nucl Acids Protein Syn 294, 497–506.
Bestor, T., Laudano, A., Mattaliano, R., and Ingram, V. (1988). Cloning and
sequencing of a cDNA encoding DNA methyltransferase of mouse
cells. J Mol Biol 203, 971–983.
Bestor, T.H., and Ingram, V.M. (1983). Two DNA methyltransferases from
murine erythroleukemia cells: purification, sequence specificity, and
mode of interaction with DNA. Proc Natl Acad Sci USA 80, 5559–
5563.
Bird, A., Taggart, M., Frommer, M., Miller, O.J., and Macleod, D. (1985).
A fraction of the mouse genome that is derived from islands of
nonmethylated, CpG-rich DNA. Cell 40, 91–99.
Bostick, M., Kim, J.K., Esteve, P.O., Clark, A., Pradhan, S., and Jacobsen,
S.E. (2007). UHRF1 plays a role in maintaining DNA methylation in
mammalian cells. Science 317, 1760–1764.
Boulay, G., Rosnoblet, C., Guérardel, C., Angrand, P.O., and Leprince, D.
(2011). Functional characterization of human Polycomb-like 3 isoforms
identifies them as components of distinct EZH2 protein complexes.
Biochem J 434, 333–342.
Bourc’his, D., Xu, G.L., Lin, C.S., Bollman, B., and Bestor, T.H. (2001).
Dnmt3L and the establishment of maternal genomic imprints. Science
294, 2536–2539.
Brandeis, M., Frank, D., Keshet, I., Siegfried, Z., Mendelsohn, M., Nemes,
A., Temper, V., Razin, A., and Cedar, H. (1994). Spl elements protect a
CpG island from de novo methylation. Nature 371, 435–438.
Brien, G.L., Gambero, G., O′Connell, D.J., Jerman, E., Turner, S.A., Egan,
C.M., Dunne, E.J., Jurgens, M.C., Wynne, K., Piao, L., et al. (2012).
Polycomb PHF19 binds H3K36me3 and recruits PRC2 and
demethylase NO66 to embryonic stem cell genes during
differentiation. Nat Struct Mol Biol 19, 1273–1281.
Brinkman, A.B., Gu, H., Bartels, S.J.J., Zhang, Y., Matarese, F., Simmer, F.,
Marks, H., Bock, C., Gnirke, A., Meissner, A., et al. (2012). Sequential
ChIP-bisulfite sequencing enables direct genome-scale investigation of
chromatin and DNA methylation cross-talk. Genome Res 22, 1128–
1138.
Burgers, P.M.J., and Kunkel, T.A. (2017). Eukaryotic DNA replication
fork. Annu Rev Biochem 86, 417–438.
Burgess, R.J., Zhou, H., Han, J., and Zhang, Z. (2010). A role for Gcn5 in
replication-coupled nucleosome assembly. Mol Cell 37, 469–480.
Cao, R., Wang, H., He, J., Erdjument-Bromage, H., Tempst, P., and Zhang,
Y. (2008). Role of hPHF1 in H3K27 methylation and Hox gene
silencing. Mol Cell Biol 28, 1862–1872.
Casanova, M., Preissner, T., Cerase, A., Poot, R., Yamada, D., Li, X.,
Appanah, R., Bezstarosti, K., Demmers, J., Koseki, H., et al. (2011).
Polycomblike 2 facilitates the recruitment of PRC2 Polycomb group
complexes to the inactive X chromosome and to target loci in
embryonic stem cells. Development 138, 1471–1482.
Chang, Y., Sun, L., Kokura, K., Horton, J.R., Fukuda, M., Espejo, A.,
Izumi, V., Koomen, J.M., Bedford, M.T., Zhang, X., et al. (2011). MPP8
mediates the interactions between DNA methyltransferase Dnmt3a and
H3K9 methyltransferase GLP/G9a. Nat Commun 2, 533.
Charlton, J., Downing, T.L., Smith, Z.D., Gu, H., Clement, K., Pop, R.,
Akopian, V., Klages, S., Santos, D.P., Tsankov, A.M., et al. (2018).
Global delay in nascent strand DNA methylation. Nat Struct Mol Biol
25, 327–332.
Chatterjee, A., Rodger, E.J., and Eccles, M.R. (2018). Epigenetic drivers of
tumourigenesis and cancer metastasis. Semin Cancer Biol 51, 149–159.
Chedin, F., Lieber, M.R., and Hsieh, C.L. (2002). The DNA
methyltransferase-like protein DNMT3L stimulates de novo
methylation by Dnmt3a. Proc Natl Acad Sci USA 99, 16916–16921.
Cheloufi, S., Elling, U., Hopfgartner, B., Jung, Y.L., Murn, J., Ninova, M.,
Hubmann, M., Badeaux, A.I., Euong Ang, C., Tenen, D., et al. (2015).
The histone chaperone CAF-1 safeguards somatic cell identity. Nature
528, 218–224.
Chen, C.C., Carson, J.J., Feser, J., Tamburini, B., Zabaronick, S., Linger, J.,
and Tyler, J.K. (2008). Acetylated lysine 56 on histone H3 drives
2182
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
chromatin assembly after repair and signals for the completion of repair.
Cell 134, 231–243.
Chen, P., Dong, L., Hu, M., Wang, Y.Z., Xiao, X., Zhao, Z., Yan, J., Wang,
P.Y., Reinberg, D., Li, M., et al. (2018). Functions of FACT in breaking
the nucleosome and maintaining its integrity at the single-nucleosome
level. Mol Cell 71, 284–293.e4.
Chen, T., Tsujimoto, N., and Li, E. (2004). The PWWP domain of Dnmt3a
and Dnmt3b is required for directing DNA methylation to the major
satellite repeats at pericentric heterochromatin. Mol Cell Biol 24, 9048–
9058.
Chen, T., Ueda, Y., Xie, S., and Li, E. (2002). A novel Dnmt3a isoform
produced from an alternative promoter localizes to euchromatin and its
expression correlates with active de novo methylation. J Biol Chem 277,
38746–38754.
Chen, Z., Yin, Q., Inoue, A., Zhang, C., and Zhang, Y. (2019). Allelic
H3K27me3 to allelic DNA methylation switch maintains noncanonical
imprinting in extraembryonic cells. Sci Adv 5, eaay7246.
Cheng, J., Yang, H., Fang, J., Ma, L., Gong, R., Wang, P., Li, Z., and Xu, Y.
(2015). Molecular mechanism for USP7-mediated DNMT1 stabilization
by acetylation. Nat Commun 6, 7023.
Cheng, J., Yang, Y., Fang, J., Xiao, J., Zhu, T., Chen, F., Wang, P., Li, Z.,
Yang, H., and Xu, Y. (2013). Structural insight into coordinated
recognition of trimethylated histone H3 lysine 9 (H3K9me3) by the
plant homeodomain (PHD) and tandem tudor domain (TTD) of UHRF1
(ubiquitin-like, containing PHD and RING finger domains, 1) protein. J
Biol Chem 288, 1329–1339.
Cheng, L., Zhang, X., Wang, Y., Gan, H., Xu, X., Lv, X., Hua, X., Que, J.,
Ordog, T., and Zhang, Z. (2019). Chromatin assembly factor 1 (CAF-1)
facilitates the establishment of facultative heterochromatin during
pluripotency exit. Nucl Acids Res 47, 11114–11131.
Choe, K.N., and Moldovan, G.L. (2017). Forging ahead through darkness:
PCNA, still the principal conductor at the replication fork. Mol Cell 65,
380–392.
Chuang, L.S.H., Ian, H.I., Koh, T.W., Ng, H.H., Xu, G., and Li, B.F.L.
(1997). Human DNA-(Cytosine-5) methyltransferase-PCNA complex
as a target for p21
WAF1
. Science 277, 1996–2000.
Clément, C., and Almouzni, G. (2015). MCM2 binding to histones H3-H4
and ASF1 supports a tetramer-to-dimer model for histone inheritance at
the replication fork. Nat Struct Mol Biol 22, 587–589.
Coleman, R.T., and Struhl, G. (2017). Causal role for inheritance of
H3K27me3 in maintaining the OFF state of a Drosophila HOX gene.
Science 356.
Cooper, D.N., Taggart, M.H., and Bird, A.P. (1983). Unmethlated domains
in vertebrate DNA. Nucl Acids Res 11, 647–658.
Csankovszki, G., Nagy, A., and Jaenisch, R. (2001). Synergism of Xist
RNA, DNA methylation, and histone hypoacetylation in maintaining X
chromosome inactivation. J Cell Biol 153, 773–784.
D’Urso, A., and Brickner, J.H. (2014). Mechanisms of epigenetic memory.
Trends Genet 30, 230–236.
Dahl, J.A., Jung, I., Aanes, H., Greggains, G.D., Manaf, A., Lerdrup, M.,
Li, G., Kuan, S., Li, B., Lee, A.Y., et al. (2016). Broad histone
H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic
transition. Nature 537, 548–552.
Dekker, J., and Mirny, L. (2016). The 3D genome as moderator of
chromosomal communication. Cell 164, 1110–1121.
Dennis, K., Fan, T., Geiman, T., Yan, Q., and Muegge, K. (2001). Lsh, a
member of the SNF2 family, is required for genome-wide methylation.
Genes Dev 15, 2940–2944.
Dhayalan, A., Rajavelu, A., Rathert, P., Tamas, R., Jurkowska, R.Z.,
Ragozin, S., and Jeltsch, A. (2010). The Dnmt3a PWWP domain reads
histone 3 lysine 36 trimethylation and guides DNA methylation. J Biol
Chem 285, 26114–26120.
Disteche, C.M., and Berletch, J.B. (2015). X-chromosome inactivation and
escape. J Genet 94, 591–599.
Donham, D.C. II, Scorgie, J.K., and Churchill, M.E.A. (2011). The activity
of the histone chaperone yeast Asf1 in the assembly and disassembly of
histone H3/H4-DNA complexes. Nucl Acids Res 39, 5449–5458.
Donovan, S., Harwood, J., Drury, L.S., and Diffley, J.F.X. (1997). Cdc6p-
dependent loading of Mcm proteins onto pre-replicative chromatin in
budding yeast. Proc Natl Acad Sci USA 94, 5611–5616.
Drané, P., Ouararhni, K., Depaux, A., Shuaib, M., and Hamiche, A. (2010).
The death-associated protein DAXX is a novel histone chaperone
involved in the replication-independent deposition of H3.3. Genes Dev
24, 1253–1265.
Du, W., Dong, Q., Zhang, Z., Liu, B., Zhou, T., Xu, R.M., Wang, H., Zhu,
B., and Li, Y. (2019). Stella protein facilitates DNA demethylation by
disrupting the chromatin association of the RING finger-type E3
ubiquitin ligase UHRF1. J Biol Chem 294, 8907–8917.
Du, Z., Song, J., Wang, Y., Zhao, Y., Guda, K., Yang, S., Kao, H.Y., Xu, Y.,
Willis, J., Markowitz, S.D., et al. (2010). DNMT1 stability is regulated
by proteins coordinating deubiquitination and acetylation-driven
ubiquitination. Sci Signal 3, ra80.
Dukatz, M., Holzer, K., Choudalakis, M., Emperle, M., Lungu, C.,
Bashtrykov, P., and Jeltsch, A. (2019). H3K36me2/3 binding and DNA
binding of the DNA methyltransferase DNMT3A PWWP domain both
contribute to its chromatin interaction. J Mol Biol 431, 5063–5074.
Duymich, C.E., Charlet, J., Yang, X., Jones, P.A., and Liang, G. (2016).
DNMT3B isoforms without catalytic activity stimulate gene body
methylation as accessory proteins in somatic cells. Nat Commun 7,
11453.
Eden, A., Gaudet, F., Waghmare, A., and Jaenisch, R. (2003).
Chromosomal instability and tumors promoted by DNA
hypomethylation. Science 300, 455.
Edmunds, J.W., Mahadevan, L.C., and Clayton, A.L. (2008). Dynamic
histone H3 methylation during gene induction: HYPB/Setd2 mediates
all H3K36 trimethylation. EMBO J 27, 406–420.
Egger, G., Jeong, S., Escobar, S.G., Cortez, C.C., Li, T.W.H., Saito, Y.,
Yoo, C.B., Jones, P.A., and Liang, G. (2006). Identification of DNMT1
(DNA methyltransferase 1) hypomorphs in somatic knockouts suggests
an essential role for DNMT1 in cell survival. Proc Natl Acad Sci USA
103, 14080–14085.
Elsässer, S.J., Huang, H., Lewis, P.W., Chin, J.W., Allis, C.D., and Patel, D.
J. (2012). DAXX envelops a histone H3.3-H4 dimer for H3.3-specific
recognition. Nature 491, 560–565.
English, C.M., Adkins, M.W., Carson, J.J., Churchill, M.E.A., and Tyler, J.
K. (2006). Structural basis for the histone chaperone activity of Asf1.
Cell 127, 495–508.
English, C.M., Maluf, N.K., Tripet, B., Churchill, M.E.A., and Tyler, J.K.
(2005). ASF1 binds to a heterodimer of histones H3 and H4: a two-step
mechanism for the assembly of the H3-H4 heterotetramer on DNA.
Biochemistry 44, 13673–13682.
Epsztejn-Litman, S., Feldman, N., Abu-Remaileh, M., Shufaro, Y., Gerson,
A., Ueda, J., Deplus, R., Fuks, F., Shinkai, Y., Cedar, H., et al. (2008).
De novo DNA methylation promoted by G9a prevents reprogramming
of embryonically silenced genes. Nat Struct Mol Biol 15, 1176–1183.
Escobar, T.M., Oksuz, O., Saldaña-Meyer, R., Descostes, N., Bonasio, R.,
and Reinberg, D. (2019). Active and repressed chromatin domains
exhibit distinct nucleosome segregation during DNA replication. Cell
179, 953–963.e11.
Estève, P.O., Chang, Y., Samaranayake, M., Upadhyay, A.K., Horton, J.R.,
Feehery, G.R., Cheng, X., and Pradhan, S. (2011). A methylation and
phosphorylation switch between an adjacent lysine and serine
determines human DNMT1 stability. Nat Struct Mol Biol 18, 42–48.
Estève, P.O., Chin, H.G., Benner, J., Feehery, G.R., Samaranayake, M.,
Horwitz, G.A., Jacobsen, S.E., and Pradhan, S. (2009). Regulation of
DNMT1 stability through SET7-mediated lysine methylation in
mammalian cells. Proc Natl Acad Sci USA 106, 5076–5081.
Estève, P.O., Chin, H.G., Smallwood, A., Feehery, G.R., Gangisetty, O.,
Karpf, A.R., Carey, M.F., and Pradhan, S. (2006). Direct interaction
between DNMT1 and G9a coordinates DNA and histone methylation
during replication. Genes Dev 20, 3089–3103.
Evrin, C., Maman, J.D., Diamante, A., Pellegrini, L., and Labib, K. (2018).
Histone H2A-H2B binding by Pol α in the eukaryotic replisome
contributes to the maintenance of repressive chromatin. EMBO J 37.
2183
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Fang, J., Cheng, J., Wang, J., Zhang, Q., Liu, M., Gong, R., Wang, P.,
Zhang, X., Feng, Y., Lan, W., et al. (2016). Hemi-methylated DNA
opens a closed conformation of UHRF1 to facilitate its histone
recognition. Nat Commun 7, 11197.
Fares, J., Fares, M.Y., Khachfe, H.H., Salhab, H.A., and Fares, Y. (2020).
Molecular principles of metastasis: a hallmark of cancer revisited. Sig
Transduct Target Ther 5, 28.
Fazly, A., Li, Q., Hu, Q., Mer, G., Horazdovsky, B., and Zhang, Z. (2012).
Histone chaperone Rtt106 promotes nucleosome formation using (H3-
H4)
2
tetramers. J Biol Chem 287, 10753–10760.
Feinberg, A.P., and Vogelstein, B. (1983). Hypomethylation distinguishes
genes of some human cancers from their normal counterparts. Nature
301, 89–92.
Fellinger, K., Rothbauer, U., Felle, M., Längst, G., and Leonhardt, H.
(2009). Dimerization of DNA methyltransferase 1 is mediated by its
regulatory domain. J Cell Biochem 106, 521–528.
Ferguson-Smith, A.C., and Bourc’his, D. (2018). The discovery and
importance of genomic imprinting. eLife 7, e42368.
Ferry, L., Fournier, A., Tsusaka, T., Adelmant, G., Shimazu, T., Matano, S.,
Kirsh, O., Amouroux, R., Dohmae, N., Suzuki, T., et al. (2017).
Methylation of DNA ligase 1 by G9a/GLP recruits UHRF1 to
replicating DNA and regulates DNA methylation. Mol Cell 67, 550–
565.e5.
Foltman, M., Evrin, C., De Piccoli, G., Jones, R.C., Edmondson, R.D.,
Katou, Y., Nakato, R., Shirahige, K., and Labib, K. (2013). Eukaryotic
replisome components cooperate to process histones during
chromosome replication. Cell Rep 3, 892–904.
Formosa, T., and Winston, F. (2020). The role of FACT in managing
chromatin: disruption, assembly, or repair? Nucl Acids Res 48, 11929–
11941.
Foster, B.M., Stolz, P., Mulholland, C.B., Montoya, A., Kramer, H.,
Bultmann, S., and Bartke, T. (2018). Critical role of the UBL domain in
stimulating the E3 ubiquitin ligase activity of uhrf1 toward chromatin.
Mol Cell 72, 739–752.e9.
Fraga, M.F., Ballestar, E., Villar-Garea, A., Boix-Chornet, M., Espada, J.,
Schotta, G., Bonaldi, T., Haydon, C., Ropero, S., Petrie, K., et al.
(2005). Loss of acetylation at Lys16 and trimethylation at Lys20 of
histone H4 is a common hallmark of human cancer. Nat Genet 37, 391–
400.
Fu, Y.V., Yardimci, H., Long, D.T., Guainazzi, A., Bermudez, V.P.,
Hurwitz, J., van Oijen, A., Schärer, O.D., and Walter, J.C. (2011).
Selective bypass of a lagging strand roadblock by the eukaryotic
replicative DNA helicase. Cell 146, 931–941.
Fuks, F., Hurd, P.J., Deplus, R., and Kouzarides, T. (2003). The DNA
methyltransferases associate with HP1 and the SUV39H1 histone
methyltransferase. Nucl Acids Res 31, 2305–2312.
Galupa, R., and Heard, E. (2018). X-Chromosome inactivation: a
crossroads between chromosome architecture and gene regulation.
Annu Rev Genet 52, 535–566.
Gambus, A., Jones, R.C., Sanchez-Diaz, A., Kanemaki, M., van Deursen,
F., Edmondson, R.D., and Labib, K. (2006). GINS maintains association
of Cdc45 with MCM in replisome progression complexes at eukaryotic
DNA replication forks. Nat Cell Biol 8, 358–366.
Gan, H., Serra-Cardona, A., Hua, X., Zhou, H., Labib, K., Yu, C., and
Zhang, Z. (2018). The Mcm2-Ctf4-Polα axis facilitates parental histone
H3-H4 transfer to lagging strands. Mol Cell 72, 140–151.e3.
Gao, Y., Gan, H., Lou, Z., and Zhang, Z. (2018). Asf1a resolves bivalent
chromatin domains for the induction of lineage-specific genes during
mouse embryonic stem cell differentiation. Proc Natl Acad Sci USA
115.
Gaydos, L.J., Rechtsteiner, A., Egelhofer, T.A., Carroll, C.R., and Strome,
S. (2012). Antagonism between MES-4 and Polycomb repressive
complex 2 promotes appropriate gene expression in C. elegans germ
cells. Cell Rep 2, 1169–1177.
Ge, Y.Z., Pu, M.T., Gowher, H., Wu, H.P., Ding, J.P., Jeltsch, A., and Xu,
G.L. (2004). Chromatin targeting of de novo DNA methyltransferases
by the PWWP domain. J Biol Chem 279, 25447–25454.
Gerace, E.L., Halic, M., and Moazed, D. (2010). The methyltransferase
activity of Clr4Suv39h triggers RNAi independently of histone H3K9
methylation. Mol Cell 39, 360–372.
Ginno, P.A., Lott, P.L., Christensen, H.C., Korf, I., and Chédin, F. (2012).
R-loop formation is a distinctive characteristic of unmethylated human
CpG island promoters. Mol Cell 45, 814–825.
Goldberg, A.D., Banaszynski, L.A., Noh, K.M., Lewis, P.W., Elsaesser, S.
J., Stadler, S., Dewell, S., Law, M., Guo, X., Li, X., et al. (2010).
Distinct factors control histone variant H3.3 localization at specific
genomic regions. Cell 140, 678–691.
Gomes, A.P., Ilter, D., Low, V., Rosenzweig, A., Shen, Z.J., Schild, T.,
Rivas, M.A., Er, E.E., McNally, D.R., Mutvei, A.P., et al. (2019).
Dynamic incorporation of histone H3 variants into chromatin is
essential for acquisition of aggressive traits and metastatic
colonization. Cancer Cell 36, 402–417.e13.
González-Magaña, A., de Opakua, A.I., Merino, N., Monteiro, H., Diercks,
T., Murciano-Calles, J., Luque, I., Bernadó, P., Cordeiro, T.N., Biasio,
A.D., et al. (2019). Double monoubiquitination modifies the molecular
recognition properties of p15
PAF
promoting binding to the reader module
of Dnmt1. ACS Chem Biol acschembio.9b00679.
Gopalakrishnan, S., Van Emburgh, B.O., Shan, J., Su, Z., Fields, C.R.,
Vieweg, J., Hamazaki, T., Schwartz, P.H., Terada, N., and Robertson, K.
D. (2009). A novel DNMT3B splice variant expressed in tumor and
pluripotent cells modulates genomic DNA methylation patterns and
displays altered DNA binding. Mol Cancer Res 7, 1622–1634.
Gottschling, D.E., Aparicio, O.M., Billington, B.L., and Zakian, V.A.
(1990). Position effect at S. cerevisiae telomeres: reversible repression
of Pol II transcription. Cell 63, 751–762.
Greer, E.L., Maures, T.J., Ucar, D., Hauswirth, A.G., Mancini, E., Lim, J.P.,
Benayoun, B.A., Shi, Y., and Brunet, A. (2011). Transgenerational
epigenetic inheritance of longevity in Caenorhabditis elegans. Nature
479, 365–371.
Grewal, S.I.S., and Jia, S. (2007). Heterochromatin revisited. Nat Rev
Genet 8, 35–46.
Grewal, S.I.S., and Klar, A.J.S. (1996). Chromosomal inheritance of
epigenetic states in fission yeast during mitosis and meiosis. Cell 86,
95–101.
Grewal, S.I.S., and Moazed, D. (2003). Heterochromatin and epigenetic
control of gene expression. Science 301, 798–802.
Groth, A., Corpet, A., Cook, A.J.L., Roche, D., Bartek, J., Lukas, J., and
Almouzni, G.. (2007). Regulation of replication fork progression
through histone supply and demand. Science 318, 1928–1931.
Gruszka, D.T., Xie, S., Kimura, H., and Yardimci, H. (2020). Single-
molecule imaging reveals control of parental histone recycling by free
histones during DNA replication. Sci Adv 6, eabc0330.
Guo, X., Wang, L., Li, J., Ding, Z., Xiao, J., Yin, X., He, S., Shi, P., Dong,
L., Li, G., et al. (2015). Structural insight into autoinhibition and histone
H3-induced activation of DNMT3A. Nature 517, 640–644.
Hall, I.M., Shankaranarayana, G.D., Noma, K.I., Ayoub, N., Cohen, A., and
Grewal, S.I.S. (2002). Establishment and maintenance of a
heterochromatin domain. Science 297, 2232–2237.
Han, J., Zhang, H., Zhang, H., Wang, Z., Zhou, H., and Zhang, Z. (2013). A
CUL4 E3 ubiquitin ligase regulates histone hand-off during nucleosome
assembly. Cell 155, 817–829.
Han, J., Zhou, H., Horazdovsky, B., Zhang, K., Xu, R.M., and Zhang, Z.
(2007a). Rtt109 acetylates histone H3 lysine 56 and functions in DNA
replication. Science 315, 653–655.
Han, J., Zhou, H., Li, Z., Xu, R.M., and Zhang, Z. (2007b). The Rtt109-
Vps75 histone acetyltransferase complex acetylates non-nucleosomal
histone H3. J Biol Chem 282, 14158–14164.
Han, M., Li, J., Cao, Y., Huang, Y., Li, W., Zhu, H., Zhao, Q., Han, J.D.J.,
Wu, Q., Li, J., et al. (2020). A role for LSH in facilitating DNA
methylation by DNMT1 through enhancing UHRF1 chromatin
association. Nucl Acids Res 48, 12116–12134.
Hashimoto, H., Horton, J.R., Zhang, X., Bostick, M., Jacobsen, S.E., and
Cheng, X. (2008). The SRA domain of UHRF1 flips 5-methylcytosine
out of the DNA helix. Nature 455, 826–829.
2184
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Hashimoto, H., Horton, J.R., Zhang, X., and Cheng, X. (2009). UHRF1, a
modular multi-domain protein, regulates replication-coupled crosstalk
between DNA methylation and histone modifications. Epigenetics 4, 8–
14.
He, C., Liu, N., Xie, D., Liu, Y., Xiao, Y., and Li, F. (2019). Structural basis
for histone H3K4me3 recognition by the N-terminal domain of the PHD
finger protein Spp1. Biochem J 476, 1957–1973.
He, H., Li, Y., Dong, Q., Chang, A.Y., Gao, F., Chi, Z., Su, M., Zhang, F.,
Ban, H., Martienssen, R., et al. (2017). Coordinated regulation of
heterochromatin inheritance by Dpb3-Dpb4 complex. Proc Natl Acad
Sci USA 114, 12524–12529.
Heard, E., and Martienssen, R.A. (2014). Transgenerational epigenetic
inheritance: myths and mechanisms. Cell 157, 95–109.
Heyn, P., Logan, C.V., Fluteau, A., Challis, R.C., Auchynnikava, T., Martin,
C.A., Marsh, J.A., Taglini, F., Kilanowski, F., Parry, D.A., et al. (2019).
Gain-of-function DNMT3A mutations cause microcephalic dwarfism
and hypermethylation of Polycomb-regulated regions. Nat Genet 51,
96–105.
Højfeldt, J.W., Hedehus, L., Laugesen, A., Tatar, T., Wiehle, L., and Helin,
K. (2019). Non-core subunits of the PRC2 complex are collectively
required for its target-site specificity. Mol Cell 76, 423–436.e3.
Holoch, D., and Margueron, R. (2017). Mechanisms regulating prc2
recruitment and enzymatic activity. Trends Biochem Sci 42, 531–542.
Hon, G.C., Rajagopal, N., Shen, Y., McCleary, D.F., Yue, F., Dang, M.D.,
and Ren, B. (2013). Epigenetic memory at embryonic enhancers
identified in DNA methylation maps from adult mouse tissues. Nat
Genet 45, 1198–1206.
Hörmanseder, E., Simeone, A., Allen, G.E., Bradshaw, C.R., Figlmüller,
M., Gurdon, J., and Jullien, J. (2017). H3K4 methylation-dependent
memory of somatic cell identity inhibits reprogramming and
development of nuclear transfer embryos. Cell Stem Cell 21, 135–
143.e6.
Horsthemke, B. (2018). A critical view on transgenerational epigenetic
inheritance in humans. Nat Commun 9, 2973.
Horsthemke, B., and Buiting, K. (2008). Genomic imprinting and im-
printing defects in humans. Adv Genet 61, 225–246.
Hu, Y., Tareen, A., Sheu, Y.J., Ireland, W.T., Speck, C., Li, H., Joshua-Tor,
L., Kinney, J.B., and Stillman, B. (2020). Evolution of DNA replication
origin specification and gene silencing mechanisms. Nat Commun 11,
5175.
Huang, C., and Zhu, B. (2018). Roles of H3K36-specific histone
methyltransferases in transcription: antagonizing silencing and
safeguarding transcription fidelity. Biophys Rep 4, 170–177.
Huang, H., Strømme, C.B., Saredi, G., Hödl, M., Strandsby, A., González-
Aguilera, C., Chen, S., Groth, A., and Patel, D.J. (2015). A unique
binding mode enables MCM2 to chaperone histones H3-H4 at
replication forks. Nat Struct Mol Biol 22, 618–626.
Huang, S., Zhou, H., Katzmann, D., Hochstrasser, M., Atanasova, E., and
Zhang, Z. (2005). Rtt106p is a histone chaperone involved in
heterochromatin-mediated silencing. Proc Natl Acad Sci USA 102,
13410–13415.
Huang, S., Zhou, H., Tarara, J., and Zhang, Z. (2007). A novel role for
histone chaperones CAF-1 and Rtt106p in heterochromatin silencing.
EMBO J 26, 2274–2283.
Hunkapiller, J., Shen, Y., Diaz, A., Cagney, G., McCleary, D., Ramalho-
Santos, M., Krogan, N., Ren, B., Song, J.S., and Reiter, J.F. (2012).
Polycomb-like 3 promotes Polycomb repressive complex 2 binding to
CpG islands and embryonic stem cell self-renewal. PLoS Genet 8,
e1002576.
Ilves, I., Petojevic, T., Pesavento, J.J., and Botchan, M.R. (2010).
Activation of the MCM2-7 helicase by association with Cdc45 and
GINS proteins. Mol Cell 37, 247–258.
Inoue, A., Jiang, L., Lu, F., Suzuki, T., and Zhang, Y. (2017a). Maternal
H3K27me3 controls DNA methylation-independent imprinting. Nature
547, 419–424.
Inoue, A., Jiang, L., Lu, F., and Zhang, Y. (2017b). Genomic imprinting of
Xist by maternal H3K27me3. Genes Dev 31, 1927–1932.
Ishimi, Y., Komamura-Kohno, Y., Arai, K., and Masai, H. (2001).
Biochemical activities associated with mouse Mcm2 protein. J Biol
Chem 276, 42744–42752.
Ishimi, Y., Komamura, Y., You, Z., and Kimura, H. (1998). Biochemical
function of mouse minichromosome maintenance 2 protein. J Biol
Chem 273, 8369–8375.
Ishiuchi, T., Enriquez-Gasca, R., Mizutani, E., Bošković, A., Ziegler-
Birling, C., Rodriguez-Terrones, D., Wakayama, T., Vaquerizas, J.M.,
and Torres-Padilla, M.E. (2015). Early embryonic-like cells are induced
by downregulating replication-dependent chromatin assembly. Nat
Struct Mol Biol 22, 662–671.
Ishiyama, S., Nishiyama, A., Saeki, Y., Moritsugu, K., Morimoto, D.,
Yamaguchi, L., Arai, N., Matsumura, R., Kawakami, T., Mishima, Y., et
al. (2017). Structure of the DNMT1 reader module complexed with a
unique two-mono-ubiquitin mark on histone H3 reveals the basis for
DNA methylation maintenance. Mol Cell 68, 350–360.e7.
Jackson, J.P., Lindroth, A.M., Cao, X., and Jacobsen, S.E. (2002). Control
of CpNpG DNA methylation by the KRYPTONITE histone H3
methyltransferase. Nature 416, 556–560.
Jackson, M., Krassowska, A., Gilbert, N., Chevassut, T., Forrester, L.,
Ansell, J., and Ramsahoye, B. (2004). Severe global DNA
hypomethylation blocks differentiation and induces histone
hyperacetylation in embryonic stem cells. Mol Cell Biol 24, 8862–
8871.
Jégu, T., Aeby, E., and Lee, J.T. (2017). The X chromosome in space. Nat
Rev Genet 18, 377–389.
Jeltsch, A., and Jurkowska, R.Z. (2014). New concepts in DNA
methylation. Trends Biochem Sci 39, 310–318.
Jia, D., Jurkowska, R.Z., Zhang, X., Jeltsch, A., and Cheng, X. (2007).
Structure of Dnmt3a bound to Dnmt3L suggests a model for de novo
DNA methylation. Nature 449, 248–251.
Jia, S., Noma, K., and Grewal, S.I.S. (2004). RNAi-independent
heterochromatin nucleation by the stress-activated ATF/CREB family
proteins. Science 304, 1971–1976.
Jones, P.A., Issa, J.P.J., and Baylin, S. (2016). Targeting the cancer
epigenome for therapy. Nat Rev Genet 17, 630–641.
Jones, P.L., Veenstra, G.J.C., Wade, P.A., Vermaak, D., Kass, S.U.,
Landsberger, N., Strouboulis, J., and Wolffe, A.P. (1998). Methylated
DNA and MeCP2 recruit histone deacetylase to repress transcription.
Nat Genet 19, 187–191.
Jurkowska, R.Z., Rajavelu, A., Anspach, N., Urbanke, C., Jankevicius, G.,
Ragozin, S., Nellen, W., and Jeltsch, A. (2011). Oligomerization and
binding of the Dnmt3a DNA methyltransferase to parallel DNA
molecules. J Biol Chem 286, 24200–24207.
Kang, B., Pu, M., Hu, G., Wen, W., Dong, Z., Zhao, K., Stillman, B., and
Zhang, Z. (2011). Phosphorylation of H4 Ser 47 promotes HIRA-
mediated nucleosome assembly. Genes Dev 25, 1359–1364.
Karg, E., Smets, M., Ryan, J., Forné, I., Qin, W., Mulholland, C.B.,
Kalideris, G., Imhof, A., Bultmann, S., and Leonhardt, H. (2017).
Ubiquitome analysis reveals PCNA-associated factor 15 (PAF15) as a
specific ubiquitination target of UHRF1 in embryonic stem cells. J Mol
Biol 429, 3814–3824.
Kaufman, P.D., Kobayashi, R., and Stillman, B. (1997). Ultraviolet
radiation sensitivity and reduction of telomeric silencing in
Saccharomyces cerevisiae cells lacking chromatin assembly factor-I.
Genes Dev 11, 345–357.
Kaya, H., Shibahara, K., Taoka, K., Iwabuchi, M., Stillman, B., and Araki,
T. (2001). FASCIATA genes for chromatin assembly factor-1 in
arabidopsis maintain the cellular organization of apical meristems.
Cell 104, 131–142.
Kim, H.S., Choi, E.S., Shin, J.A., Jang, Y.K., and Park, S.D. (2004).
Regulation of Swi6/HP1-dependent heterochromatin assembly by
cooperation of components of the mitogen-activated protein kinase
pathway and a histone deacetylase Clr6. J Biol Chem 279, 42850–
42859.
Ku, M., Koche, R.P., Rheinbay, E., Mendenhall, E.M., Endoh, M.,
Mikkelsen, T.S., Presser, A., Nusbaum, C., Xie, X., Chi, A.S., et al.
2185
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
(2008). Genomewide analysis of PRC1 and PRC2 occupancy identifies
two classes of bivalent domains. PLoS Genet 4, e1000242.
Kuo, A.J., Cheung, P., Chen, K., Zee, B.M., Kioi, M., Lauring, J., Xi, Y.,
Park, B.H., Shi, X., Garcia, B.A., et al. (2011). NSD2 links
dimethylation of histone H3 at lysine 36 to oncogenic programming.
Mol Cell 44, 609–620.
Kurat, C.F., Yeeles, J.T.P., Patel, H., Early, A., and Diffley, J.F.X. (2017).
Chromatin controls DNA replication origin selection, lagging-strand
synthesis, and replication fork rates. Mol Cell 65, 117–130.
La Salle, S., and Trasler, J.M. (2006). Dynamic expression of DNMT3a and
DNMT3b isoforms during male germ cell development in the mouse.
Dev Biol 296, 71–82.
Laprell, F., Finkl, K., and Müller, J. (2017). Propagation of Polycomb-
repressed chromatin requires sequence-specific recruitment to DNA.
Science 356, 85–88.
Lehnertz, B., Ueda, Y., Derijck, A.A.H.A., Braunschweig, U., Perez-
Burgos, L., Kubicek, S., Chen, T., Li, E., Jenuwein, T., and Peters, A.H.
F.M. (2003). Suv39h-mediated histone H3 lysine 9 methylation directs
DNA methylation to major satellite repeats at pericentric
heterochromatin. Curr Biol 13, 1192–1200.
LeRoy, G., Orphanides, G., Lane, W.S., and Reinberg, D. (1998).
Requirement of RSF and FACT for transcription of chromatin
templates in vitro. Science 282, 1900–1904.
Lewis, P.W., Elsaesser, S.J., Noh, K.M., Stadler, S.C., and Allis, C.D.
(2010). Daxx is an H3.3-specific histone chaperone and cooperates with
ATRX in replication-independent chromatin assembly at telomeres.
Proc Natl Acad Sci USA 107, 14075–14080.
Li, B.Z., Huang, Z., Cui, Q.Y., Song, X.H., Du, L., Jeltsch, A., Chen, P., Li,
G., Li, E., and Xu, G.L. (2011). Histone tails regulate DNA methylation
by allosterically activating de novo methyltransferase. Cell Res 21,
1172–1181.
Li, H., Liefke, R., Jiang, J., Kurland, J.V., Tian, W., Deng, P., Zhang, W.,
He, Q., Patel, D.J., Bulyk, M.L., et al. (2017). Polycomb-like proteins
link the PRC2 complex to CpG islands. Nature 549, 287–291.
Li, H., Rauch, T., Chen, Z.X., Szabó, P.E., Riggs, A.D., and Pfeifer, G.P.
(2006). The histone methyltransferase SETDB1 and the DNA
methyltransferase DNMT3A interact directly and localize to
promoters silenced in cancer cells. J Biol Chem 281, 19489–19500.
Li, M., Gan, J., Sun, Y., Xu, Z., Yang, J., Sun, Y., and Li, C. (2020a).
Architectural proteins for the formation and maintenance of the 3D
genome. Sci China Life Sci 63, 795–810.
Li, Q., Burgess, R., and Zhang, Z. (2013). All roads lead to chromatin:
multiple pathways for histone deposition. Biochim Biophys Acta
(BBA)-Gene Regulatory Mech 1819, 238–246.
Li, Q., Zhou, H., Wurtele, H., Davies, B., Horazdovsky, B., Verreault, A.,
and Zhang, Z. (2008). Acetylation of histone H3 lysine 56 regulates
replication-coupled nucleosome assembly. Cell 134, 244–255.
Li, T., Wang, L., Du, Y., Xie, S., Yang, X., Lian, F., Zhou, Z., and Qian, C.
(2018a). Structural and mechanistic insights into UHRF1-mediated
DNMT1 activation in the maintenance DNA methylation. Nucl Acids
Res 46, 3218–3231.
Li, Y., Zhang, Z., Chen, J., Liu, W., Lai, W., Liu, B., Li, X., Liu, L., Xu, S.,
Dong, Q., et al. (2018b). Stella safeguards the oocyte methylome by
preventing de novo methylation mediated by DNMT1. Nature 564,
136–140.
Li, Z., Hua, X., Serra-Cardona, A., Xu, X., Gan, S., Zhou, H., Yang, W.S.,
Chen, C.L., Xu, R.M., and Zhang, Z. (2020b). DNA polymerase α
interacts with H3-H4 and facilitates the transfer of parental histones to
lagging strands. Sci Adv 6, eabb5820.
Liang, G., Chan, M.F., Tomigahara, Y., Tsai, Y.C., Gonzales, F.A., Li, E.,
Laird, P.W., and Jones, P.A. (2002). Cooperativity between DNA
methyltransferases in the maintenance methylation of repetitive
elements. Mol Cell Biol 22, 480–491.
Liu, C.P., Xiong, C., Wang, M., Yu, Z., Yang, N., Chen, P., Zhang, Z., Li,
G., and Xu, R.M. (2012a). Structure of the variant histone H3.3-H4
heterodimer in complex with its chaperone DAXX. Nat Struct Mol Biol
19, 1287–1292.
Liu, N., Zhang, Z., Wu, H., Jiang, Y., Meng, L., Xiong, J., Zhao, Z., Zhou,
X., Li, J., Li, H., et al. (2015). Recognition of H3K9 methylation by
GLP is required for efficient establishment of H3K9 methylation, rapid
target gene repression, and mouse viability. Genes Dev 29, 379–393.
Liu, S., Xu, Z., Leng, H., Zheng, P., Yang, J., Chen, K., Feng, J., and Li, Q.
(2017). RPA binds histone H3-H4 and functions in DNA replication–
coupled nucleosome assembly. Science 355, 415–420.
Liu, W.H., Roemer, S.C., Port, A.M., and Churchill, M.E.A. (2012b). CAF-
1-induced oligomerization of histones H3/H4 and mutually exclusive
interactions with Asf1 guide H3/H4 transitions among histone
chaperones and DNA. Nucl Acids Res 40, 11229–11239.
Liu, W.H., Roemer, S.C., Zhou, Y., Shen, Z.J., Dennehey, B.K., Balsbaugh,
J.L., Liddle, J.C., Nemkov, T., Ahn, N.G., Hansen, K.C., et al. (2016).
The Cac1 subunit of histone chaperone CAF-1 organizes CAF-1-H3/H4
architecture and tetramerizes histones. eLife 5, e18023.
Liu, Y., Zhou, K., Zhang, N., Wei, H., Tan, Y.Z., Zhang, Z., Carragher, B.,
Potter, C.S., D′Arcy, S., and Luger, K. (2020). FACT caught in the act of
manipulating the nucleosome. Nature 577, 426–431.
Long, H., Zhang, L., Lv, M., Wen, Z., Zhang, W., Chen, X., Zhang, P., Li,
T., Chang, L., Jin, C., et al. (2020). H2A.Z facilitates licensing and
activation of early replication origins. Nature 577, 576–581.
Loyola, A., and Almouzni, G. (2007). Marking histone H3 variants: how,
when and why? Trends Biochem Sci 32, 425–433.
Loyola, A., Bonaldi, T., Roche, D., Imhof, A., and Almouzni, G. (2006).
PTMs on H3 variants before chromatin assembly potentiate their final
epigenetic state. Mol Cell 24, 309–316.
Lu, C., Jain, S.U., Hoelper, D., Bechet, D., Molden, R.C., Ran, L., Murphy,
D., Venneti, S., Hameed, M., Pawel, B.R., et al. (2016). Histone H3K36
mutations promote sarcomagenesis through altered histone methylation
landscape. Science 352, 844–849.
Lucchini, R., Wellinger, R.E., and Sogo, J.M. (2001). Nucleosome
positioning at the replication fork. EMBO J 20, 7294–7302.
MacAlpine, D.M. (2021). Stochastic initiation of DNA replication across
the human genome. Mol Cell 81, 2873–2874.
MacAlpine, D.M., and Almouzni, G. (2013). Chromatin and DNA
replication. Cold Spring Harbor Perspectives Biol 5, a010207.
Macleod, D., Charlton, J., Mullins, J., and Bird, A.P. (1994). Sp1 sites in
the mouse APRT gene promoter are required to prevent methylation of
the CpG island. Genes Dev 8, 2282–2292.
Madamba, E.V., Berthet, E.B., and Francis, N.J. (2017). Inheritance of
histones H3 and H4 during DNA replication in vitro. Cell Rep 21,
1361–1374.
Margueron, R., Justin, N., Ohno, K., Sharpe, M.L., Son, J., Drury William
J., I., Voigt, P., Martin, S.R., Taylor, W.R., de Marco, V., et al. (2009).
Role of the Polycomb protein EED in the propagation of repressive
histone marks. Nature 461, 762–767.
Margueron, R., and Reinberg, D. (2010). Chromatin structure and the
inheritance of epigenetic information. Nat Rev Genet 11, 285–296.
Margueron, R., and Reinberg, D. (2011). The Polycomb complex PRC2
and its mark in life. Nature 469, 343–349.
Masumoto, H., Hawke, D., Kobayashi, R., and Verreault, A. (2005). A role
for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA
damage response. Nature 436, 294–298.
Mattiroli, F., Gu, Y., Yadav, T., Balsbaugh, J.L., Harris, M.R., Findlay, E.S.,
Liu, Y., Radebaugh, C.A., Stargell, L.A., Ahn, N.G., et al. (2017).
DNA-mediated association of two histone-bound complexes of yeast
chromatin assembly factor-1 (CAF-1) drives tetrasome assembly in the
wake of DNA replication. eLife 6, e22799.
McDonald, O.G., Li, X., Saunders, T., Tryggvadottir, R., Mentch, S.J.,
Warmoes, M.O., Word, A.E., Carrer, A., Salz, T.H., Natsume, S., et al.
(2017). Epigenomic reprogramming during pancreatic cancer
progression links anabolic glucose metabolism to distant metastasis.
Nat Genet 49, 367–376.
McKnight, S.L., and Miller, O.L. Jr. (1977). Electron microscopic analysis
of chromatin replication in the cellular blastoderm Drosophila
melanogaster embryo. Cell 12, 795–804.
Mello, J.A., Silljé, H.H.W., Roche, D.M.J., Kirschner, D.B., Nigg, E.A.,
2186
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
and Almouzni, G. (2002). Human Asf1 and CAF-1 interact and
synergize in a repair-coupled nucleosome assembly pathway. EMBO
Rep 3, 329–334.
Ming, X., Zhang, Z., Zou, Z., Lv, C., Dong, Q., He, Q., Yi, Y., Li, Y., Wang,
H., and Zhu, B. (2021a). Author correction: kinetics and mechanisms of
mitotic inheritance of DNA methylation and their roles in aging-
associated methylome deterioration. Cell Res 31, 373.
Ming, X., Zhu, B., and Li, Y. (2021b). Mitotic inheritance of DNA
methylation: more than just copy and paste. J Genet Genomics 48, 1–
13.
Moazed, D. (2011). Mechanisms for the inheritance of chromatin states.
Cell 146, 510–518.
Mohn, F., Weber, M., Rebhan, M., Roloff, T.C., Richter, J., Stadler, M.B.,
Bibel, M., and Schübeler, D. (2008). Lineage-specific Polycomb targets
and de novo DNA methylation define restriction and potential of
neuronal progenitors. Mol Cell 30, 755–766.
Monk, D., Mackay, D.J.G., Eggermann, T., Maher, E.R., and Riccio, A.
(2019). Genomic imprinting disorders: lessons on how genome,
epigenome and environment interact. Nat Rev Genet 20, 235–248.
Morrison, A.J., and Shen, X. (2009). Chromatin remodelling beyond
transcription: the INO80 and SWR1 complexes. Nat Rev Mol Cell Biol
10, 373–384.
Motamedi, M.R., Verdel, A., Colmenares, S.U., Gerber, S.A., Gygi, S.P.,
and Moazed, D. (2004). Two RNAi complexes, RITS and RDRC,
physically interact and localize to noncoding centromeric RNAs. Cell
119, 789–802.
Muotri, A.R., Marchetto, M.C.N., Coufal, N.G., Oefner, R., Yeo, G.,
Nakashima, K., and Gage, F.H. (2010). L1 retrotransposition in neurons
is modulated by MeCP2. Nature 468, 443–446.
Murayama, A., Sakura, K., Nakama, M., Yasuzawa-Tanaka, K., Fujita, E.,
Tateishi, Y., Wang, Y., Ushijima, T., Baba, T., Shibuya, K., et al. (2006).
A specific CpG site demethylation in the human interleukin 2 gene
promoter is an epigenetic memory. EMBO J 25, 1081–1092.
Nakayama, J., Klar, A.J., and Grewal, S.I. (2000). A chromodomain pro-
tein, Swi6, performs imprinting functions in fission yeast during mitosis
and meiosis. Cell 101, 307–317.
Nakayama, J., Rice, J.C., Strahl, B.D., Allis, C.D., and Grewal, S.I.S.
(2001). Role of histone H3 lysine 9 methylation in epigenetic control of
heterochromatin assembly. Science 292, 110–113.
Nan, X., Ng, H.H., Johnson, C.A., Laherty, C.D., Turner, B.M., Eisenman,
R.N., and Bird, A. (1998). Transcriptional repression by the methyl-
CpG-binding protein MeCP2 involves a histone deacetylase complex.
Nature 393, 386–389.
Neri, F., Incarnato, D., Krepelova, A., Rapelli, S., Pagnani, A., Zecchina,
R., Parlato, C., and Oliviero, S. (2013). Genome-wide analysis
identifies a functional association of Tet1 and Polycomb repressive
complex 2 in mouse embryonic stem cells. Genome Biol 14, R91.
Ng, R.K., and Gurdon, J.B. (2008). Epigenetic memory of an active gene
state depends on histone H3.3 incorporation into chromatin in the
absence of transcription. Nat Cell Biol 10, 102–109.
Nishiyama, A., Mulholland, C.B., Bultmann, S., Kori, S., Endo, A., Saeki,
Y., Qin, W., Trummer, C., Chiba, Y., Yokoyama, H., et al. (2020). Two
distinct modes of DNMT1 recruitment ensure stable maintenance DNA
methylation. Nat Commun 11, 1222.
Nishiyama, A., Yamaguchi, L., Sharif, J., Johmura, Y., Kawamura, T.,
Nakanishi, K., Shimamura, S., Arita, K., Kodama, T., Ishikawa, F., et al.
(2013). Uhrf1-dependent H3K23 ubiquitylation couples maintenance
DNA methylation and replication. Nature 502, 249–253.
Noma, K., Allis, C.D., and Grewal, S.I.S. (2001). Transitions in distinct
histone H3 methylation patterns at the heterochromatin domain
boundaries. Science 293, 1150–1155.
Ohm, J.E., McGarvey, K.M., Yu, X., Cheng, L., Schuebel, K.E., Cope, L.,
Mohammad, H.P., Chen, W., Daniel, V.C., Yu, W., et al. (2007). A stem
cell-like chromatin pattern may predispose tumor suppressor genes to
DNA hypermethylation and heritable silencing. Nat Genet 39, 237–242.
Okano, M., Bell, D.W., Haber, D.A., and Li, E. (1999). DNA
methyltransferases Dnmt3a and Dnmt3b are essential for de novo
methylation and mammalian development. Cell 99, 247–257.
Oksuz, O., Narendra, V., Lee, C.H., Descostes, N., LeRoy, G., Raviram, R.,
Blumenberg, L., Karch, K., Rocha, P.P., Garcia, B.A., et al. (2018).
Capturing the onset of PRC2-mediated repressive domain formation.
Mol Cell 70, 1149–1162.e5.
Ooi, S.K.T., Qiu, C., Bernstein, E., Li, K., Jia, D., Yang, Z., Erdjument-
Bromage, H., Tempst, P., Lin, S.P., Allis, C.D., et al. (2007). DNMT3L
connects unmethylated lysine 4 of histone H3 to de novo methylation of
DNA. Nature 448, 714–717.
Orphanides, G., LeRoy, G., Chang, C.H., Luse, D.S., and Reinberg, D.
(1998). FACT, a factor that facilitates transcript elongation through
nucleosomes. Cell 92, 105–116.
Otani, J., Nankumo, T., Arita, K., Inamoto, S., Ariyoshi, M., and
Shirakawa, M. (2009). Structural basis for recognition of H3K4
methylation status by the DNA methyltransferase 3A ATRX-DNMT3-
DNMT3L domain. EMBO Rep 10, 1235–1241.
Papamichos-Chronakis, M., and Peterson, C.L. (2008). The Ino80
chromatin-remodeling enzyme regulates replisome function and
stability. Nat Struct Mol Biol 15, 338–345.
Papp, B., and Müller, J. (2006). Histone trimethylation and the maintenance
of transcriptional ON and OFF states by trxG and PcG proteins. Genes
Dev 20, 2041–2054.
Peters, J. (2014). The role of genomic imprinting in biology and disease: an
expanding view. Nat Rev Genet 15, 517–530.
Petryk, N., Dalby, M., Wenger, A., Stromme, C.B., Strandsby, A.,
Andersson, R., and Groth, A. (2018). MCM2 promotes symmetric
inheritance of modified histones during DNA replication. Science 361,
1389–1392.
Pinter, S.F., Sadreyev, R.I., Yildirim, E., Jeon, Y., Ohsumi, T.K., Borowsky,
M., and Lee, J.T. (2012). Spreading of X chromosome inactivation via a
hierarchy of defined Polycomb stations. Genome Res 22, 1864–1876.
Plath, K., Mlynarczyk-Evans, S., Nusinow, D.A., and Panning, B. (2002).
Xist RNA and the Mechanism of X Chromosome Inactivation. Annu
Rev Genet 36, 233–278.
Polo, S.E., Theocharis, S.E., Grandin, L., Gambotti, L., Antoni, G.,
Savignoni, A., Asselain, B., Patsouris, E., and Almouzni, G. (2010).
Clinical significance and prognostic value of chromatin assembly
factor-1 overexpression in human solid tumours. Histopathology 57,
716–724.
Popovic, R., Martinez-Garcia, E., Giannopoulou, E.G., Zhang, Q., Zhang,
Q., Ezponda, T., Shah, M.Y., Zheng, Y., Will, C.M., Small, E.C., et al.
(2014). Histone methyltransferase MMSET/NSD2 alters EZH2 binding
and reprograms the myeloma epigenome through global and focal
changes in H3K36 and H3K27 methylation. PLoS Genet 10, e1004566.
Pradhan, S., Bacolla, A., Wells, R.D., and Roberts, R.J. (1999).
Recombinant human DNA (cytosine-5) methyltransferase. J Biol
Chem 274, 33002–33010.
Ptashne, M. (2013). Faddish stuff: epigenetics and the inheritance of
acquired characteristics. FASEB J 27, 1–2.
Putiri, E.L., Tiedemann, R.L., Thompson, J.J., Liu, C., Ho, T., Choi, J.H.,
and Robertson, K.D. (2014). Distinct and overlapping control of 5-
methylcytosine and 5-hydroxymethylcytosine by the TET proteins in
human cancer cells. Genome Biol 15, R81.
Qian, C., Li, S., Jakoncic, J., Zeng, L., Walsh, M.J., and Zhou, M.M.
(2008). Structure and hemimethylated CpG binding of the SRA domain
from human UHRF1. J Biol Chem 283, 34490–34494.
Qin, S., and Min, J. (2014). Structure and function of the nucleosome-
binding PWWP domain. Trends Biochem Sci 39, 536–547.
Qin, W., Wolf, P., Liu, N., Link, S., Smets, M., La Mastra, F., Forné, I.,
Pichler, G., Hörl, D., Fellinger, K., et al. (2015). DNA methylation
requires a DNMT1 ubiquitin interacting motif (UIM) and histone
ubiquitination. Cell Res 25, 911–929.
Ragunathan, K., Jih, G., and Moazed, D. (2015). Epigenetic inheritance
uncoupled from sequence-specific recruitment. Science 348, 1258699.
Raiymbek, G., An, S., Khurana, N., Gopinath, S., Larkin, A., Biswas, S.,
Trievel, R.C., Cho, U.S., and Ragunathan, K. (2020). An H3K9
methylation-dependent protein interaction regulates the non-enzymatic
2187
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
functions of a putative histone demethylase. eLife 9, e53155.
Rajakumara, E., Wang, Z., Ma, H., Hu, L., Chen, H., Lin, Y., Guo, R., Wu,
F., Li, H., Lan, F., et al. (2011). PHD finger recognition of unmodified
histone H3R2 links UHRF1 to regulation of euchromatic gene
expression. Mol Cell 43, 275–284.
Ramachandran, S., and Henikoff, S. (2016). Transcriptional regulators
compete with nucleosomes post-replication. Cell 165, 580–592.
Ramsahoye, B.H., Biniszkiewicz, D., Lyko, F., Clark, V., Bird, A.P., and
Jaenisch, R. (2000). Non-CpG methylation is prevalent in embryonic
stem cells and may be mediated by DNA methyltransferase 3a. Proc
Natl Acad Sci USA 97, 5237–5242.
Rasmussen, K.D., and Helin, K. (2016). Role of TET enzymes in DNA
methylation, development, and cancer. Genes Dev 30, 733–750.
Rea, S., Eisenhaber, F., O’Carroll, D., Strahl, B.D., Sun, Z.W., Schmid, M.,
Opravil, S., Mechtler, K., Ponting, C.P., Allis, C.D., et al. (2000).
Regulation of chromatin structure by site-specific histone H3
methyltransferases. Nature 406, 593–599.
Reddy, B.D., Wang, Y., Niu, L., Higuchi, E.C., Marguerat, S.B., Bähler, J.,
Smith, G.R., and Jia, S. (2011). Elimination of a specific histone H3K14
acetyltransferase complex bypasses the RNAi pathway to regulate
pericentric heterochromatin functions. Genes Dev 25, 214–219.
Reinberg, D., and Vales, L.D. (2018). Chromatin domains rich in
inheritance. Science 361, 33–34.
Remacha, L., Currás-Freixes, M., Torres-Ruiz, R., Schiavi, F., Torres-
Pérez, R., Calsina, B., Letón, R., Comino-Méndez, I., Roldán-Romero,
J.M., Montero-Conde, C., et al. (2018). Gain-of-function mutations in
DNMT3A in patients with paraganglioma. Genet Med 20, 1644–1651.
Ren, W., Fan, H., Grimm, S.A., Guo, Y., Kim, J.J., Yin, J., Li, L., Petell, C.
J., Tan, X.F., Zhang, Z.M., et al. (2020). Direct readout of
heterochromatic H3K9me3 regulates DNMT1-mediated maintenance
DNA methylation. Proc Natl Acad Sci USA 117, 18439–18447.
Ren, W., Fan, H., Grimm, S.A., Kim, J.J., Li, L., Guo, Y., Petell, C.J., Tan,
X.F., Zhang, Z.M., Coan, J.P., et al. (2021). DNMT1 reads
heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation.
Nat Commun 12, 2490.
Rodriguez-Martin, B., Alvarez, E.G., Baez-Ortega, A., Zamora, J., Supek,
F., Demeulemeester, J., Santamarina, M., Ju, Y.S., Temes, J., Garcia-
Souto, D., et al. (2020). Pan-cancer analysis of whole genomes
identifies driver rearrangements promoted by LINE-1
retrotransposition. Nat Genet 52, 306–319.
Rondelet, G., Dal Maso, T., Willems, L., and Wouters, J. (2016). Structural
basis for recognition of histone H3K36me3 nucleosome by human de
novo DNA methyltransferases 3A and 3B. J Struct Biol 194, 357–367.
Rose, N.R., and Klose, R.J. (2014). Understanding the relationship between
DNA methylation and histone lysine methylation. Biochim Biophys
Acta (BBA) - Gene Regulatory Mech 1839, 1362–1372.
Rothbart, S.B., Dickson, B.M., Ong, M.S., Krajewski, K., Houliston, S.,
Kireev, D.B., Arrowsmith, C.H., and Strahl, B.D. (2013). Multivalent
histone engagement by the linked tandem Tudor and PHD domains of
UHRF1 is required for the epigenetic inheritance of DNA methylation.
Genes Dev 27, 1288–1298.
Rothbart, S.B., Krajewski, K., Nady, N., Tempel, W., Xue, S., Badeaux, A.
I., Barsyte-Lovejoy, D., Martinez, J.Y., Bedford, M.T., Fuchs, S.M., et
al. (2012). Association of UHRF1 with methylated H3K9 directs the
maintenance of DNA methylation. Nat Struct Mol Biol 19, 1155–1160.
Rottach, A., Frauer, C., Pichler, G., Bonapace, I.M., Spada, F., and
Leonhardt, H. (2010). The multi-domain protein Np95 connects DNA
methylation and histone modification. Nucl Acids Res 38, 1796–1804.
Sadeghi, L., Prasad, P., Ekwall, K., Cohen, A., and Svensson, J.P. (2015).
The Paf1 complex factors Leo1 and Paf1 promote local histone
turnover to modulate chromatin states in fission yeast. EMBO Rep 16,
1673–1687.
Sado, T., Okano, M., Li, E., and Sasaki, H. (2004). De novo DNA
methylation is dispensable for the initiation and propagation of X
chromosome inactivation. Development 131, 975–982.
Schlesinger, Y., Straussman, R., Keshet, I., Farkash, S., Hecht, M.,
Zimmerman, J., Eden, E., Yakhini, Z., Ben-Shushan, E., Reubinoff, B.
E., et al. (2007). Polycomb-mediated methylation on Lys27 of histone
H3 pre-marks genes for de novo methylation in cancer. Nat Genet 39,
232–236.
Schlissel, G., and Rine, J. (2019). The nucleosome core particle remembers
its position through DNA replication and RNA transcription. Proc Natl
Acad Sci USA 116, 20605–20611.
Schmitges, F.W., Prusty, A.B., Faty, M., Stützer, A., Lingaraju, G.M.,
Aiwazian, J., Sack, R., Hess, D., Li, L., Zhou, S., et al. (2011). Histone
methylation by PRC2 is inhibited by active chromatin marks. Mol Cell
42, 330–341.
Seale, R.L. (1976). Studies on the mode of segregation of histone nu bodies
during replication in HeLa cells. Cell 9, 423–429.
Sendžikaitė, G., Hanna, C.W., Stewart-Morgan, K.R., Ivanova, E., and
Kelsey, G. (2019). A DNMT3A PWWP mutation leads to methylation
of bivalent chromatin and growth retardation in mice. Nat Commun 10,
1884.
Serra-Cardona, A., Duan, S., Yu, C., and Zhang, Z. (2022). H3K4me3
recognition by the COMPASS complex facilitates the restoration of this
histone mark following DNA replication. Sci Adv 8.
Serra-Cardona, A., and Zhang, Z. (2017). Replication-coupled nucleosome
assembly in the passage of epigenetic information and cell identity.
Trends Biochem Sci 43, 136–148.
Shan, C.M., Bao, K., Diedrich, J., Chen, X., Lu, C., Yates Iii, J.R., and Jia,
S. (2020). The INO80 complex regulates epigenetic inheritance of
heterochromatin. Cell Rep 33, 108561.
Shan, C.M., Fang, Y., and Jia, S. (2021). Leaving histone unturned for
epigenetic inheritance. FEBS J, doi: 10.1111/febs.16260.
Sharif, J., Muto, M., Takebayashi, S.I., Suetake, I., Iwamatsu, A., Endo, T.
A., Shinga, J., Mizutani-Koseki, Y., Toyoda, T., Okamura, K., et al.
(2007). The SRA protein Np95 mediates epigenetic inheritance by
recruiting Dnmt1 to methylated DNA. Nature 450, 908–912.
Shibahara, K., and Stillman, B. (1999). Replication-dependent marking of
DNA by PCNA facilitates CAF-1-coupled inheritance of chromatin.
Cell 96, 575–585.
Shilatifard, A. (2012). The COMPASS family of histone H3K4 methylases:
mechanisms of regulation in development and disease pathogenesis.
Annu Rev Biochem 81, 65–95.
Shirohzu, H., Kubota, T., Kumazawa, A., Sado, T., Chijiwa, T., Inagaki, K.,
Suetake, I., Tajima, S., Wakui, K., Miki, Y., et al. (2002). Three novel
DNMT3B mutations in Japanese patients with ICF syndrome. Am J
Med Genet 112, 31–37.
Simon, A.C., Zhou, J.C., Perera, R.L., van Deursen, F., Evrin, C., Ivanova,
M.E., Kilkenny, M.L., Renault, L., Kjaer, S., Matak-Vinković, D., et al.
(2014). A Ctf4 trimer couples the CMG helicase to DNA polymerase α
in the eukaryotic replisome. Nature 510, 293–297.
Smith, S., and Stillman, B. (1991). Stepwise assembly of chromatin during
DNA replication in vitro. EMBO J 10, 971–980.
Soares, L.M., He, P.C., Chun, Y., Suh, H., Kim, T.S., and Buratowski, S.
(2017). Determinants of histone H3K4 methylation patterns. Mol Cell
68, 773–785.e6.
Sobel, R.E., Cook, R.G., Perry, C.A., Annunziato, A.T., and Allis, C.D.
(1995). Conservation of deposition-related acetylation sites in newly
synthesized histones H3 and H4. Proc Natl Acad Sci USA 92, 1237–
1241.
Sorida, M., Hirauchi, T., Ishizaki, H., Kaito, W., Shimada, A., Mori, C.,
Chikashige, Y., Hiraoka, Y., Suzuki, Y., Ohkawa, Y., et al. (2019).
Regulation of ectopic heterochromatin-mediated epigenetic
diversification by the JmjC family protein Epe1. PLoS Genet 15,
e1008129.
Statham, A.L., Robinson, M.D., Song, J.Z., Coolen, M.W., Stirzaker, C.,
and Clark, S.J. (2012). Bisulfite sequencing of chromatin
immunoprecipitated DNA (BisChIP-seq) directly informs methylation
status of histone-modified DNA. Genome Res 22, 1120–1127.
Stillman, B. (1986). Chromatin assembly during SV40 DNA replication in
vitro. Cell 45, 555–565.
Su, D., Hu, Q., Li, Q., Thompson, J.R., Cui, G., Fazly, A., Davies, B.A.,
Botuyan, M.V., Zhang, Z., and Mer, G. (2012). Structural basis for
2188
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
recognition of H3K56-acetylated histone H3-H4 by the chaperone
Rtt106. Nature 483, 104–107.
Sun, X.J., Wei, J., Wu, X.Y., Hu, M., Wang, L., Wang, H.H., Zhang, Q.H.,
Chen, S.J., Huang, Q.H., and Chen, Z. (2005). Identification and
characterization of a novel human histone H3 lysine 36-specific
methyltransferase. J Biol Chem 280, 35261–35271.
Syeda, F., Fagan, R.L., Wean, M., Avvakumov, G.V., Walker, J.R., Xue, S.,
Dhe-Paganon, S., and Brenner, C. (2011). The replication focus
targeting sequence (RFTS) domain is a DNA-competitive inhibitor of
Dnmt1. J Biol Chem 286, 15344–15351.
Sykaras, A.G., Pergaris, A., and Theocharis, S. (2021). Challenging,
accurate and feasible: CAF-1 as a tumour proliferation marker of
diagnostic and prognostic value. Cancers 13, 2575.
Tackett, A.J., Dilworth, D.J., Davey, M.J., O′Donnell, M., Aitchison, J.D.,
Rout, M.P., and Chait, B.T. (2005). Proteomic and genomic
characterization of chromatin complexes at a boundary. J Cell Biol
169, 35–47.
Tagami, H., Ray-Gallet, D., Almouzni, G., and Nakatani, Y. (2004).
Histone H3.1 and H3.3 complexes mediate nucleosome assembly
pathways dependent or independent of DNA synthesis. Cell 116, 51–61.
Takeshita, K., Suetake, I., Yamashita, E., Suga, M., Narita, H., Nakagawa,
A., and Tajima, S. (2011). Structural insight into maintenance
methylation by mouse DNA methyltransferase 1 (Dnmt1). Proc Natl
Acad Sci USA 108, 9055–9059.
Talbert, P.B., and Henikoff, S. (2010). Histone variants — ancient wrap
artists of the epigenome. Nat Rev Mol Cell Biol 11, 264–275.
Talbert, P.B., and Henikoff, S. (2021). The Yin and Yang of histone marks
in transcription. Annu Rev Genom Hum Genet 22, 147–170.
Tamaru, H., and Selker, E.U. (2001). A histone H3 methyltransferase
controls DNA methylation in Neurospora crassa. Nature 414, 277–283.
Tamaru, H., Zhang, X., McMillen, D., Singh, P.B., Nakayama, J., Grewal,
S.I., Allis, C.D., Cheng, X., and Selker, E.U. (2003). Trimethylated
lysine 9 of histone H3 is a mark for DNA methylation in Neurospora
crassa. Nat Genet 34, 75–79.
Tan, B.C.M., Chien, C.T., Hirose, S., and Lee, S.C. (2006). Functional
cooperation between FACT and MCM helicase facilitates initiation of
chromatin DNA replication. EMBO J 25, 3975–3985.
Tanaka, T., Knapp, D., and Nasmyth, K. (1997). Loading of an Mcm
protein onto DNA replication origins is regulated by Cdc6p and CDKs.
Cell 90, 649–660.
Tartof, K.D., Hobbs, C., and Jones, M. (1984). A structural basis for
variegating position effects. Cell 37, 869–878.
Tate, P.H., and Bird, A.P. (1993). Effects of DNA methylation on DNA-
binding proteins and gene expression. Curr Opin Genet Dev 3, 226–
231.
Thomassin, H., Flavin, M., Espinás, M.L., and Grange, T. (2001).
Glucocorticoid-induced DNA demethylation and gene memory during
development. EMBO J 20, 1974–1983.
Trewick, S.C., Minc, E., Antonelli, R., Urano, T., and Allshire, R.C. (2007).
The JmjC domain protein Epe1 prevents unregulated assembly and
disassembly of heterochromatin. EMBO J 26, 4670–4682.
Tsukada, Y.I., Fang, J., Erdjument-Bromage, H., Warren, M.E., Borchers,
C.H., Tempst, P., and Zhang, Y. (2006). Histone demethylation by a
family of JmjC domain-containing proteins. Nature 439, 811–816.
Tyler, J.K., Adams, C.R., Chen, S.R., Kobayashi, R., Kamakaka, R.T., and
Kadonaga, J.T. (1999). The RCAF complex mediates chromatin
assembly during DNA replication and repair. Nature 402, 555–560.
Tyler, J.K., Collins, K.A., Prasad-Sinha, J., Amiott, E., Bulger, M., Harte, P.
J., Kobayashi, R., and Kadonaga, J.T. (2001). Interaction between the
Drosophila CAF-1 and ASF1 Chromatin Assembly Factors. Mol Cell
Biol 21, 6574–6584.
VanDemark, A.P., Blanksma, M., Ferris, E., Heroux, A., Hill, C.P., and
Formosa, T. (2006). The structure of the yFACT Pob3-M domain, its
interaction with the DNA replication factor RPA, and a potential role in
nucleosome deposition. Mol Cell 22, 363–374.
Vaughan, R.M., Dickson, B.M., Whelihan, M.F., Johnstone, A.L., Cornett,
E.M., Cheek, M.A., Ausherman, C.A., Cowles, M.W., Sun, Z.W., and
Rothbart, S.B. (2018). Chromatin structure and its chemical
modifications regulate the ubiquitin ligase substrate selectivity of
UHRF1. Proc Natl Acad Sci USA 115, 8775–8780.
Verdel, A.., Jia, S., Gerber, S., Sugiyama, T., Gygi, S., Grewal, S.I.S., and
Moazed, D. (2004). RNAi-mediated targeting of heterochromatin by the
RITS complex. Science 303, 672–676.
Verma, N., Pan, H., Doré, L.C., Shukla, A., Li, Q.V., Pelham-Webb, B.,
Teijeiro, V., González, F., Krivtsov, A., Chang, C.J., et al. (2018). TET
proteins safeguard bivalent promoters from de novo methylation in
human embryonic stem cells. Nat Genet 50, 83–95.
Verreault, A., Kaufman, P.D., Kobayashi, R., and Stillman, B. (1996).
Nucleosome assembly by a complex of CAF-1 and acetylated histones
H3/H4. Cell 87, 95–104.
Vincent, J.A., Kwong, T.J., and Tsukiyama, T. (2008). ATP-dependent
chromatin remodeling shapes the DNA replication landscape. Nat Struct
Mol Biol 15, 477–484.
Volk, A., Liang, K., Suraneni, P., Li, X., Zhao, J., Bulic, M., Marshall, S.,
Pulakanti, K., Malinge, S., Taub, J., et al. (2018). A CHAF1B-
dependent molecular switch in hematopoiesis and leukemia
pathogenesis. Cancer Cell 34, 707–723.e7.
Volpe, T.A., Kidner, C., Hall, I.M., Teng, G., Grewal, S.I.S., and
Martienssen, R.A. (2002). Regulation of heterochromatic silencing
and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–
1837.
Waddington, C.H. (1942). The epigenotype. Endeavour 1, 18–20.
Wagner, E.J., and Carpenter, P.B. (2012). Understanding the language of
Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13, 115–126.
Wang, C., Zhu, B., and Xiong, J. (2018a). Recruitment and reinforcement:
maintaining epigenetic silencing. Sci China Life Sci 61, 515–522.
Wang, J., Reddy, B.D., and Jia, S. (2015). Rapid epigenetic adaptation to
uncontrolled heterochromatin spreading. eLife 4, e06179.
Wang, J., Tadeo, X., Hou, H., Tu, P.G., Thompson, J., Yates III, J.R., and
Jia, S. (2013). Epe1 recruits BET family bromodomain protein Bdf2 to
establish heterochromatin boundaries. Genes Dev 27, 1886–1902.
Wang, Q., Yu, G., Ming, X., Xia, W., Xu, X., Zhang, Y., Zhang, W., Li, Y.,
Huang, C., Xie, H., et al. (2020). Imprecise DNMT1 activity coupled
with neighbor-guided correction enables robust yet flexible epigenetic
inheritance. Nat Genet 52, 828–839.
Wang, T., Liu, Y., Edwards, G., Krzizike, D., Scherman, H., and Luger, K.
(2018b). The histone chaperone FACT modulates nucleosome structure
by tethering its components. Life Sci Alliance 1, e201800107.
Wang, X., and Moazed, D. (2017). DNA sequence-dependent epigenetic
inheritance of gene silencing and histone H3K9 methylation. Science
356, 88–91.
Wang, X., Paulo, J.A., Li, X., Zhou, H., Yu, J., Gygi, S.P., and Moazed, D.
(2021). A composite DNA element that functions as a maintainer
required for epigenetic inheritance of heterochromatin. Mol Cell 81,
3979–3991.e4.
Weinberg, D.N., Papillon-Cavanagh, S., Chen, H., Yue, Y., Chen, X.,
Rajagopalan, K.N., Horth, C., McGuire, J.T., Xu, X., Nikbakht, H., et
al. (2019). The histone mark H3K36me2 recruits DNMT3A and shapes
the intergenic DNA methylation landscape. Nature 573, 281–286.
Weinberg, D.N., Rosenbaum, P., Chen, X., Barrows, D., Horth, C.,
Marunde, M.R., Popova, I.K., Gillespie, Z.B., Keogh, M.C., Lu, C., et
al. (2021). Two competing mechanisms of DNMT3A recruitment
regulate the dynamics of de novo DNA methylation at PRC1-targeted
CpG islands. Nat Genet 53, 794–800.
Williams, K., Christensen, J., and Helin, K. (2012). DNA methylation: TET
proteins—guardians of CpG islands? EMBO Rep 13, 28–35.
Wu, H., Zeng, H., Lam, R., Tempel, W., Amaya, M.F., Xu, C., Dombrovski,
L., Qiu, W., Wang, Y., and Min, J. (2011). Structural and histone
binding ability characterizations of human PWWP domains. PLoS ONE
6, e18919.
Xu, C., and Corces, V.G. (2018). Nascent DNA methylome mapping
reveals inheritance of hemimethylation at CTCF/cohesin sites. Science
359, 1166–1170.
Xu, M., Long, C., Chen, X., Huang, C., Chen, S., and Zhu, B. (2010).
2189
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
Partitioning of histone H3-H4 tetramers during DNA replication-
dependent chromatin assembly. Science 328, 94–98.
Xu, M., Wang, W., Chen, S., and Zhu, B. (2011). A model for mitotic
inheritance of histone lysine methylation. EMBO Rep 13, 60–67.
Yamaguchi, L., Nishiyama, A., Misaki, T., Johmura, Y., Ueda, J., Arita, K.,
Nagao, K., Obuse, C., and Nakanishi, M. (2017). Usp7-dependent
histone H3 deubiquitylation regulates maintenance of DNA
methylation. Sci Rep 7, 55.
Yang, J., Zhang, X., Feng, J., Leng, H., Li, S., Xiao, J., Liu, S., Xu, Z., Xu,
J., Li, D., et al. (2016). The histone chaperone fact contributes to DNA
replication-coupled nucleosome assembly. Cell Rep 16, 3414.
Yarychkivska, O., Shahabuddin, Z., Comfort, N., Boulard, M., and Bestor,
T.H. (2018). BAH domains and a histone-like motif in DNA
methyltransferase 1 (DNMT1) regulate de novo and maintenance
methylation in vivo. J Biol Chem 293, 19466–19475.
Youmans, D.T., Schmidt, J.C., and Cech, T.R. (2018). Live-cell imaging
reveals the dynamics of PRC2 and recruitment to chromatin by SUZ12-
associated subunits. Genes Dev 32, 794–805.
Yu, C., Gan, H., Serra-Cardona, A., Zhang, L., Gan, S., Sharma, S.,
Johansson, E., Chabes, A., Xu, R.M., and Zhang, Z. (2018a). A
mechanism for preventing asymmetric histone segregation onto
replicating DNA strands. Science 361, 1386–1389.
Yu, M., and Ren, B. (2017). The three-dimensional organization of
mammalian genomes. Annu Rev Cell Dev Biol 33, 265–289.
Yu, R., Wang, X., and Moazed, D. (2018b). Epigenetic inheritance
mediated by coupling of RNAi and histone H3K9 methylation. Nature
558, 615–619.
Yuan, W., Xu, M., Huang, C., Liu, N., Chen, S., and Zhu, B. (2011). H3K36
methylation antagonizes PRC2-mediated H3K27 methylation. J Biol
Chem 286, 7983–7989.
Zaratiegui, M., Irvine, D.V., and Martienssen, R.A. (2007). Noncoding
RNAs and gene silencing. Cell 128, 763–776.
Zee, B.M., Levin, R.S., DiMaggio, P.A., and Garcia, B.A. (2010). Global
turnover of histone post-translational modifications and variants in
human cells. Epigenet Chromatin 3, 22.
Zhang, B., Zheng, H., Huang, B., Li, W., Xiang, Y., Peng, X., Ming, J., Wu,
X., Zhang, Y., Xu, Q., et al. (2016). Allelic reprogramming of the
histone modification H3K4me3 in early mammalian development.
Nature 537, 553–557.
Zhang, H., Gao, Q., Tan, S., You, J., Lyu, C., Zhang, Y., Han, M., Chen, Z.,
Li, J., Wang, H., et al. (2019). SET8 prevents excessive DNA
methylation by methylation-mediated degradation of UHRF1 and
DNMT1. Nucleic Acids Res, doi: 10.1093/nar/gkz626.
Zhang, H., Han, J., Kang, B., Burgess, R., and Zhang, Z. (2012). Human
histone acetyltransferase 1 protein preferentially acetylates H4 histone
molecules in H3.1-H4 over H3.3-H4. J Biol Chem 287, 6573–6581.
Zhang, K., Mosch, K., Fischle, W., and Grewal, S.I.S. (2008). Roles of the
Clr4 methyltransferase complex in nucleation, spreading and
maintenance of heterochromatin. Nat Struct Mol Biol 15, 381–388.
Zhang, L., Serra-Cardona, A., Zhou, H., Wang, M., Yang, N., Zhang, Z.,
and Xu, R.M. (2018). Multisite substrate recognition in Asf1-dependent
acetylation of histone H3 K56 by Rtt109. Cell 174, 818–830.e11.
Zhang, Y., Jurkowska, R., Soeroes, S., Rajavelu, A., Dhayalan, A., Bock, I.,
Rathert, P., Brandt, O., Reinhardt, R., Fischle, W., et al. (2010).
Chromatin methylation activity of Dnmt3a and Dnmt3a/3L is guided by
interaction of the ADD domain with the histone H3 tail. Nucl Acids Res
38, 4246–4253.
Zhang, Z., Shibahara, K., and Stillman, B. (2000). PCNA connects DNA
replication to epigenetic inheritance in yeast. Nature 408, 221–225.
Zhao, J., Wang, M., Chang, L., Yu, J., Song, A., Liu, C., Huang, W., Zhang,
T., Wu, X., Shen, X., et al. (2020a). RYBP/YAF2-PRC1 complexes and
histone H1-dependent chromatin compaction mediate propagation of
H2AK119ub1 during cell division. Nat Cell Biol 22, 439–452.
Zhao, Q., Zhang, J., Chen, R., Wang, L., Li, B., Cheng, H., Duan, X., Zhu,
H., Wei, W., Li, J., et al. (2016). Dissecting the precise role of H3K9
methylation in crosstalk with DNA maintenance methylation in
mammals. Nat Commun 7, 12464.
Zhao, Y., and Garcia, B.A. (2015). Comprehensive catalog of currently
documented histone modifications. Cold Spring Harb Perspect Biol 7,
a025064.
Zhao, Z., Lan, M., Li, J., Dong, Q., Li, X., Liu, B., Li, G., Wang, H., Zhang,
Z., and Zhu, B. (2019). The proinflammatory cytokine TNFα induces
DNA demethylation-dependent and -independent activation of
interleukin-32 expression. J Biol Chem 294, 6785–6795.
Zhao, Z., Zhang, Z., Li, J., Dong, Q., Xiong, J., Li, Y., Lan, M., Li, G., and
Zhu, B. (2020b). Sustained TNF-α stimulation leads to transcriptional
memory that greatly enhances signal sensitivity and robustness. eLife 9,
e61965.
Zhou, H., Madden, B.J., Muddiman, D.C., and Zhang, Z. (2006).
Chromatin assembly factor 1 interacts with histone H3 methylated at
lysine 79 in the processes of epigenetic silencing and DNA repair.
Biochemistry 45, 2852–2861.
Zhou, K., Gaullier, G., and Luger, K. (2019). Nucleosome structure and
dynamics are coming of age. Nat Struct Mol Biol 26, 3–13.
Zhu, B., and Reinberg, D. (2011). Epigenetic inheritance: uncontested? Cell
Res 21, 435–441.
Ziller, M.J., Gu, H., Müller, F., Donaghey, J., Tsai, L.T.Y., Kohlbacher, O.,
De Jager, P.L., Rosen, E.D., Bennett, D.A., Bernstein, B.E., et al.
(2013). Charting a dynamic DNA methylation landscape of the human
genome. Nature 500, 477–481.
2190
Du, W., et al. Sci China Life Sci November (2022) Vol.65 No.11
