
Research
Global DNA hypomethylation coupled to repressive
chromatin domain formation and gene silencing
in breast cancer
Gary C. Hon,
1
R. David Hawkins,
1
Otavia L. Caballero,
2
Christine Lo,
3
Ryan Lister,
4
Mattia Pelizzola,
4
Armand Valsesia,
5
Zhen Ye,
1
Samantha Kuan,
1
Lee E. Edsall,
1
Anamaria Aranha Camargo,
6
Brian J. Stevenson,
5
Joseph R. Ecker,
4
Vineet Bafna,
3
Robert L. Strausberg,
2,7
Andrew J. Simpson,
2,7
and Bing Ren
1,8,9
1
Ludwig Institute for Cancer Research, La Jolla, California 92093, USA;
2
Ludwig Collaborative Laboratory for Cancer Biology
and Therapy, Department of Neurosurgery, Johns Hopkins University School of Medicine, Baltimore, Maryland 21231, USA;
3
Department of Computer Science, University of California San Diego, San Diego, California 92093, USA;
4
Genomic Analysis
Laboratory, Howard Hughes Medical Institute, and The Salk Institute for Biological Studies, La Jolla, California 92037, USA;
5
Swiss
Institute of Bioinformatics, Ludwig Institute for Cancer Research, University of Lausanne, 1011 Lausanne, Switzerland;
6
Ludwig
Institute for Cancer Research, 01323-903 Sao Paulo, SP, Brazil;
7
Ludwig Institute for Cancer Research Ltd., New York, New York
10017, USA;
8
Department of Cellular and Molecular Medicine, Moores Cancer Center, and Institute of Genomic Medicine, University
of California San Diego, La Jolla, California 92093, USA
While genetic mutation is a hallmark of cancer, many cancers also acquire epigenetic alterations during tumorigenesis
including aberrant DNA hypermethylation of tumor suppressors, as well as changes in chromatin modifications as caused
by genetic mutations of the chromatin-modifying machinery. However, the extent of epigenetic alterations in cancer cells
has not been fully characterized. Here, we describe complete methylome maps at single nucleotide resolution of a low-
passage breast cancer cell line and primary human mammary epithelial cells. We find widespread DNA hypomethylation
in the cancer cell, primarily at partially methylated domains (PMDs) in normal breast cells. Unexpectedly, genes within
these regions are largely silenced in cancer cells. The loss of DNA methylation in these regions is accompanied by
formation of repressive chromatin, with a significant fraction displaying allelic DNA methylation where one allele is DNA
methylated while the other allele is occupied by histone modifications H3K9me3 or H3K27me3. Our results show
a mutually exclusive relationship between DNA methylation and H3K9me3 or H3K27me3. These results suggest that
global DNA hypomethylation in breast cancer is tightly linked to the formation of repressive chromatin domains and
gene silencing, thus identifying a potential epigenetic pathway for gene regulation in cancer cells.
[Supplemental material is available for this article.]
Breast cancer is characterized by both genetic and epigenetic al-
terations (Sjoblom et al. 2006; Esteller 2007, 2008; Wood et al.
2007; Stephens et al. 2009). While a large number of genetic mu-
tations are linked to breast cancer, there is clear evidence that
epigenetic alterations, such as hypo- or hypermethylation of
DNA, occur early in the initiation or development of the tumors
(Kohonen-Corish et al. 2007). Some genes commonly hyper-
methylated in breast cancers are involved in evasion of apoptosis
(RASSF1, HOXA5, TWIST1) and cellular senescence (CCND2,
CDKN2A), while others regulate DNA repair (BRCA1), cell growth
(ESR1, PGR), and tissue invasion (CDH1) (Dworkin et al. 2009;
Jovanovic et al. 2010). Further underscoring the role of the epige-
netic mechanisms in tumorigenesis, such epigenetic events have
been exploited for early diagnosis or treatment. For example, a
therapeutic strategy blocking DNA methylation with 5-azacytidine
( Jones et al. 1983) has been approved for treatment of preleukemic
myelodysplastic syndrome (Kaminskas et al. 2005) and is in clin-
ical trials for several forms of cancer (Kelly et al. 2010).
DNA methylation (Holliday 1979; Feinberg and Vogelstein
1983; Laird 2003, 2010) is the most studied epigenetic event in
cancer. Bisulfite sequencing (Frommer et al. 1992) of targeted loci
such as the breast cancer susceptibility gene BRCA1 (Tapia et al.
2008) supports the notion that tumor suppressors are frequently
inactivated by DNA methylation at CpG islands and promoters.
Genome-scale methods including MeDIP-seq (Ruike et al. 2010)
and CHARM (Irizarry et al. 2009) have confirmed global hypo-
methylation and focal hypermethylation as hallmarks of breast
and colon cancer. More recently, whole genome shotgun bisulfite
sequencing offers single-nucleotide resolution of DNA methyla-
tion in human cells (Lister et al. 2009). This unprecedented reso-
lution has revealed that cytosines methylated in the CG context
(mCG) are nearly completely methylated in pluripotent cells but
are frequently in a partially methylated state in somatic cells. These
partially methylated cytosines are clustered to form partially
methylated domains (PMDs), which can span nearly 40% the ge-
nome (Lister et al. 2009). Interestingly, genes within PMDs are
found to be generally repressed, though the mechanism is unclear
(Lister et al. 2011). This method has also recently been used to
9
Corresponding author.
E-mail [email protected].
Article published online before print. Article, supplemental material, and pub-
lication date are at http://www.genome.org/cgi/doi/10.1101/gr.125872.111.
246 Genome Research
www.genome.org
22:246–258 Ó 2012 by Cold Spring Harbor Laboratory Press; ISSN 1088-9051/12; www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
observe increased epigenetic variation at hypomethylated regions
in cancer cells (Hansen et al. 2011). Finally, further underscoring
the role of DNA methylation in cancer, a recent genetic study
showed that mutations in the DNA methyltransferase DNMT3A
are frequently found in acute myeloid leukemia (Ley et al. 2010).
Chromatin state can also be altered in cancer cells (Parsons
et al. 2010; Jiao et al. 2011; Varela et al. 2011). For example,
heterochromatin-associated H3K9me3 and Polycomb-associated
H3K27me3 mark large repressed domains in somatic cells (Hawkins
et al. 2010) that are misregulated in cancer. H3K9me3 is deposited
by a family of histone methyltransferases including SUV39H1,
inhibition of which in acute myeloid leukemia cells is sufficient for
re-expression of the transcriptionally silenced tumor suppressor genes
CDKN2B and CDH1 marked by H3K9me3 (Lakshmikuttyamma
et al. 2009). EZH2, the enzyme responsible for depositing H3K27me3,
is often overexpressed in aggressive breast cancers (Kleer et al. 2003;
Chang et al. 2011), and mutations in the H3K27me3 demethylase
KDM6A are common in clear cell renal cell carcinoma (Dalgliesh
et al. 2010). Thus, misregulation of repressive chromatin modifica-
tions may also play a role in tumorigenesis.
While both DNA methylation and chromatin modifications
have been associated with tumorigenesis, few studies have in-
tegrated both aspects on a global scale to investigate their co-
ordinated role in cancer progression. Here, we report use of high-
throughput sequencing technology to map DNA methylation at
base resolution and two repressive chromatin modifications in
a breast cancer cell line and primary mammary epithelial cells.
Comparative analysis of the two epigenomes reveals widespread
DNA hypomethylation that is tightly coupled to the formation of
repressive chromatin domains and gene silencing in the cancer
cells. We propose that such large scale alterations of the epigenetic
landscape may play an important role in tumorigenesis by inhib-
iting expression of tumor suppressor genes. We also suggest that
global hypomethylation may occur through a passive mechanism.
Further, we show that, while hypomethylation of repetitive ele-
ments is common, it is not the only explanation for increased
transcription from such repetitive sequences.
Results
Global DNA hypomethylation in the breast cancer cell
line HCC1954
The HCC1954 cell line is derived from the primary tissue of a
ductal breast carcinoma of an East Indian female (Neve et al. 2006).
It belongs to the subtype of estrogen receptor (ER)/progesterone
receptor (PR) negative and ERBB2 (HER2) positive breast cancers
characterized by poor prognosis. We performed whole genome
shotgun sequencing of bisulfite-treated DNA [MethylC-seq (Lister
et al. 2008, 2009)] in HCC1954, generating 889,012,059 uniquely
mapped monoclonal reads, with an average of 27-fold genome
coverage. As a control, we also performed MethylC-seq on human
mammary epithelial cells (HMECs) at 20-fold coverage.
DNA methylation in both HCC1954 and HMEC exists almost
exclusively (>99.8%) in the CG context. To compare the global
profiles of DNA methylation between HCC1954 and HMEC, we
quantified mCG by dividing the genome into 10-kb windows and
calculating the percentage of cytosine bases methylated in se-
quenced reads (%mCG). We found that a large fraction of the
HCC1954 genome is differentially methylated as compared to nor-
mal cells (Fig. 1A; Supplemental Fig. S1). In agreement with pre-
vious studies (Esteller 2008; Ruike et al. 2010), we observed global
hypomethylation and local hypermethylation in HCC1954 com-
pared to HMEC. Denoting the four quartiles of %mCG from 0% to
100% as low, mid-low, mid-high, and high (Fig. 1A), we observed
that a striking 22.3% of these 10-kb windows have low mCG in
HCC1954, compared to 0.64% in HMEC. On the other hand,
hypermethylation is more limited, with 3.1% of HCC1954 win-
dows exhibiting %mCG >95%, compared to 0.3% in HMEC.
These domains of hypomethylation span known tumor
suppressors such as DACH1 (Fig. 1B). On a chromosome-wide view, it
is clear that hypomethylated regions form large domains, which are
punctuated by regions where DNA methylation is high (Fig. 1C).
Chromosome-wide views also suggest that loci with the most
pronounced hypomethylation in HCC1954 coincide with regions
that are not fully methylated in HMEC (Fig. 1C). Genome-wide,
88.2% of all low mCG windows in HCC1954 are in a PMD state
(defined as mid-low or mid-high %mCG) in HMEC (Fig. 1D). Also,
of all hypermethylated HCC1954 regions that are not highly
methylated in HMEC, 99.6% belong to the PMD state in HMEC.
These results suggest that PMDs in HMEC are the most likely regions
to either gain or lose DNA methylation during tumorigenesis.
To test if epigenetically unstable regions in breast cancer (BC)
might coincide with those in other cancers, we compared our re-
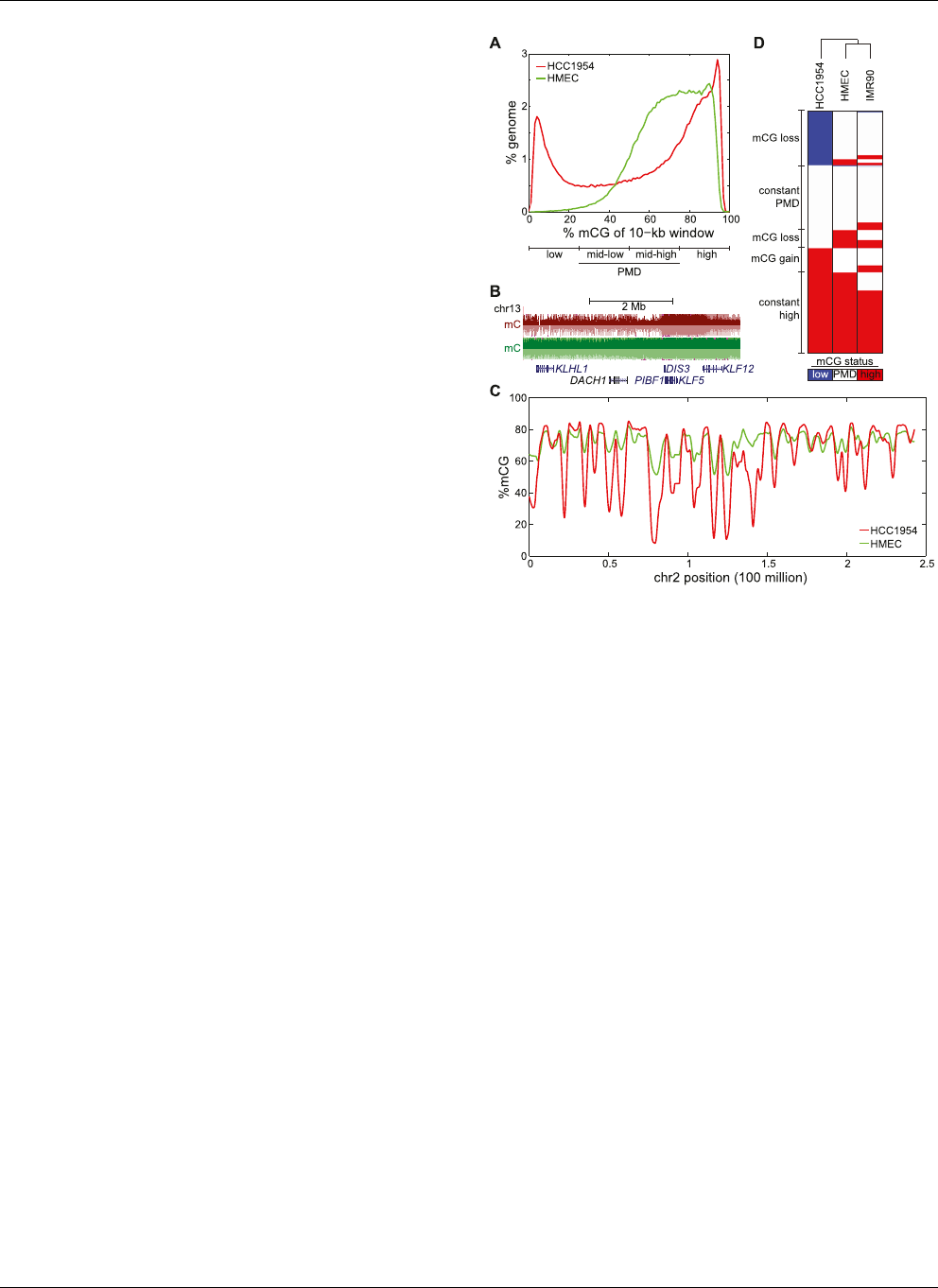
Figure 1. Global hypomethylation in breast cancer. (A) Distribution of
%mCG for all 10-kb windows in the human genome. Quartiles of in-
creasing %mCG are labeled as low, mid -low, mid-high, and high %mCG,
with mid-low and mid-high representing partially methylated domains
(PMDs). (B) A large domain of hypomethylation near the DACH1 tumor
suppressor. (Red) HCC1954, (green) HMEC. (C ) Distribution of %mCG on
chromosome 2 for the breast cancer cell line HCC1954 and the normal
breast line HMEC. (D) A heatmap of low, PMD, and high %mCG for all 10-
kb windows in the human genome. Each of the 282,109 rows represents
the mCG status for a 10-kb window in HCC1954, HMEC, and IMR90 fi-
broblast cells. The dendrogram represents the similarity (Pearson corre-
lation) of the profiles across different cells.
Genome Research 247
www.genome.org
DNA hypomethylation linkedtochromatinsilencing
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
sults to those obtained by a recent study
which cataloged epigenetically unstable
regions in colon cancer (CC)(Irizarry et al.
2009). Regions showing hypomethylation
in CC (N = 994) are also generally hypo-
methylated in BC: Of 730 hypomethyated
CC loci showing methylation bias in BC
(Fisher’s P-value # 0.01), 537 (73.6%) are
hypomethylated compared to 193 that
are hypermethylated (Supplemental Fig.
S2) (P < 10
100
, binomial). In contrast,
hypermethylated CC regions (N = 704)
tend to be hypermethylated in BC: of 555
hypermethylated CC loci with methyla-
tion bias in BC, 485 (87.4%) are hyper-
methylated and 70 are hypomethylated
(Supplemental Fig. S2) (P < 10
100
,bino-
mial). This suggests that certain regions of
the genome exhibit inherent epigenetic
instability and consistently gain or lose
mCG in multiple types of cancers. In
agreement with Irizarry et al. (2009),
overlapping hypo- and hypermethylated
genes are enriched for embryonic and
developmental proteins, including the
reprogramming factor SOX2,neural
developmental genes NEGR1 and NRG1,
and members of the HOXA cluster (HOXA1
through HOXA7,andHOXA9).
DNA hypomethylation is associated
with decreased gene expression
in breast cancers
DNA hypermethylation at promoters is
generally correlated with gene silencing.
Recently, DNA methylation at gene bod-
ies has been shown to be prevalent in
mammalian cells, and some evidence has
suggested it is correlated with gene ac-
tivity (Hellman and Chess 2007; Ball et al.
2009), but the mechanisms have not
been clearly understood (Jones 1999;
Maunakea et al. 2010; Wu et al. 2010). T o
explore the effect of DNA hypomethylation on gene expression in
breast cancer cells, we performed RNA-seq experiments to de-
termine genome-wide steady-state transcript abundance in
both HCC1954 and HMEC cells. We then examined the transcript
abundance for genes overlapping domains with DNA hypo-
methylation. Unexpectedly, these genes have a greater tendency to
be repressed in HCC1954: While only 3.8% of all genes transition
from highly methylated to PMD states, this represents 14.1% of all
genes losing expression (p
high!PMD
< 10
16
, hypergeometric) (Fig.
2A). Likewise, the 220 genes undergoing PMD to low %mCG
transition represent 12.1% of all down-regulated genes, compared
to 8.1% expected by chance (p
PMD!low
= 1.08 3 10
9
, hyper-
geometric). In contrast, hypomethylated genes are less likely to
gain expression HCC1954 (Fig. 2A) (p
high!PMD
= 5.7 3 10
6
,
p
PMD!low
= 1.03 3 10
30
, hypergeometric). There is also significant
overlap between down-regulated, hypomethylated genes in
HCC1954 with hypomethylated genes in colon cancer (P = 1.03 3
10
7
, hypergeometric).
To test if this pattern of gene expression is representative of
a broader set of breast cancers, we examined the gene expression
profiles of a panel of 50 ERBB2 positive breast tumors and 23
normal breast samples (Hennessy et al. 2009; Parker et al. 2009;
Prat et al. 2010). In this data set, 13,262 genes could be un-
ambiguously assigned to transcripts from our RNA-seq experi-
ments. At 935 of these genes undergoing hypomethylation from
HMEC to HCC1954, we similarly observe an enrichment of cancer-
specifically repressed genes (p
high!PMD
= 8.86 3 10
7
,p
PMD!low
=
1.38 3 10
3
, hypergeometric) and a depletion of cancer-specifically
expressed genes (Fig. 2B) (p
high!PMD
= 2.79 3 10
4
,p
PMD!low
=
4.43 3 10
5
, hypergeometric). Interestingly, similar results are also
observed when comparing non-ERBB2 positive breast tumors with
normal breast samples (data not shown), suggesting that genes
undergoing hypomethylation in HCC1954 are generally repressed
in breast cancers.
To understand whether specific cellular pathways are partic-
ularly affected by the abnormal hypomethylation in HCC1954, we
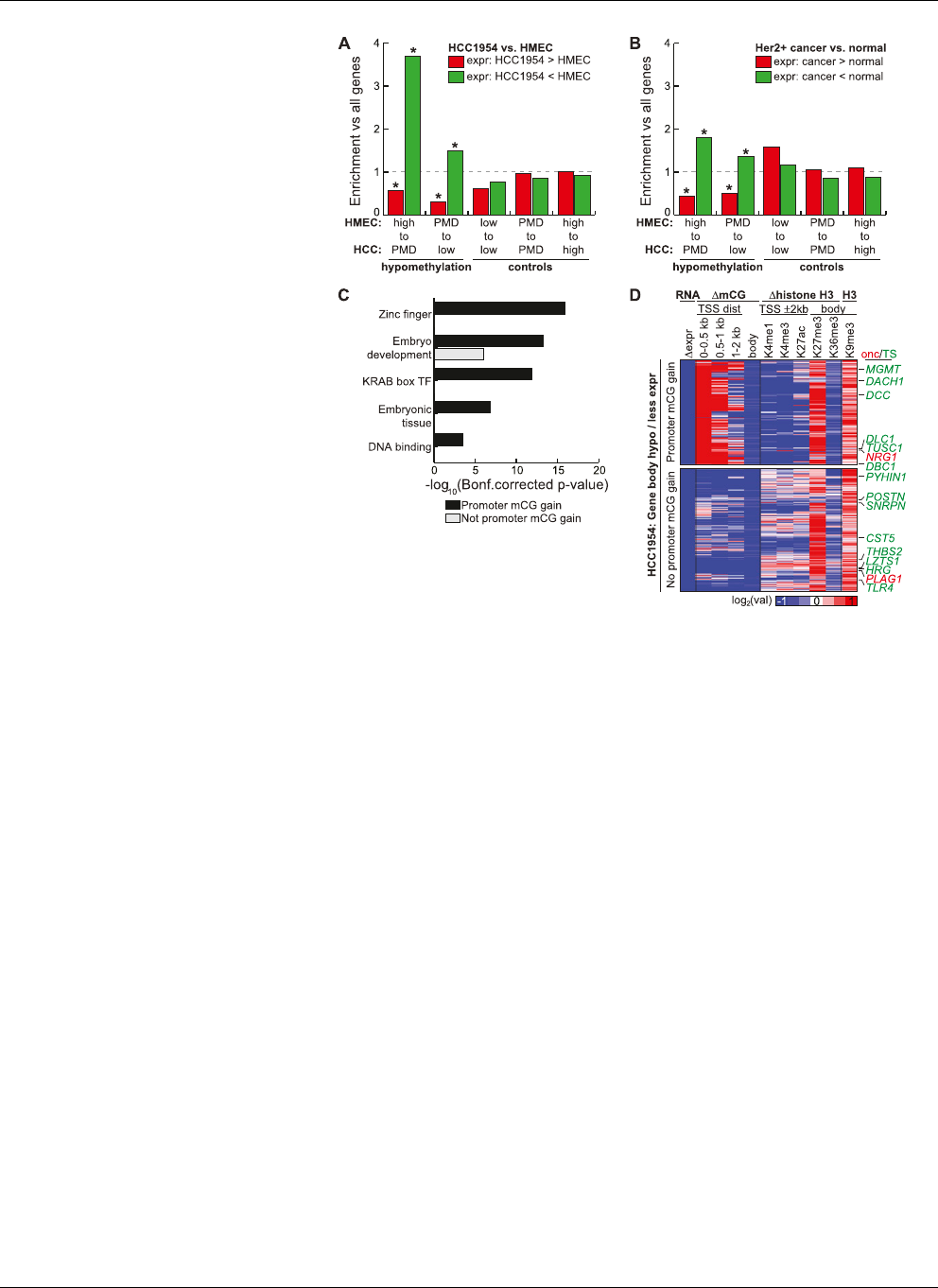
Figure 2. Gene body hypomethylation is associated with gene repression. (A) Genes were overlapped
with 10-kb domains undergoing hypomethylation (high to PMD, PMD to low) or control (low to low,
PMD to PMD, high to high) transitions. Shown is the enrichment of each transition state with genes having
HCC1954 expression at least eightfold more (red) or less (green) than HMEC, when compared to the
global enrichment of all transition states. (*) P-value # 0.01 (hypergeometric). (B)Usingthesametran-
sition states in A, but comparing the expression of genes in a panel of 50 ERBB2+ (HER2+)breastcancers
compared to a panel of 23 normal breast samples (Weigelt et al. 2005; Oh et al. 2006; Perreard et al. 2006;
Herschkowitz et al. 2007, 2008; Hoadley et al. 2007; Mullins et al. 2007; Hennessy et al. 2009; Hu et al.
2009; Parker et al. 2009; Prat et al. 2010). Significantly more (red) or less (green) expressed genes are
defined by a Wilcoxon rank sum test (P-value # 0.01) between the expression values of the two different
panels. (*) Enrichment P-value # 0.01 (hypergeometric). (C ) Functional enrichment of hypomethylated
genes having loss of expression by the DAVID analysis tool (Dennis et al. 2003). (D) Epigenetic status of
genes undergoing hypomethylation in gene bodies with loss of expression. With the exception of
H3K9me3, all values are of HCC1954 relative to HMEC. RNA, log
2
(HCC1954 RPKM/HMEC RPKM);
DmCG, log
2
(HCC1954%mCG/HMEC %mCG); Dhistone, log
2
(HCC1954 ChIP/input) log
2
(HMEC
ChIP/input); H3K9me3, log
2
(HCC1954 ChIP/input). (Onc) oncogene, (TS) tumor suppressor.
Hon et al.
248 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

performed gene ontology (GO) analysis for the genes found within
the hypomethylated domains. Consistent with previous analysis
of colon cancer-specifically methylated regions (Irizarry et al.
2009), these genes are significantly enriched in embryonic de-
velopment, including the highly divergent homeobox gene HDX
along with several neuronal growth factors NEGR1, NETO1, and
NTM. In addition, we observe significant enrichment for zinc
finger genes, Kruppel-associated box (KRAB) transcription factors,
and DNA binding proteins (Dennis et al. 2003; Fig. 2C). This en-
richment for transcription factors indicates a drastic alteration of
the HCC1954 regulatory program from HMEC concordant with
hypomethylation.
Formation of repressive chromatin domains
at hypomethylated genomic regions
To explore the molecular processes potentially responsible for
lower transcript abundance of genes in the hypomethylated do-
mains, we examined the status of DNA methylation at promoters
of the genes. Among the 627 gene-body hypomethylated genes
with lower transcript abundance in HCC1954, promoters of 289
(46%) genes are hypermethylated, consistent with a role for pro-
moter DNA hypermethylation in transcriptional repression. These
genes include many well-known tumor suppressors such as MGMT
(Esteller et al. 1999; Everhard et al. 2009; Hibi et al. 2009a, b), DCC
(deleted in colorectal carcinoma) (Fearon et al. 1990), and DLC1
(deleted in liver cancer 1) (Yuan et al. 1998).
Surprisingly, the remaining 54% repressed genes (N = 338)
exhibit no change (N = 70) or loss (N = 268) of DNA methylation at
promoters, yet still display lower transcript abundance in cancer
cells. These genes also include well-known tumor suppressor
genes, including the interferon-inducible gene PYHIN1 (Ding et al.
2006), the leucine zipper gene LZTS1 (Ishii et al. 1999), the anti-
angiogenesis factors THBS2 (Streit et al. 1999) and HRG (Rolny
et al. 2011), and the cysteine protease inhibitor CST5 (Alvarez-Diaz
et al. 2009). Importantly, genes with intragenic hypomethylation
(HMEC
high
! HCC1954
PMD
) but lacking promoter hyper-
methylation are also significantly enriched in down-regulated
genes (P = 2.0 3 10
7
, hypergeometric) and significantly depleted
in up-regulated genes (P = 5.0 3 10
3
, hypergeometric), therefore
excluding the possibility that promoter hypermethylation ex-
plains these observations. We hypothesized that these genes may
be aberrantly repressed by other epigenetic mechanisms, such as
repressive chromatin modifications. To address this possibility, we
performed chromatin immunoprecipitation followed by se-
quencing (ChIP-seq) for several histone modifications including
the repressive H3K9me3 and H3K27me3 marks in HCC1954 and
the active chromatin marks H3K4me1, H3K4me3, H3K27ac, and
H3K36me3. Comparing these modifications to those of HMEC
produced by the ENCODE Consortium (Birney et al. 2007; Ernst
et al. 2011), we observed increased H3K27me3 in HCC1954 at
hypomethylated gene bodies, independent of promoter methyla-
tion status (Fig. 2D). Although H3K9me3 was not mapped in
HMECs, we also observed enrichment of this modification at both
sets of genes in HCC1954.
The results reveal that large-scale hypomethylation is corre-
lated with an increase in repressive chromatin formation. To fur-
ther verify this obser vation on a gl obal s cale, we plott ed the
enrichment of H3K9me3 and H3K27me3 relative to input
[log
2
(ChIP/input)]. The genomic loci with the strongest H3K9me3
and H3K27me3 enrichment also correspond to low mCG. Con-
versely, the greatest depletion of these repressive modifications oc-
curs at loci with high mCG for both HCC1954 and HMEC (Fig. 3A).
As seen on a chromosome-wide view (Fig. 3B), regions showing
hypomethylated DNA exhibit the greatest enrichment of H3K9me3
or H3K27me3 in HCC1954. Inspection of the DACH1 tumor sup-
pressor gene illustrates strong enrichment of these two repressive
marks in a large hypomethylated domain, which abruptly transi-
tions to a fully methylated domain coincident with loss of the re-
pressive modifications and gain of the mCG-associated histone
modification H3K36me3 (Ball et al. 2009; Lister et al. 2009) (Fig. 3C).
It has previously been observed that regions of hypo-
methylation are biased toward gene poor regions (Aran et al. 2011).
We also observe a small but positive correlation between %mCG
with promoters (R = 0.03), exons (R = 0.17), and gene bodies (R =
0.26) (Supplemental Fig. S3). However, the magnitude of correla-
tion between %mCG with H3K9me3 (R = 0.66) and H3K27me3
(R = 0.53) is noticeably greater, suggesting a stronger link be-
tween hypomethylation with chromatin than with genic features.
Allelic basis of repressive chromatin and DNA methylation
The above observations led us to hypothesize that formation of
H3K9me3 and H3K27me3 domains are closely coupled to the loss
of DNA methylation in HCC1954 cells. This hypothesis is sup-
ported by several global analyses: (1) For HCC1954, H3K9me3/
H3K27me3 enrichment increases as mCG decreases (Fig. 3D,E;
Supplemental Fig. S4); (2) 10-kb windows gaining the most mCG
also lose the most H3K9me3 and H3K27me3, and vice versa (Fig.
3F,G); and (3) 10-kb windows gaining repressive chromatin also
tend to lose mCG, and vice versa (Fig. 3H,I).
However, it has been observed that loci exhibiting partial
methylation overlap with H3K27me3, as previously reported in
IMR90 fetal lung fibroblasts (Lister et al. 2009). To resolve this
apparent contradiction, we tested the possibility that PMDs in
HCC1954 coinciding with repressive chromatin may have one
allele in a fully methylated state and another allele marked with
repressive chromatin.
We first distinguished the different alleles in HCC1954 by
identifying the haplotype blocks in the genome of these cancer
cells. We obtained 1,000,880,493 paired-end, nonclonal, uniquely
mapped reads corresponding to a 27.6-fold genome coverage
(Supplemental Fig. S5). Using the bam2mpg genotyping program
(Teer et al. 2010), we found 1,211,258 high-confidence single nu-
cleotide polymorphisms (SNPs) in the genome. Together with the
sequenced reads to link SNPs together, we employed the previously
developed error-correcting algorithms HASH and HapCUT to
identify 269,392 phased haplotype blocks (Bansal and Bafna 2008;
Bansal et al. 2008) with an N50 of 290 bp (Fig. 4A).
To assess if PMDs consist of one fully methylated allele and
another fully unmethylated allele, we focused on the 15,309 par-
tially methylated haplotype blocks with an allele-combined aver-
age %mCG between 40% and 60% (Fig. 4B). Plotting the %mCG of
each phased allele illustrates a clear bias between the methylation
status of each allele (Fig. 4B). Almost half (47%) of these partially
methylated haplotype blocks exhibit significant allelic bias in DNA
methylation (P # 0.05, Fisher’s exact test), with the majority (60%)
having at least a 50% difference between %mCG on the two alleles
(Fig. 4C). These results suggest that allele-specific methylation is
prevalent and that PMDs frequently consist of two differentially
methylated alleles.
To examine the status of histone modifications at the hap-
lotype blocks identified above, we sequenced between 76.5 and
92.3 million ChIP-seq reads for H3K9me3, H3K27me3, and
DNA hypomethylation linkedtochromatinsilencing
Genome Research 249
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
H3K36me3 in HCC1954 cells to gain adequate coverage and ex-
amined the allelic bias among the haplotype blocks that display
allelic DNA methylation. By distinguishing the ChIP-seq reads
corresponding to different haplotype blocks, we first identified
2100, 2479, and 2104 blocks that exhibit significant allelic bias for
H3K9me3, H3K27me3, and H3K36me3, respectively (Fig. 4A) (P #
0.05, Fisher’s exact test). Overlapping these blocks with those
showing allele-specific DNA methylation, we observe that 78% of
overlapping H3K36me3 blocks are on the same allele as mCG
(p
H3K36me3
= 1.38 3 10
17
, binomial). For example, ubiquitously
expressed MAGED2 on chrX is located within a PMD, and DNA
methylation is on the same allele as H3K36me3 (Fig. 4E). In con-
trast, the majority of overlapping H3K9me3 (75%) and H3K27me3
(79%) blocks are on the opposite allele from mCG (Fig. 4D)
(p
H3K9me3
= 1.75 3 10
12
,p
H3K27me3
= 2.11 3 10
15
, binomial). For
example, the MGMT tumor suppressor gene is partially methylated
in HCC1954 and marked by H3K27me3, with these modifications
belonging on opposite alleles (Fig. 4F). Similarly, a partially meth-
ylated haplotype block near the CADM1 tumor suppressor gene is
marked by H3K9me3 on a different allele as mCG (Fig. 4G).
To further validate the mutual exclusiveness of H3K9me3 and
H3K27me3 with mCG, we performed ChIP with these chromatin
modifications, then determined the DNA methylation status of
the resulting DNA by bisulfite sequencing (ChIP-methylC-seq). Of
the 13,494 and 17,805 10-kb windows exhibiting significant dif-
ferences in DNA methylation status between H3K9me3 and
H3K27me3 ChIP-methylC-seq compared to methylC-seq reads
(Fisher’s exact test, P # 0.01), 87.0% and 84.3% have less DNA
methylation in the ChIP sample compared to the methylC-seq
sample, respectively (p
H3K9me3
< 10
100
,p
H3K27me3
< 10
100
) (Fig. 5).
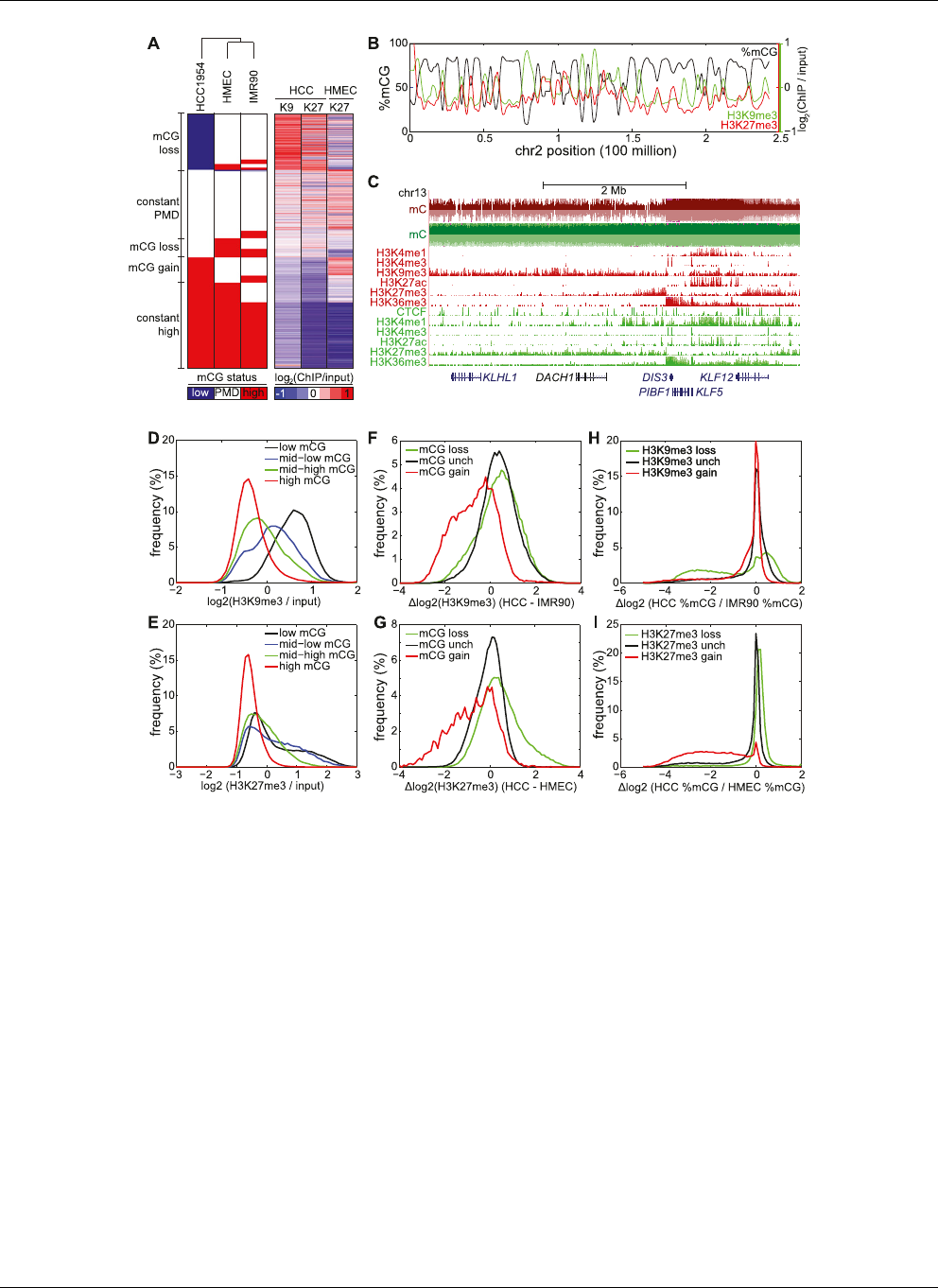
Figure 3. Repressive chromatin is depleted of mCG. (A)(Left) A reproduction of Figure 1D; (right) each of the 282,109 rows represents the H3K9me3
(denoted K9) and H3K27me3 (denoted K27) status in HCC1954 and HMEC, in the same order as the left panel. (B) Distribution of %mCG, H3K9me3, and
H3K27me3 on chromosome 2 for HCC1954. (C ) A large domain of DNA hypomethylation near the DACH1 tumor suppressor coincides with H3K9me3
and H3K27me3 in HCC1954. (Red) HCC1954, (green) HMEC. (D) Distribution of H3K9me3 enrichment for four quantiles of DNA methylation status in
HCC1954. (E)AsinD, but for H3K27me3. (F ) Distribution of change in H3K9me3 in HCC1954 compared with IMR90 for 10-kb windows that lose mCG,
gain mCG, or are unchanged for mCG. Dlog
2
(H3K9me3) = log
2
(HCC1954 H3K9me3/input) log
2
(IMR90 H3K9me3/input). (G)AsinF, but for
H3K27me3 comparing HCC1954 with HMEC. (H) Distribution of change in mCG in HCC1954 compared with IMR90 for 10-kb windows that lose
H3K9me3, gain H3K9me3, or are unchanged for H3K9me 3. (I)AsinH, but for H3K27me3 comparing HCC1954 with HMEC.
Hon et al.
250 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
In contrast, most (63.2%) of the 20,585 10-kb bins with differ-
ences in H3K36me3 ChIP-methylC-seq versus background are
more enriched for mCG (p
H3K36me3
< 10
100
)(Fig.5).Thus,in
support of our observations, DNA en-
riched for H3K9me3 or H3K27me3 is
depleted of mCG compared to DNA
enriched for H3K36me3.
A higher-order organization
of H3K9me3 and H3K27me3
The mutually exclusive nature of
H3K9me3 and H3K27me3 with mCG is
one example of the organization of the
HCC1954 epigenome. While H3K9me3
and H3K27me3 rarely overlapped (Sup-
plemental Fig. S6), we also observed
frequent examples of a higher-order
organizati on between H3K9me3 and
H3K27me3: These marks appear to be
spatially organized such that large do-
mains of H3K9me3 are flanked by regions
of H3K27me3 enrichment. Examples in-
clude the DCC and DLC1 tumor sup-
pressor genes (Fig. 6A,B).
To assess if this is a global phe-
nomenon, we defined large domains of
H3K9me3 by merging H3K9me3-enriched
10-kb windows, resulting in 376 domains
spanning 589 Mb. To exclude H3K9me3
domains having internal enrichment of
H3K27me3, we removed those domains
with an average (H3K27me3 RPKM)/(input
RPKM) $ 1.5, resulting in a final list of
322 H3K9me3 domains spanning 536 Mb
of the human genome. We observed en-
richment of H3K9me3 (Fig. 6C), de-
pletion of mCG (Fig. 6E), and background
levels of H3K27me3 within the bodies of
these H3K9me3 domains (Fig. 6D). How-
ever, H3K27me3 was enriched in re-
gions directly flanking the boundaries of
H3K9me3 domains (Fig. 6D). Similar re-
sults are observed in IMR90 H3K9me3
domains, though the extent of H3K27me3
enrichment at H3K9me3 flanks is weaker
(Fig. 6C–F). As H3K9me3 was not mapped
in HMEC, we cannot assess this phenom-
enon directly in these cells. However, to
identify potential H3K9me3 domains, we
searched for regions depleted of DNA
methylation that are simultaneously de-
pleted of H3K27me3. These large do-
mains, putatively similar to the H3K9me3
enriched domains found in HCC1956,
also exhibit flanking peaks of H3K27me3
in HMEC, as in HCC1954 and IMR90
(Supplemental Fig. S7), suggesting that
H3K27me3 flanking of H3K9me3 may be
acommonphenomenon.
A passive model of global hypomethylation
Thus far, we have focused on gross changes of DNA methylation
across large 10-kb bins. To gain insight into which genomic fea-
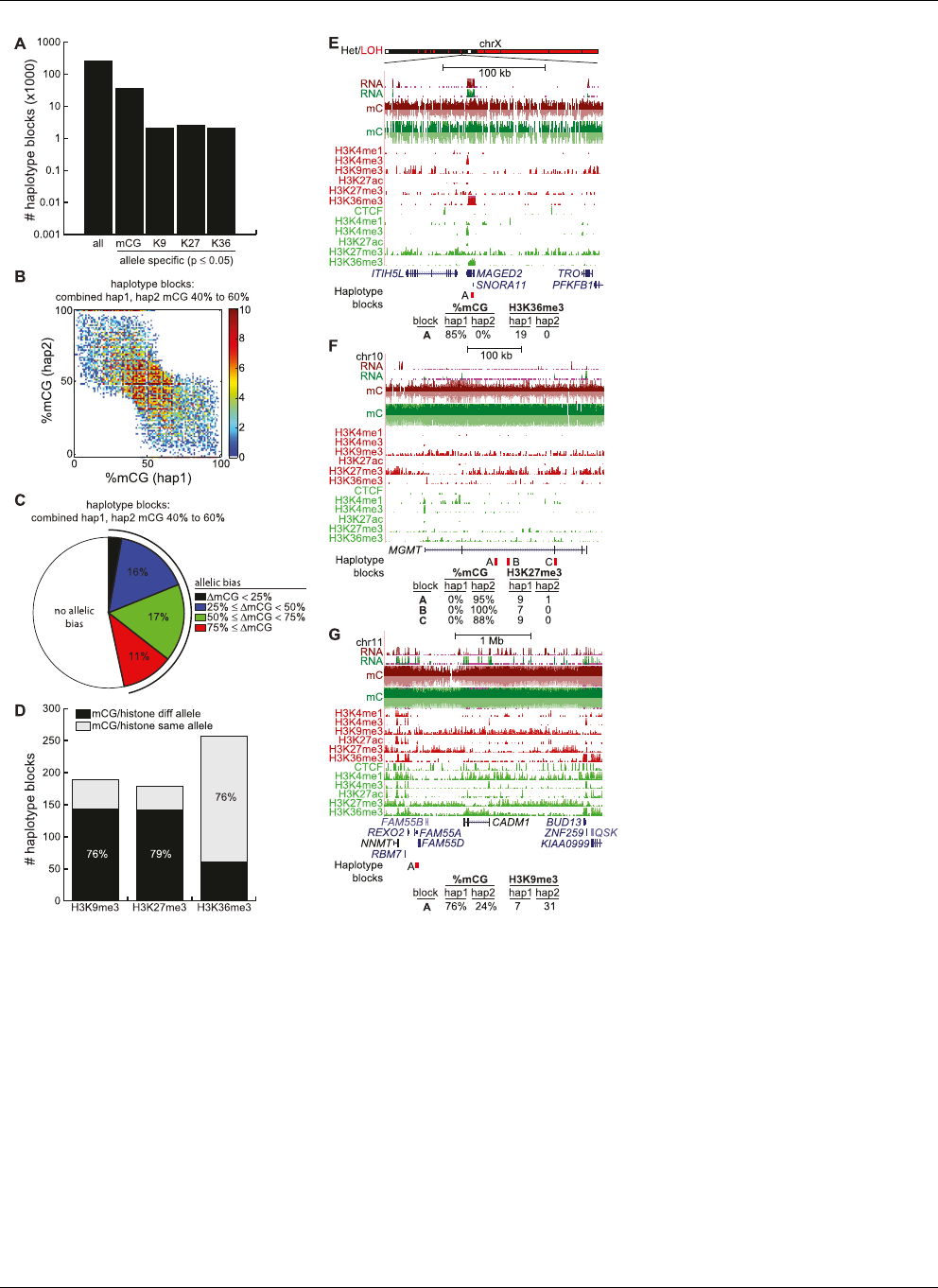
Figure 4. Allelic distribution of epigenetic modifications. (A) Number of haplotype blocks found by
SNP phasing (denoted all), and the number of these blocks showing significant allelic bias for mCG,
H3K9me3 (denoted K9), H3K27me3 (denoted K27), and H3K36me3 (denoted K36) (Fisher’s exact test
P-value # 0.05). (B) For haplotype blocks where the combined frequency of methylation of both
haplotypes is between 40% and 60%, shown is the density plot of %mCG on haplotype 1 versus %mCG
on haplotype 2. (C ) For haplotype blocks in B, shown is the fraction with allelic bias (Fisher’s exact test P-
value # 0.05), for different possible values of allelic bias. DmCG = |%mCG(hap1) %mCG(hap2)|. (D)
Of haplotype blocks simultaneously showing allelic bias in mCG (Fisher’s exact test P-value # 0.05,
DmCG $ 50%) and a histone modification (Fisher’s exact test P-value # 0.05), the number of haplotype
blocks where mCG is on the same or different allele as the histone modification. (E) Allelic H3K36me3 at
the MAGED2 gene. The bar at top indicates where one arm of chrX was lost. (Red) HCC1954, (green)
HMEC. (Bottom) Number of H3K36me3 reads belonging to haplotype 1 or 2. (F ) Allelic H3K27me3 at
the MGMT tumor suppressor. (Red) HCC1954, (green) HMEC. (Bottom) Number of H3K27me3 reads
belonging to haplotype 1 or 2. (G) Allelic H3K9me3 near the CADM1 tumor suppressor. (Red) HCC1954,
(green) HMEC. (Bottom) Number of H3K9me3 reads belonging to haplotype 1 or 2.
DNA hypomethylation linkedtochromatinsilencing
Genome Research 251
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

tures are more likely to undergo hypomethylation, we examined
the methylation level at single cytosine residues spanning exons,
intragenic regions, or intergenic regions. The distribution of cy-
tosine methylation frequency at exons in HCC1954 is nearly in-
distinguishable from HMEC (Fig. 7A). At genic regions, defined to
be the entire interval between transcription start site (TSS) and
transcription terminal site (TTS), the frequency of lowly methyl-
ated cytosines is higher in HCC1954 than HMEC (Fig. 7A) (20.6%
vs 14.5%, P < 10
16
, binomial). However, hypomethylation is most
pronounced in intergenic regions, where 30.2% of HCC1954 cy-
tosines are lowly methylated compared to 16.2% in HMEC (Fig.
7A) (P < 10
16
, binomial).
Two potential mechanisms can be envisioned to account for
the global hypomethylation in cancer cells. One possibility is that
the methyl group is actively removed from methylated cytosines.
Several proteins, including AICDA (also known as AID), GADD45A,
Tet family of proteins, ELP3, and TDG, have recently been impli-
cated in demethylation of methylated cytosines (Barreto et al. 2007;
Bhutani et al. 2010; Ito et al. 2010; Cortellino et al. 2011). None of
these genes is significantly more expressed in HCC1943 than
HMEC, nor do they harbor somatic mutations in HCC1954, sug-
gesting that an active mechanism may not be responsible for global
hypomethylation in HCC1954. Alternatively, DNA methylation
may be gradually lost during replication of the cancer cells, which
would be biased toward late-replicating regions of the genome. To
test whether our data support this model, we compared the DNA
methylation levels in HCC1954 to a compilation of regions that are
consistently early-, middle-, and late- replicating in a panel of four
cell lines (hESC, erythroid, lymphoid, and fibroblast cells) (Hansen
et al. 2009). We find that 58.2% of consistently late-replicating
regions have low methylation in HCC1954, significantly more
than that expected by chance (22.3%, P < 10
16
, binomial). In
contrast, only 1.2% and 4.0% of consistently early- and middle-
replicating regions, respectively, are lowly methylated. These re-
sults are consistent with a previous report (Aran et al. 2011) and the
model that DNA methylation at late-replicating regions is gradu-
ally lost through many rounds of cell divisions. Further experi-
ments are necessary to understand how late-replication timing
leads to partial loss of DNA methylation.
DNA found near lamina-associated domains (LADs) has pre-
viously been shown to be late-replicating (Moir et al. 1994). As
LADs have been shown to be stable across different cell types
(Peric-Hupkes et al. 2010), to provide further support that hypo-
methylated domains in HCC1954 are late-replicating, we com-
pared them to a previously published map of LADs in Tig3 human
fibroblast cells (Guelen et al. 2008). We observe remarkable con-
cordance between LADs in Tig3 cells and hypomethylated do-
mains in HCC1954 (Fig. 7C). On a global scale, 37.2% of LADs are
found in hypomethylated domains, compared to 22.3% expected
by chance (P < 10
16
, binomial), consistent with previous obser-
vations (Hansen et al. 2011).
Assessing the epigenetic contribution to increased
transcription of repeat elements
Increased expression of repetitive elements is a hallmark of cancer
cells, and this aspect of cancer has frequently been attributed
to global hypomethylation (Prak and Kazazian 2000). To assess
the frequency of hypomethylation at repeats, we examined the
methylation level at each cytosine residue contained within re-
peats. Dramatically, 21.2% of repeat-associated cytosines are lowly
methylated in HCC1954, compared to 8.7% in HMEC (P < 10
16
,
binomial) (Fig. 8A). Lowly methylated repeats are also more fre-
quently found in intergenic regions than expected by chance:
while 55.2% of all repeat-associated cytosines are intergenic, the
subset consisting of lowly methylated residues is 69.2% intergenic
(P < 10
16
, binomial) (Fig. 8B).
To study the expression of repetitive elements, we focused on
intergenic repeats, thereby avoiding ambiguities with genic fea-
tures. The med ian expression level o f intergenic repeats in
HCC1954 (0.43 RPKM) is 5.2 times more than in HMEC (0.08
RPKM) (P < 10
16
, rank sum test) (Fig. 8C), supporting previous
observations of extensive repeat expression in cancer cells. Remi-
niscent of chromatin signatures at actively transcribed genes, we
also observe hypomethylation at repeats concordant with peaks of
H3K4me3 and H3K27ac (Fig. 8D,E). Strand-specific expression
emanates from these peaks and coincides with enrichment of
H3K36me3, extending to contiguous blocks of transcription span-
ning ;31 kb and ;7kbnearSERINC2 and PLA2G2F,respectively.
Notably, these large blocks of repeat transcr iption sp an nu-
merous adjacent repeat sequences and lack hypomethylation.
These observations suggest several explanations for repeat ex-
pression: (1) loss of a repressive epigenetic mark, (2) gain of an active
chromatin mark, or (3) read-through transcription of neighboring
repeats. To further explore these three possibilities, we quantified
how much each component explains the expression of a set of 4592
intergenic repeats highly (RPKM
HCC1954
$ 1) and specifically
(RPKM
HCC1954
$ 10 3 RPKM
HMEC
) expressed in HCC1954. About
one-third (34.2%) of these repeats exhibited at least a 50% change in
an epigenetic modification. Of these, loss of DNA methylation is
the most prominent epigenetic change, accounting for 62.7%. Gain
of active chromatin modifications accounts for 42.1% of these re-
peats, while loss of H3K27me3 only makes up 8.7% (Fig. 8F).
To explain repeat expression by read-through of neighboring
repeats, we first identified 6354 intergenic domains containing
clustered and strand-specific RNA-seq reads (see Methods). While
these domains only span 5.4% of all intergenic regions, they ac-
count for 31.0% of all repeats specifically expressed in HCC1954 (P <
10
16
, binomial). As further evidence that these repeats are the re-
sult of read-through transcription, 94.0% are oriented in the domi-
nant direction of RNA-seq reads in the domains, while the expectation
is only 50% (P < 10
16
, binomial). In total, these oriented repeats
account for 29.2% of all HCC1954-specific repeats, and altogether
about half (52.3%) of all repeats could be explained by either epi-
genetic changes or read-through repeat transcription (Fig. 8F).
Figure 5. Mutual exclusiveness of chromatin modifications and mCG
by ChIP-methylC-seq. ChIP was performed for H3K9me3, H3K27me3, or
H3K36me3 in HCC1954, followed by bisulfite conversion and sequenc-
ing. For 10-kb windows, mCG bias was compared to genomic back-
ground from methylC-seq (P # 0.01, Fisher’s exact test). Shown is the
number of 10-kb windows showing significant differences, split by
whether there is more or less mCG in the ChIP-methylC-seq sample.
Hon et al.
252 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
Discussion
Global DNA hypomethylation is a hallmark in human cancer, but
its functional consequences have been unclear (Hinshelwood and
Clark 2008). By comparing the methylomes of a breast cancer line
HCC1954 and the primary breast cell HMEC, we find a link be-
tween hypomethylation and the formation of repressive chroma-
tin domains. Affected genes are frequently repressed in cancer cells
and include tumor suppressors such as DNA repair gene MGMT
(Esteller et al. 1999), the deleted in colorectal carcinoma gene, DCC
(Fearon et al. 1990), and the deleted in liver cancer gene, DLC1
(Yuan et al. 1998). Thus, hypomethylation is not always an acti-
vating epigenetic change as previously assumed (Esteller et al.
2000; Esteller 2007, 2008). Rather, when it is accompanied
by a gain of repressive chromatin, the result can be a decrease in
expression.
The mechanism of global hypomethylation is a long-standing
question in cancer epigenetics. Our data seem to support a passive
mechanism whereby methylation is gradually lost over successive
cell divisions, as opposed to an active mechanism involving
demethylating enzymes (Wild and Flanagan 2010). Evidence for
this passive mechanism includes a bias for hypomethylation at
intergenic, late-replicating, and lamina-associated regions of the
genome. Furthermore, given that global hypomethylation is a
consistent feature of many diverse cancers (Gama-Sosa et al. 1983),
the alternative hypothesis involving activation of demethylating
enzymes appears less plausible. In light of the abnormal growth
typical of cancer, a passive mechanism is more intuiti ve: Cancer
cells grow faster than methylation can be copied from the rep-
licating parental DNA, resulting in progressive loss of DNA
methylation.
Increased expression of repetitive DNA sequences has been
frequently observed in cancer cells, and this has often been linked
to changes in DNA methylation or chromatin structure (Florl et al.
1999; Menendez et al. 2004; Schulz 2006; Ting et al. 2011). By
integrating the transcriptome and epigenome data sets, our anal-
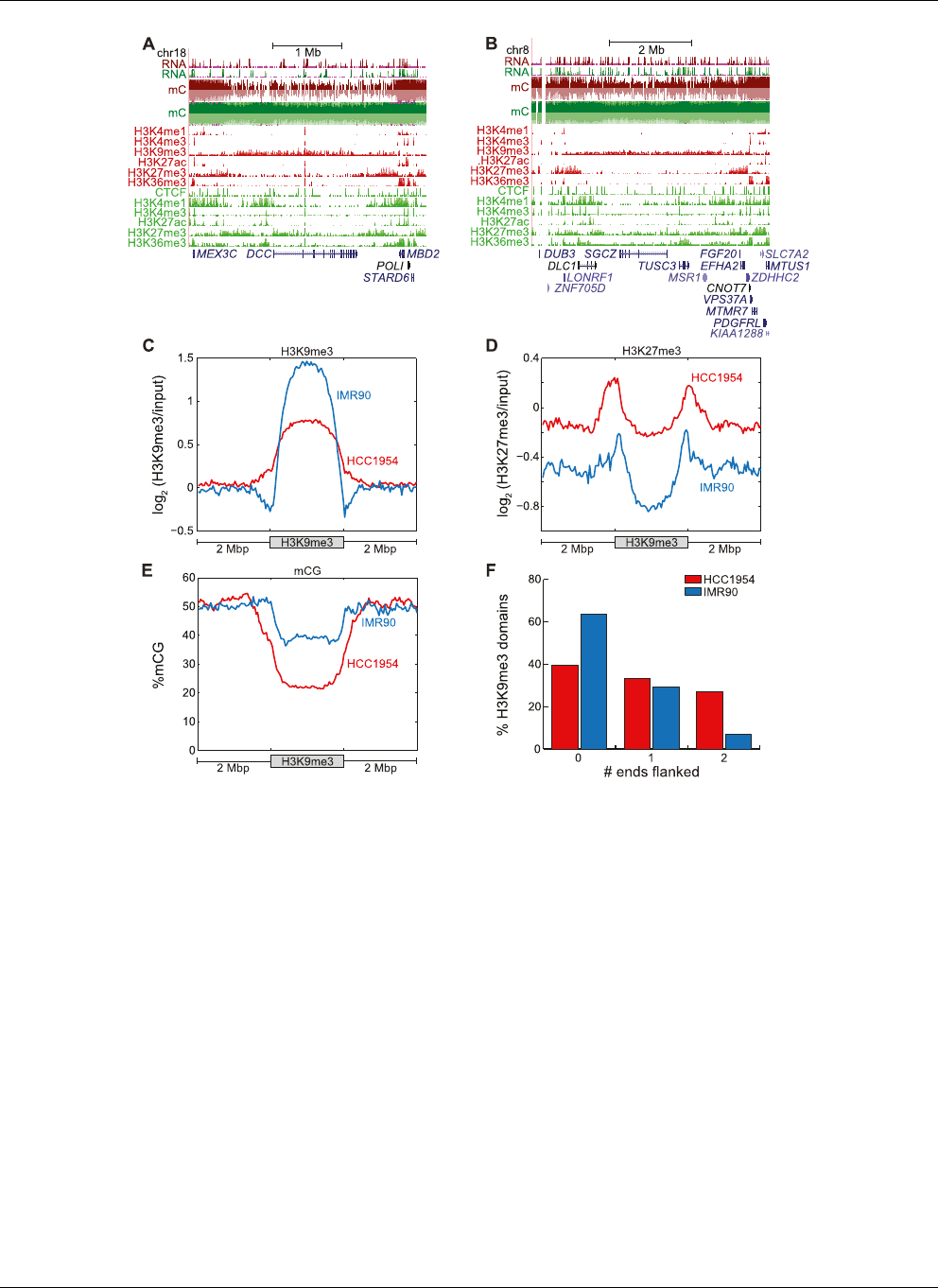
Figure 6. Large-scale organizatio n of H3K9me3 and H3K27me3. (A) Hypomethylation near the DCC tumor suppressor coincides with H3K9me3, which
is flanked by H3K27me3. (B) Hypomethylation near the DLC1 tumor suppressor coincides with H3K9me3, which is flanked by H3K27me3. (C ) Enrichment
of H3K9me3 within 2 Mb of H3K9me3 domains in HCC1954 and IMR90. (D) Enrichment of H3K27me3 within 2 Mb of H3K9me3 domains in HCC1954
and IMR90. (E) Enrichment of mCG within 2 Mb of H3K9me3 domains in HCC1954 and IMR90. (F ) Fraction of H3K9me3 domains flanked by 0, 1, or 2
H3K27me3 domains.
DNA hypomethylation linkedtochromatinsilencing
Genome Research 253
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
ysis suggests several possible mechanisms of repeat expression.
Epigenetics can explain the expression of about one-third of
HCC1954-specifically expressed repeats. Additionally, transcrip-
tion read-through may be involved, as we observe that spatially
clustered repeats are often expressed as a single transcriptional
unit. One end of this unit is marked with the canonical epige-
netic marks of transcription initiation (H3K4me3, H3K27ac, and
hypomethylation), and the transcribed body with numerous re-
petitive elements contains the elongation mark H3K36me3. These
repeats are potentially transcribed as a result of read-through
transcription and span nearly a third of HCC1954-specifically
expressed repetitive elements.
Partial DNA methylation is a recent observation made possi-
ble by whole genome shotgun bisulfite sequencing (Lister et al.
2009). Since PMDs are not observed in pluripotent stem cells but
span about one-third of the genome in differentiated cell types
such as fetal lung fibroblasts, the gradual loss of DNA methylation
in PMDs during development may be interpreted as another form
of global hypomethylation. However, the biological relevance and
formative processes of PMDs has yet to be understood. We show
that a significant fraction of PMDs correspond to allele-specifically
methylated DNA. In these regions, the unmethylated allele is oc-
cupied by H3K9me3 and H3K27me3, while the hypermethylated
allele is devoid of the repressive chromatin marks. This argues
for a mutually exclusive relationship between DNA methylation
and H3K9me3 or H3K27me3 at PMDs and is also in agreement
with previous observations that highly methylated ES cells have
much lower levels of H3K9me3 and H3K27me3 compared to dif-
ferentiated cells (Lister et al. 2009; Hawkins et al. 2010). This ex-
clusivity of H3K9me3 and DNA methylation is not incompatible
with previous studies showing a close mechanistic tie between
H3K9me3 and DNA methylation, particularly at constitutive het-
erochromatin, which is generally centro-
meric or telomeric (Schotta et al. 2004;
Volkel and Angrand 2007). For example,
H3K9me3 directs DNMT3B-dependent
DNA methylation at pericentric satellite
repeats in mouse ES cells (Lehnertz et al.
2003). As short-read sequencing tech-
nologies cannot map to these highly
repetitive regions of the genome, our
analyses are restricted to regions of fac-
ultative heterochromatin and euchro-
matin. Thus, at least in HCC1954 and
IMR90, H3K9me3 appears to be exclusive
of DNA methylation outside of constitu-
tive heterochromatin.
Our observations suggest that DNA
methylation changes in breast cancer
cells may be mechanistically linked to
the pathways responsible for H3K9me3
or H3K27me3. Interestingly, the genes
encoding EZH2 and other PcG proteins
responsible for H3K27me3 are frequently
overexpressed in breast cancers (Kleer
et al. 2003; Chang et al. 2011), and point
mutations that lead to loss or gain of
function have also been reported in these
genes (Dalgliesh et al. 2010; Morin et al.
2010; Yap et al. 2010). The exclusive na-
ture of repression, together with the ob-
servation that hypomethylation is co-
incident with an increase in repressive chromatin, suggests that
these repressive mechanisms may be compensatory for each other
and that therapies targeting one modification may not be suffi-
cient to acquire gene activation.
Given the mutual exclusion of repressive epigenetic modi-
fications, an unresolved question is whether global hypo-
methylation in cancer results in gain of H3K9me3/H3K27me3, or
the other way around. Recently, it has been shown that DNA
methylation prevents binding of the PRC2 complex to chromatin
to deposit H3K27me3 (Lindroth et al. 2008; Bartke et al. 2010; Wu
et al. 2010). Further, Komashko and Farnham observed that dis-
ruption of DNA methylation by 5-azacytidine in human embry-
onic kidney cells results in global increases of H3K9me3 and
H3K27me3 (Komashko and Farnham 2010). Taken together, these
data suggest that loss of DNA methylation in cancer cells may lead
to the formation of repressive chromatin domains and silencing of
tumor suppressor genes. We postulate, therefore, that inhibition of
the H3K27me3 or H3K9me3 methyltransferases could be a new
cancer therapeutic strategy. Further research is needed to identify
the causes of DNA hypomethylation in these cells.
Methods
Cell culture
HCC1954 cells were grown in RPMI 1640 media supplemented
with 10% FBS, 13 nonessential amino acids (Invitrogen 11140050),
and 13 L-glutamine (Invitrogen 45000-676-1). Cells submitted
to epigenetic analysis were between passage 34 and 42. Cryo-
preserved HMECs at pass age 6–7 were purchased from Lonz a
(CC-2551) and grown according to the manufacturer’s instructions
at 37°C/5% CO
2
. The cells were split two times before harvesting.
Harvested HMECs highly express p16/CDKN2A, suggesting these
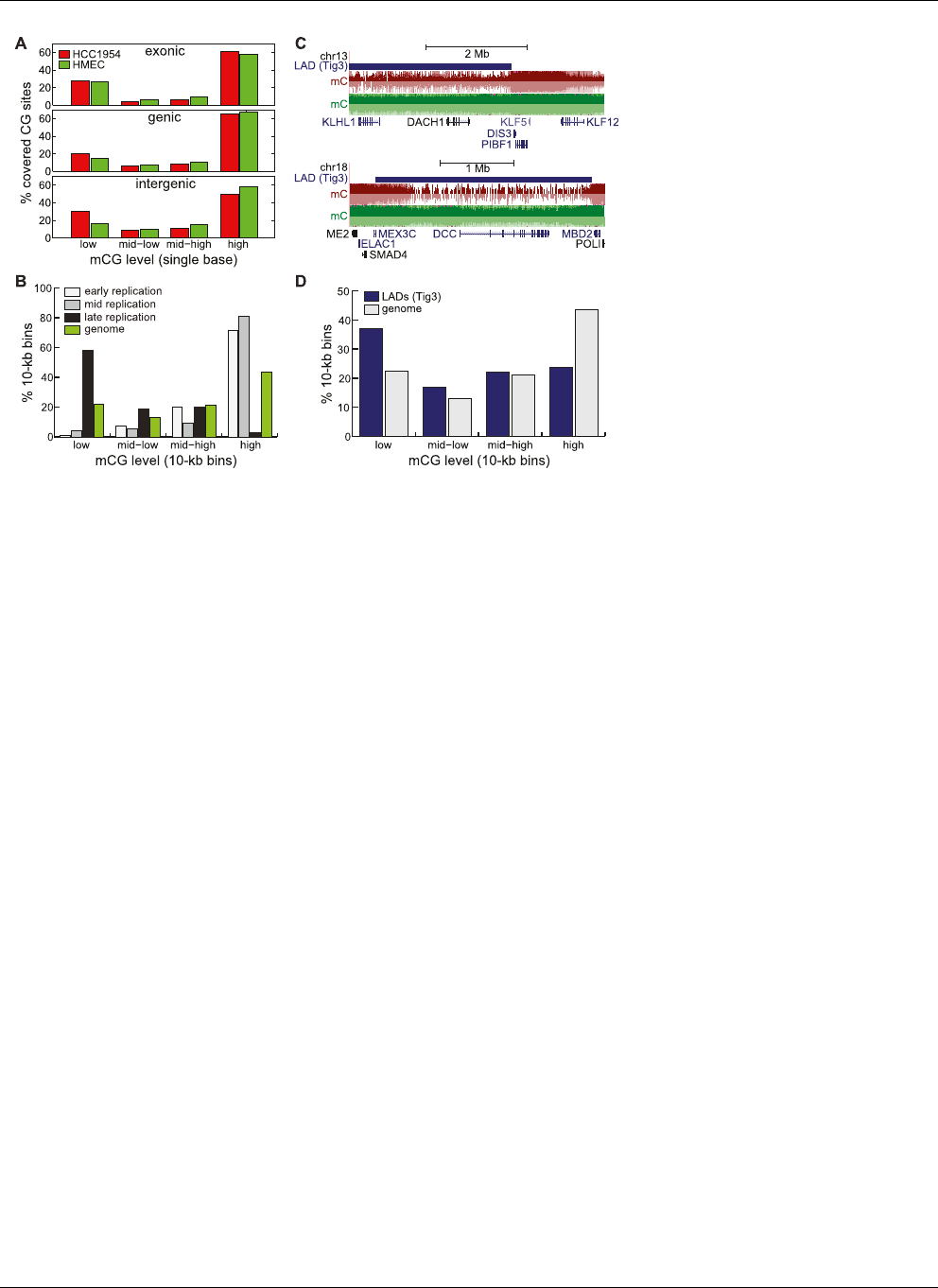
Figure 7. A passive model of hypomethylation. (A) Distribution of HCC1954 %mCG for exonic (top),
genic (middle), and intergenic (bottom) cytosine residues. (B) Distribution of HCC1954 %mCG for 10-kb
regions that are consistently early-, middle-, and late-replicating in four cell types (Hansen et al. 2009),
compared to the background genome (green). (C ) Snapshots illustrating the overlap of hypomethy-
lated regions with lamina-associated domains previously mapped in Tig3 cells (Guelen et al. 2008), at
the DACH1 (top)andDCC (bottom) genes. (D) Distribution of HCC1954 %mCG for 10-kb regions that
are found in Tig3 lamina-associated domains (blue), compared to the background genome (gray).
Hon et al.
254 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
cells have not reached stasis (Novak et al. 2008). Harvested HMECs
express KLK6, COX7A1, EPCAM, KRT19, and PRDM1, all of which
are hallmarks of early passage prestasis HMECs. Finally, harvested
HMECs express TP53, indicating that they have not entered
crisis (Garbe et al. 2009). See Supplemental Material for detailed
descriptions.
Sequencing
Sonicated genomic DNA was submitted to Illumina paired-end li-
brary preparation according to the manufacturer’s instructions.
Libraries for mRNA-seq (Parkhomchuk et al. 2009), ChIP-seq
(Hawkins et al. 2010), and methylC-seq (Lister et al. 2009) were
created as described previously. ChIP-seq was performed with an-
tibodies specific to H3K4me1, H3K4me3, H3K9me3, H3K27ac,
H3K27me3, and H3K36me3. RNA-seq reads were mapped with
TopHat (Trapnell et al. 2009), and other
experiments were mapped with Bowtie
(Langmead et al. 2009). Sequenced reads
for RNA-seq and ChIP-seq were 36 bp,
with methylC-seq at 100 bp, and genome
sequencing being a mix of 36, 80, and 101
bp. See Supplemental Material for com-
plete protocols.
Antibodies
ChIP-seq was performed using the fol-
lowing antibodies: H3K4me1 (Abcam
ab8895-50, lot 720417), H3K4me3
(Millipore CS200580, lot DAM1612220),
H3K9me3 (Abcam ab8898-100, lot
699671; Diagenode pAb-056-50, lot A93-
0041), H3K27a c (Active Motif 39133, lot
19208002), H3K27me3 (Millipore 07-
449, lot DAM161288), and H3K36me3
(Abcam ab9050-100, 707946). All anti-
bodies used here were validated by pep-
tide dot blot assays to ensure specific-
ity to the correct histone modification
(Egelhofer et al. 2011).
Read mapping
Reads from genome sequencing were
mapped by the Bowtie program in two
passes: first as paired-end mapp ing and
then as single-end mapping of unmapped
reads. ChIP-seq reads were also mapped
by Bowtie. For both genome sequencing
and ChIP-seq, onl y reads that mapped
uniquely to hg18 wi th, at most, three
mismatches were kept. MethylC-seq reads
were mapped as previously described
(Lister et a l. 2009). RNA-se q reads were
mapped by the TopHat program. For non-
RNA libraries, PCR duplicates were re-
moved with the Picard program (Picard
2011). All biological replicates were com-
bined a nd compresse d/indexed by t he
SAMtools suite into BAM format (Li et al.
2009). See Supplemental Material for de-
tailed descriptions.
Quantifying RNA-seq expression
To quantify RPKM expression (reads per kilobase of model per
million base pairs sequenced) at UCSC Known Genes (Hsu et al.
2006) and RefSeq genes (Pruitt et al. 2005), the Cufflinks program
(Trapnell et al. 2010) was applied to the mapped TopHat reads. To
allow for division between RPKM values, the lowest RPKM value
was set to 5 3 10
6
. To reduce redundancy of the gene lists, genes
were merged and their expression values summed if (1) they had
the same common gene name with the same TSS, or (2) they had
the same common gene name and overlap each other. A gene is
a member of a 10-kb DNA methylation domain category (low/mid-
low/mid-high/high) if there is any overlap of the gene body with
a member of the methylation category. Therefore, a gene can be-
long to multiple categories. To quantify expression of repetitive
elements, we counted the number of reads mapping in each repeat
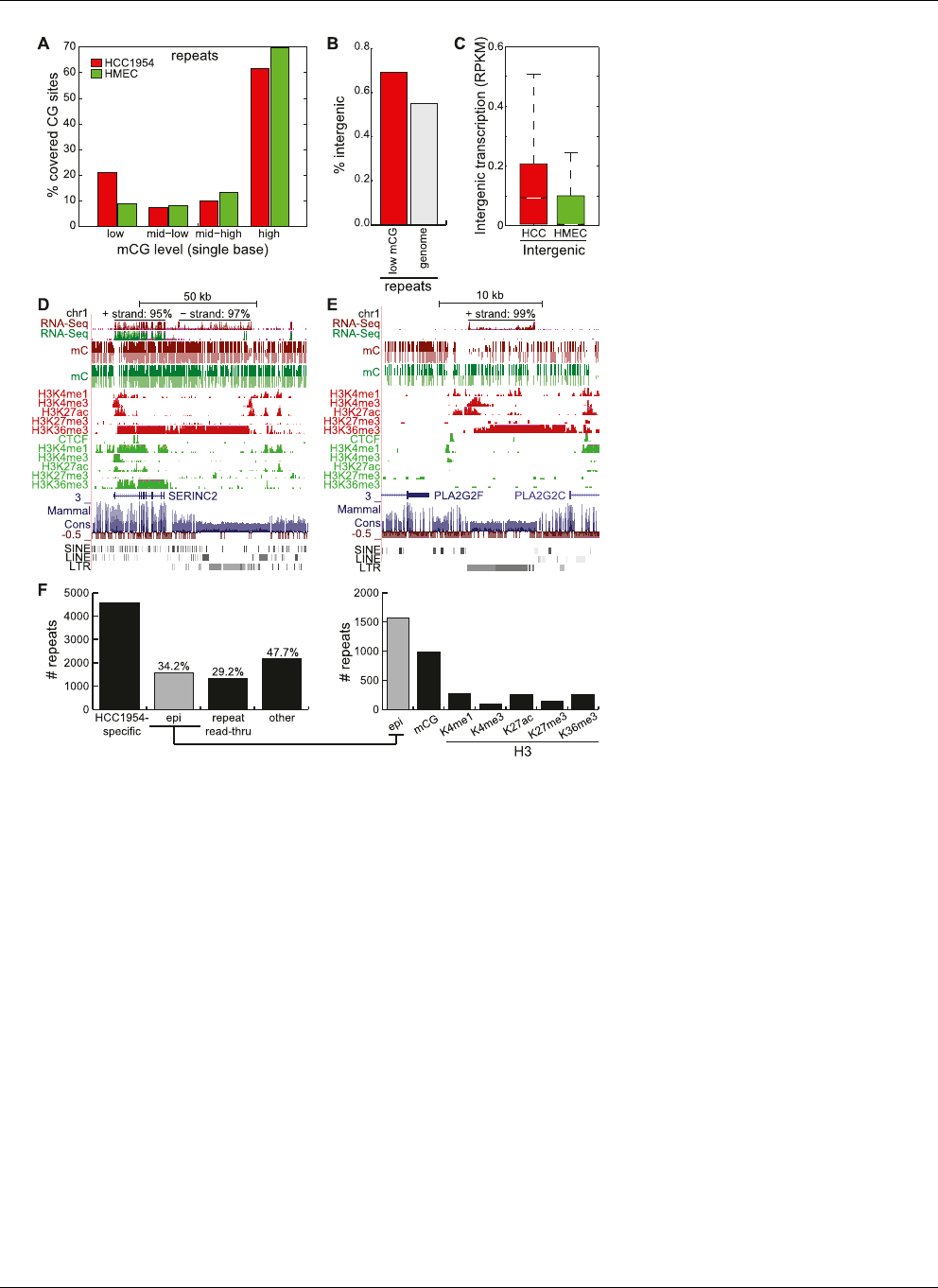
Figure 8. Epigenetic contribution of increased repeat expression. (A) Distribution of %mCG of each
repeat-associated cytosine residue spanned by at least 10 methylC-seq reads. (B) Percentage of all re-
peats (gray) and of lowly methylated repeats in HCC1954 (red) that are intergenic. (C ) Distribution of
intergenic repeat expression in HCC1954 and HMEC, expressed in RPKM. (White line) median, (box)
25th to 75th percentiles. (D) Snapshot of repeat expression and read-through with epigenetic signa-
tures near the SERINC2 gene. RNA-seq strand distribution shown on top.(E) Snapshot of repeat ex-
pression and read-through with epigenetic signatures near the PLA2G2F gene. RNA-seq strand
distribution shown on top.(F ) Number of highly expressed repeats specific to HCC1954 and the number
that can be explained by at least a 50% change in epigenetic modifications or read-through tran-
scription from nearby repeats. (Epi) epigenetic.
DNA hypomethylation linkedtochromatinsilencing
Genome Research 255
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

and expressed the result in RPKM units, using only the subset of
RNA-seq reads that are uniquely mapping to hg18.
Quantifying ChIP-seq enrichment
Enrichment of histone modifications in a specified region of the
genome was quantified as log
2
(ChIP RPKM/input RPKM). To avoid
division by zero, a pseudocount was added depending on the
depth of sequencing: Pseudocount = (# million base pairs map-
ped)/p, where p is the pseudocount factor, here set to 2. Thus,
RPKM = (# reads in bin + pseudocount)/(# kb in model)/(# million
base pairs mapped). This method of pseudocounts guarantees that
bins having (1) no ChIP and input reads or (2) ChIP and input
reads perfectly proportional to the number of reads sequenced will
have log
2
(ChIP RPKM/input RPKM) = 0.
Quantifying mCG enrichment
DNA methylation in the CG context was quantified as %mCG of
all cytosine bases in the reference sequence. Specifically, for each of
the Watson and Crick strands, all cytosine bases in the CG context
in the reference sequence hg18 were identified. For all mapped
methylC-seq reads in a given genomic interval, the number of
cytosines in CG context that were called methylated as well as the
number called unmethylated were counted. To reduce noise from
sequencing error, only those bases with phred score $20, indicating
sequencing confidence of at least 99%, were counted. Then %mCG =
(# methylated CGs on both strands)/(# methylated CGs on both
strands + # unmethylated CGs on both strands) 3 100. This method
assures that genomic intervals spanned by many methylC-seq reads
devoid of mCG are accurately given a low %mCG value.
Defining haplotype blocks
To generate haplotypes, aligned reads were first used to determine
SNPs in HCC1954 using the genotyping program bam2mpg (Teer
et al. 2010). To safeguard against incorrect SNP calls due to se-
quencing error, only considered bases with phred score $20 were
considered. Only genotypes with score $10 were kept, assuring
that reported genotypes are at least exp(10) » 22,000 times as prob-
ableasthenextmostprobablegenotype.WithalistofSNPsandthe
sequence read information to link them together, previously de-
veloped error-correcting algorithms HASH and HapCUTwere used to
assemble haplotypes (Bansal and V afna 2008; Bansal et al. 2008).
Assessing allele-specific epigenetic modifications
To assess allele-specific ChIP-seq enrichment at a given haplotype
block, the number of ChIP reads landing in each of the two hap-
lotypes hap1 and hap2 were counted. The background consisted of
the number of occurrences of hap1 and hap2 from genome se-
quencing. Fisher’s Exact Test was used to assess allele-specificity of
ChIP-seq (P # 0.05). To assess allele-specific DNA methylation at
a given haplotype block, the number of methylated and unmeth-
ylated cytosines in CG context (given the reference genome se-
quence hg18 and with a phred score $20) were counted for hap1
and hap2. Fisher’s Exact Test was used to assess allele-specificity of
methylC-seq (P # 0.05). See Supplemental Material for detailed
descriptions.
To assess allele-specific DNA methylation from ChIP-methylC-
seq data, the number of methylated and unmethylated cytosines
in CG context (given the reference genome sequence hg18 and
with a phred score $20) were counted for each 10-kb window
using ChIP-methylC-seq and methylC-seq data, and Fisher’s Exact
test (P # 0.01) was used to assess differential methylation.
Identifying large chromatin domains
To find large domains of H3K9me3 enrichment, log
2
(ChIP RPKM/
input RPKM) as described above was calculated for all 10-kb win-
dows spanning the human genome. H3K9me3-enriched windows
are defined as having (H3K9me3 RPKM)/(input RPKM) $ 1.5 . The
binomial distribution with p = (# H3K9me3-enriched bins)/(# total
bins) was used to find all domains of size 250 kb spanning these
10-kb windows that were significantly enriched for H3K9me3-
enriched bins (P-value # 0.01). To reduce redundancy of over-
lapping 250-kb domains, domains with no more than 50 kb of
nonoverlapping regions were merged. As visual inspection in-
dicated H3K27me3 domains were smaller than H3K9me3 do-
mains, H3K27me3 domains were identified similarly but with
a domain size of 100 kb rather than 250 kb.
Quantifying repeat-associated read-through transcription
We counted the number of reads mapping in each intergenic 1-kb
bin and expressed the result in RPKM units, using only the subset
of RNA-seq reads that are uniquely mapping to hg18. To reduce
noise, we filtered for strand-specific bins containing at least three
reads and having at least five times as many reads on one strand as
on the other. To consolidate strand-specific transcribed intergenic
clusters, bins within 5 kb on the same strand were merged, and
merged regions smaller than 5 kb were discarded. Finally, repeats
associated with read-through transcription were defined as repeats
belonging to a merged region that are oriented in the same di-
rection of transcription.
Data access
All the sequencing data generated here have been submitted to the
NCBI Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.
nih.gov/geo/) under accession no. GSE29069.
Acknowledgments
This work was supported by the Ludwig Institute for Cancer Re-
search and the Mary K. Chapman Foundation (J.R.E.). Work in the
laboratory of J.R.E. is supported by the Howard Hughes Medical
Institute and the Gordon and Betty Moore foundation. J.R.E. is
a HHMI-GBMF Investigator. C.L. and V.B. were supported by an
NSF fellowship, and grants 5RO1-HG004962 (NIH), and NSF-CCF-
1115206 to V.B.
References
Alvarez-Dı
´
az S, Valle N, Garcı
´
a JM, Pen
˜
a C, Freije JM, Quesada V, Astudillo A,
Bonilla F, Lo
´
pez-Otı
´
n C, Mun
˜
oz A. 2009. Cystatin D is a candidate tumor
suppressor gene induced by vitamin D in human colon cancer cells.
J Clin Invest 119: 2343–2358.
Aran D, Toperoff G, Rosenberg M, Hellman A. 2011. Replication timing-
related and gene body-specific methylation of active human genes. Hum
Mol Genet 20: 670–680.
Ball MP, Li JB, Gao Y, Lee JH, LeProust EM, Park IH, Xie B, Daley GQ, Church
GM. 2009. Targeted and genome-scale strategies reveal gene-body
methylation signatures in human cells. Nat Biotechnol 27: 361–368.
Bansal V, Bafna V. 2008. HapCUT: An efficient and accurate algorithm for
the haplotype assembly problem. Bioinformatics 24: i153–i159.
Bansal V, Halpern AL, Axelrod N, Bafna V. 2008. An MCMC algorithm for
haplotype assembly from whole-genome sequence data. Genome Res 18:
1336–1346.
Barreto G, Schafer A, Marhold J, Stach D, Swaminathan SK, Handa V,
Doderlein G, Maltry N, Wu W, Lyko F, et al. 2007. Gadd 45a promotes
epigenetic gene activation by repair-mediated DNA demethylation.
Nature 445: 671–675.
Bartke T, Vermeulen M, Xhemalce B, Robson SC, Mann M, Kouzarides T.
2010. Nucleosome-interacting proteins regulated by DNA and histone
methylation. Cell 143: 470–484.
Hon et al.
256 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

Bhutani N, Brady JJ, Damian M, Sacco A, Corbel SY, Blau HM. 2010.
Reprogramming towards pluripotency requires AID-dependent DNA
demethylation. Nature 463: 1042–1047.
Birney E JA, Stamatoyannopoulos A, Dutta R, Guigo TR, Gingeras EH,
Margulies Z, Weng M, Snyder ET, Dermitzakis RE, Thurman et al. 2007.
Identification and analysis of functional elements in 1% of the human
genome by the ENCODE pilot project. Nature 447: 799–816.
Chang CJ, Yang JY, Xia W, Chen CT, Xie X, Chao CH, Woodward WA, Hsu
JM, Hortobagyi GN, Hung MC. 2011. EZH2 promotes expansion of
breast tumor initiating cells through activation of RAF1-b-catenin
signaling. Cancer Cell 19: 86–100.
Cortellino S, Xu J, Sannai M, Moore R, Caretti E, Cigliano A, Le Coz M,
Devarajan K, Wessels A, Sopran o D, et al. 2011. Thy mine DNA
glycosylase is essential for active DNA demethylation by linked
deamination-base excision repair. Cell 146: 67–79.
Dalgliesh GL, Furge K, Greenman C, Chen L, Bignell G, Butler A, Davies H,
Edkins S, Hardy C, Latimer C, et al. 2010. Systematic sequencing of renal
carcinoma reveals inactivation of histone modifying genes. Nature 463:
360–363.
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA.
2003. DAVID: Database for Annotation, Visualization, and Integrated
Discovery. Genome Biol 4: R60. doi: 10.1186/gb-2003-4-9-r60.
Ding Y, Lee JF, Lu H, Lee MH, Yan DH. 2006. Interferon-inducible protein
IFIXalpha1 functions as a negative regulator of HDM2. Mol Cell Biol 26:
1979–1996.
Dworkin AM, Huang TH, Toland AE. 20 09. Epigenetic alterations in the
breast: Implications for breast cancer detection, prognosis, and
treatment. Semin Cancer Biol 19: 165–171.
Egelhofer TA, Minoda A, Klugman S, Lee K, Kolasinska-Zwierz P,
Alekseyenko AA, Cheung MS, Day DS, Gadel S, Gorchakov AA, et al.
2011. An assessment of histone-modification antibody quality. Nat
Struct Mol Biol 18: 91–93.
Ernst J, Khe radpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB, Zhang
X, Wang L, Issner R, Coyne M, et al. 2011. Mapping and analysis of
chromatin state dynamics in nine human cell types. Nature 473: 43–49.
Esteller M. 2007. Cancer epigenomics: DNA methylomes and histone-
modification maps. Nat Rev Genet 8: 286–298.
Esteller M. 2008. Epigenetics in cancer. N Engl J Med 358: 1148–1159.
Esteller M, Hamilton SR, Burger PC, Baylin SB, Herman JG. 1999.
Inactivation of the DNA repair gene O6-methylguanine-DNA
methyltransferase by promoter hypermethylation is a common event in
primary human neoplasia. Cancer Res 59: 793–797.
Esteller M, Silva JM, Dominguez G, Bonilla F, Matias-Guiu X, Lerma E,
Bussaglia E, Prat J, Harkes IC, Repasky EA, et al. 2000. Promoter
hypermethylation and BRCA1 inactivation in sporadic breast and
ovarian tumors. J Natl Cancer Inst 92: 564–569.
Everhard S, Tost J, El Abdalaoui H, Criniere E, Busato F, Marie Y, Gut IG,
Sanson M, Mokhtari K, Laigle-Donadey F, et al. 2009. Identification of
regions correlating MGMT promoter methylation and gene expression
in glioblastomas. Neuro-oncol 11: 348–356.
Fearon ER, Cho KR, Nigro JM, Kern SE, Simons JW, Ruppert JM, Hamilton
SR, Preisinger AC, Thomas G, Kinzler KW, et al. 1990. Identification of
a chromosome 18q gene that is altered in colorectal cancers. Science 247:
49–56.
Feinberg AP, Vogelstein B. 1983. Hypomethylation distinguishes genes of
some human cancers from their normal counterparts. Nature 301: 89–92.
Florl AR, Lower R, Schmitz-Drager BJ, Schulz WA. 1999. DNA methylation
and expression of LINE-1 and HERV-K provirus sequences in urothelial
and renal cell carcinomas. Br J Cancer 80: 1312–1321.
Frommer M, McDonald LE, Millar DS, Collis CM, Watt F, Grigg GW, Molloy
PL, Paul CL. 1992. A genomic sequencing protocol that yields a positive
display of 5-methylcytosine residues in individual DNA strands. Proc
Natl Acad Sci 89: 1827–1831.
Gama-Sosa MA, Slagel VA, Trewyn RW, Oxenhandler R, Kuo KC, Gehrke
CW, Ehrlich M. 1983. The 5-methylcytosine content of DNA from
human tumors. Nucleic
Acids Res 11: 6883–6894.
Garbe JC, Bhattacharya S, Merchant B, Bassett E, Swisshelm K, Feiler HS,
Wyrobek AJ, Stampfer MR. 2009. Molecular distinctions between stasis
and telomere attrition senescence barriers shown by long-term culture
of normal human mammary epithelial cells. Cancer Res 69: 7557–7568.
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH,
de Klein A, Wessels L, de Laat W, et al. 2008. Domain organization of
human chromosomes revealed by mappi ng of nuclear lamina
interactions. Nature 453: 948–951.
Hansen RS, Thomas S, Sandstrom R, Canfield TK, Thurman RE, Weaver M,
Dorschner MO, Gartler SM, Stamatoyannopoulos JA. 2009. Sequencing
newly replicated DNA revea ls widespread plasticity in human
replication timing. Proc Natl Acad Sci 107: 139–144.
Hansen KD, Timp W, Bravo HC, Sabunciyan S, Langmead B, McDonald OG,
Wen B, Wu H, Liu Y, Diep D, et al. 2011. Increased methylation variation
in epigenetic domains across cancer types. Nat Genet 43: 768–775.
Hawkins RD, Hon GC, Lee LK, Ngo Q, Lister R, Pelizzola M, Edsall LE, Kuan
S, Luu Y, Klugman S, et al. 2010. Distinct epigenomic landscapes of
pluripotent and lineage-committed human cells. Cell Stem Cell 6: 479–
491.
Hellman A, Chess A. 2007. Gene body-specific methylation on the active X
chromosome. Science 315: 1141–1143.
Hennessy BT, Gonzalez-Angulo AM, Stemke-Hale K, Gilcrease MZ,
Krishnamurthy S, Lee JS, Fridlyand J, Sahin A, Agarwal R, Joy C, et al.
2009. Characterization of a naturally occurring breast cancer subset
enriched in epithelial-to-mesenchymal transition and stem cell
characteristics. Cancer Res 69: 4116–4124.
Herschkowitz JI, Simin K, Weigman VJ, Mikaelian I, Usary J, Hu Z,
Rasmussen KE, Jones LP, Assefnia S, Chandrasekharan S, et al. 2007.
Identification of conserved gene expression features between murine
mammary carcinoma models and human breast tumors. Genome Biol 8:
R76. doi: 10.1186/gb-2007-8-5-r76.
Herschkowitz JI, He X, Fan C, Perou CM. 2008 . The functional loss of the
retinoblastoma tumour suppressor is a common event in basal-like and
luminal B breast carcinomas. Breast Cancer Res 10: R75. doi: 10.1186/
bcr2142.
Hibi K, Goto T, Mizukami H, Kitamura Y, Sakata M, Saito M, Ishibashi K,
Kigawa G, Nemoto H, Sanada Y. 2009a. MGMT gene is aberrantly
methylated from the early stages of colorectal cancers.
Hepatogastroenterology 56: 1642–1644.
Hibi K, Sakata M, Yokomizo K, Kitamura YH, Sakuraba K, Shirahata A, Goto
T, Mizukami H, Saito M, Ishibashi K, et al. 2009b. Methylation of the
MGMT gene is frequently detected in advanced gastric carcinoma.
Anticancer Res 29: 5053–5055.
Hinshelwood RA, Clark SJ. 2008. Breast cancer epigenetics: Normal human
mammary epithelial cells as a model system. J Mol Med 86: 1315–
1328.
Hoadley KA, Weigman VJ, Fan C, Sawyer LR, He X, Troester MA, Sartor CI,
Rieger-House T, Bernard PS, Carey LA, et al. 2007. EGFR-associated
expression profiles vary with breast tumor subtype. BMC Genomics 8:
258. doi: 10.1186/1471-2164-8-258.
Holliday R. 1979. A new theory of carcinogenesis. Br J Cancer 40: 513–522.
Hsu F, Kent WJ, Clawson H, Kuhn RM, Diekhans M, Haussler D. 2006. The
UCSC Known Genes. Bioinformatics 22: 1036–1046.
Hu Z, Fan C, Livasy C, He X, Oh DS, Ewend MG, Carey LA, Subramanian S,
West R, Ikpatt F, et al. 2009. A compact VEGF signature associated with
distant metastases and poor outcomes. BMC Med 7: 9. doi: 10.1186/
1741-7015-7-9.
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P, Cui H,
Gabo K, Rongione M, Webster M, et al. 2009. The human colon cancer
methylome shows similar hyp o- and hypermethylation at conserved
tissue-specific CpG island shores. Nat Genet 41: 178–186.
Ishii H, Baffa R, Numata SI, Murakumo Y, Rattan S, Inoue H, Mori M, Fidanza
V, Alder H, Croce CM. 1999. The FEZ1 gene at chromosome 8p22
encodes a leucine-zipper protein, and its expression is altered in
multiple human tumors. Proc Natl Acad Sci 96: 3928–3933.
Ito S, D’Alessio AC, Taranova OV, Hong K, Sowers LC, Zhang Y. 2010. Role of
Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal, and inner
cell mass specification. Nature 466: 1129–1133.
Jiao Y, Shi C, Edil BH, de Wilde RF, Klimstra DS, Maitra A, Schulick RD, Tang
LH, Wolfgang CL, Choti MA, et al. 2011. DAXX/ATRX, MEN1, and
mTOR
pathway genes are frequently altered in pancreatic
neuroendocrine tumors. Science 331: 1199–1203.
Jones PA. 1999. The DNA methylation paradox. Trends Genet 15: 34–37.
Jones PA, Taylor SM, Wilson VL. 1983. Inhibition of DNA methylation by 5-
azacytidine. Recent Results Cancer Res 84: 202–211.
Jovanovic J, Ronneberg JA, Tost J, Kristensen V. 2010. The epigenetics of
breast cancer. Mol Oncol 4: 242–254.
Kaminskas E, Farrell AT, Wang YC, Sridhara R, Pazdur R. 2005. FDA drug
approval summary: Azacitidine (5-azacytidine, Vidaza) for injectable
suspension. Oncologist 10: 176–182.
Kelly TK, De Carvalho DD, Jones PA. 2010. Epigenetic modifications as
therapeutic targets. Nat Biotechnol 28: 1069–1078.
Kleer CG, Cao Q, Varambally S, Shen R, Ota I, Tomlins SA, Ghosh D, Sewalt
RG, Otte AP, Hayes DF, et al. 2003. EZH2 is a marker of aggressive breast
cancer and promotes neoplastic transformation of breast epithelial cells.
Proc Natl Acad Sci 100: 11606–11611.
Kohonen-Corish MR, Sigglekow ND, Susanto J, Chapuis PH, Bokey EL, Dent
OF, Chan C, Lin BP, Seng TJ, Laird PW, et al. 2007. Promoter methylation
of the mutated in colorectal cancer gene is a frequent early event in
colorectal cancer. Oncogene 26: 4435–4441.
Komashko VM, Farnham PJ. 2010. 5-azacytidine treatment reorganizes
genomic histone modification patterns. Epigenetics 5: 229–240.
Laird PW. 2003. The power and the promise of DNA methylation markers.
Nat Rev Cancer 3: 253–266.
Laird PW. 2010. Principles and challenges of genome-wide DNA
methylation analysis. Nat Rev Genet 11: 191–203.
DNA hypomethylation linkedtochromatinsilencing
Genome Research 257
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

Lakshmikuttyamma A, Scott SA, DeCoteau JF, Geyer CR. 2009. Reexpression
of epigenetically silenced AML tumor suppressor genes by SUV39H1
inhibition. Oncogene 29: 576–588.
Langmead B, Trapnell C, Pop M, Salzberg SL. 2009. Ultrafast and memory-
efficient alignment of short DNA sequences to the human genome.
Genome Biol 10: R25. doi: 10.1186/gb-2009-1 0-3-r25.
Lehnertz B, Ueda Y, Derijck AA, Braunschweig U, Perez-Burgos L, Kubicek S,
Chen T, Li E, Jenuwein T, Peters AH. 2003. Suv39h-mediated histone H3
lysine 9 methylation directs DNA methylation to major satellite repeats
at pericentric heterochromatin. Curr Biol 13: 1192 –1200.
Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE, Kandoth
C, Payton JE, Baty J, Welch J, et al. 2010. DNMT3A mutations in acute
myeloid leukemia. N Engl J Med 363: 2424–2433.
Li H, Handsak er B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G,
Abecasis G, Durbin R. 2009. The Sequence Alignment/Map format and
SAMtools. Bioinformatics 25: 2078–2079.
Lindroth AM, Park YJ, McLean CM, Dokshin GA, Persson JM, Herman H,
Pasini D, Miro X, Donohoe ME, Lee JT, et al. 2008. Antagonism between
DNA and H3K27 methylation at the imprinted Rasgrf1 locus. PLoS Genet
4: e1000145. doi: 10.1371/journal.pgen.1000145.
Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH,
Ecker JR. 2008. Highly integrated single-base resolution maps of the
epigenome in Arabidopsis. Cell 133: 523–536.
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery
JR, Lee L, Ye Z, Ngo QM, et al. 2009. Human DNA methylomes at base
resolution show widespread epigenomic differences. Nature 462: 315–322.
Lister R, Pelizzola M, Kida YS, Hawkins RD, Nery JR, Hon G, Antosiewicz-
Bourget J, O’Malley R, Castanon R, Klugman S, et al. 2011. Hotspots of
aberrant epigenomic reprogramming in human induced pluripotent
stem cells. Nature 471: 68–73.
Maunakea AK, Nagarajan RP, Bilenky M, Ballinger TJ, D’Souza C, Fouse SD,
Johnson BE, Hong C, Nielsen C, Zhao Y, et al. 2010. Conserved role of
intragenic DNA methylation in regulating alternative promoters. Nature
466: 253–257.
Menendez L, Benigno BB, McDonald JF. 2004. L1 and HERV-W
retrotransposons are hypomethylated in human ovarian carcinomas.
Mol Cancer 3: 12. doi: 10.1186/1476-4598-3-12.
Moir RD, Montag-Lowy M, Goldman RD. 1994. Dynamic properties of
nuclear lamins:Lamin B is associated with sites of DNA replication. J Cell
Biol 125: 1201–1212.
Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R, Paul JE,
Boyle M, Woolcock BW, Kuchenbauer F, et al. 2010. Somatic mutations
altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas
of germinal-center origin. Nat Genet 42: 181–185.
Mullins M, Perreard L, Quackenbush JF, Gauthier N, Bayer S, Ellis M, Parker J,
Perou CM, Szabo A, Bernard PS. 2007. Agreement in breast cancer
classification between microarray and quantitative reverse transcription
PCR from fresh-frozen and formalin-fixed, paraffin-embedded tissues.
Clin Chem 53: 1273–1279.
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T, Clark L, Bayani N,
Coppe JP, Tong F, et al. 2006. A collection of breast cancer cell lines for the
study of functionally distinct cancer subtypes. Cancer Cell 10: 515–527.
Novak P, Jensen T, Oshiro MM, Watts GS, Kim CJ, Futscher BW. 2008.
Agglomerative epigenetic aberrations are a common event in human
breast cancer. Cancer Res 68: 8616–8625.
Oh DS, Troester MA, Usary J, Hu Z, He X, Fan C, Wu J, Carey LA, Perou CM.
2006. Estrogen-regulated genes predict survival in hormone receptor-
positive breast cancers. J Clin Oncol 24: 1656–1664.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S,
Fauron C, He X, Hu Z, et al. 2009. Supervised risk predictor of breast
cancer based on intrinsic subtypes. J Clin Oncol 27: 1160–1167.
Parkhomchuk D, Borodina T, Amstislavskiy V, Banaru M, Hallen L,
Krobitsch S, Lehrac h H, Soldatov A. 2009. Transcriptome analysis by
strand-specific sequencing of complementary DNA. Nucleic Acids Res 37:
e123. doi: 10.1093/nar/gkp596.
Parsons DW, Li M, Zhang X, Jones S, Lear y RJ, Lin JC, Boca SM, Carter H,
Samayoa J, Bettegowda C, et al. 2010. The genetic landscape of the
childhood cancer medulloblastoma. Science 331: 435–439.
Peric-Hupkes D, Meuleman W, Pagie L, Bruggeman SW, Solovei I, Brugman
W, Graf S, Flicek P, Kerkhoven RM, van Lohuizen M, et al. 2010.
Molecular maps of the reorganization of genome-nuclear lamina
interactions during differentiation. Mol Cell 38: 603–613.
Perreard L, Fan C, Quackenbush JF, Mullins M, Gauthier NP, Nelson E, Mone
M, Hansen H, Buys SS, Rasmussen K, et al. 2006. Classification and risk
stratification of invasive breast carcinomas using a real-time quantitative
RT-PCR assay. Breast Cancer Res 8: R23. doi: 10.1186/bcr1399.
Picard . 2011. http://picard.sourceforge.net.
Prak ET, Kazazian HH Jr. 2000. Mobile elements and the human genome. Nat
Rev Genet 1: 134–144.
Prat A, Parker JS, Karginova O, Fan C, Livasy C, Herschkowitz JI, He X, Perou
CM. 2010. Phenotypic and molecular characterization of the claudin-
low intrinsic subtype of breast cancer. Breast Cancer Res 12: R68. doi:
10.1186/bcr2635.
Pruitt KD, Tatusova T, Maglott DR. 2005. NCBI Reference Sequence (RefSeq):
A curated non-redundant sequence database of genomes, transcripts,
and proteins. Nucleic Acids Res 33: D501–D504.
Rolny C, Mazzone M, Tugues S, Laoui D, Johansson I, Coulon C, Squadrito
ML, Segura I, Li X, Knevels E, et al. 2011. HRG inhibits tumor growth and
metastasis by inducing macrophage polarizatio n and vessel
normalization through downregulation of PlGF. Cancer Cell 19: 31–44.
Ruike Y, Imanaka Y, Sato F, Shimizu K, Tsujimoto G. 2010. Genome-wide
analysis of aberrant methylation in human breast cancer cells using
methyl-DNA immunoprecipitation combined with high-throughput
sequencing. BMC Genomics 11: 137. doi: 10.1186/1471-2164-11-137.
Schotta G, Lachner M, Sarma K, Ebert A, Sengupta R, Reuter G, Reinberg D,
Jenuwein T. 2004. A silencing pathway to induce H3-K9 and H4-K20
trimethylation at constitutive heterochromatin. Genes Dev 18: 1251–1262.
Schulz WA. 2006. L1 retrotransposons in human cancers. J Biomed Biotechnol
2006: 83672. doi: 10.1155/JBB/2006/83672.
Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD, Mandelker D,
Leary RJ, Ptak J, Silliman N, et al. 2006. The consensus coding sequences
of human breast and colorectal cancers. Science 314: 268–274.
Stephens PJ, McBride DJ, Lin ML, Varela I, Pleasance ED, Simpson JT,
Stebbings LA, Leroy C, Edkins S, Mudie LJ, et al. 2009. Complex
landscapes of somatic rearrangement in human breast cancer genomes.
Nature 462: 1005–1010.
Streit M, Riccardi L, Velasco P, Brown LF, Hawighorst T, Bornstein P, Detmar
M. 1999. Thrombospondin-2: A potent endogenous inhibitor of tumor
growth and angiog enesis. Proc Natl Acad Sci 96: 14888–14893.
Tapia T, Smalley SV, Kohen P, Munoz A, Solis LM, Corvalan A, Faundez P,
Devoto L, Camus M, Alvarez M, et al. 2008. Promoter hypermethylation
of BRCA1 correlates with absence of expression in hereditary breast
cancer tumors. Epigenetics 3: 157–163.
Teer JK, Bonnycastle LL, Chines PS, Hansen NF, Aoyama N, Swift AJ, Abaan
HO, Albert TJ, Margulies EH, Green ED, et al. 2010. Systematic
compa
rison of three genomic enrichment methods for massively
parallel DNA sequencing. Genome Res 20: 1420–1431.
Ting DT, Lipson D, Paul S, Brannigan BW, Akhavanfard S, Coffman EJ,
Contino G, Deshpande V, Iafrate AJ, Letovsky S, et al. 2011. Aberrant
overexpression of satellite repeats in pancreatic and other epithelial
cancers. Science 331: 593–596.
Trapnell C, Pachter L, Salzberg SL. 2009. TopHat: Discovering splice
junctions with RNA-Seq. Bioinformatics 25: 1105–1111.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ,
Salzberg SL, Wold BJ, Pachter L. 2010. Transcript assembly and
quantification by RNA-Seq reveals unannotated transcripts and isoform
switching during cell differentiation. Nat Biotechno l 28: 511–515.
van Haaften G, Dalgliesh GL, Davies H, Chen L, Bignell G, Greenman C,
Edkins S, Hardy C, O’Meara S, Teague J, et al. 2009. Somatic mutations of
the histone H3K27 demethylase gene UTX in human cancer. Nat Genet
41: 521–523.
Varela I, Tarpey P, Raine K, Huang D, Ong CK, Stephens P, Davies H, Jones D,
Lin ML, Teague J, et al. 2011. Exome sequencing identifies frequent
mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma.
Nature 469: 539–542.
Volkel P, Angrand PO. 2007. The control of histone lysine methylation in
epigenetic regulation. Biochimie 89: 1–20.
Weigelt B, Hu Z, He X, Livasy C, Carey LA, Ewend MG, Glas AM, Perou CM,
Van’t Veer LJ. 2005. Molecular portraits and 70-gene prognosis signature
are preserved throughout the metastatic process of breast cancer. Cancer
Res 65: 9155–9158.
Wild L, Flanagan JM. 2010. Genome -wide hypomethylation in cancer may
be a passive consequence of transformation. Biochim Biophys Acta 1806:
50–57.
Wood LD, Parsons DW, Jones S, Lin J, Sjoblom T, Leary RJ, Shen D, Boca SM,
Barber T, Ptak J, et al. 2007. The genomic landscapes of human breast
and colorectal cancers. Science 318: 1108–1113.
Wu H, Coskun V, Tao J, Xie W, Ge W, Yoshikawa K, Li E, Zhang Y, Sun YE.
2010. Dnmt3a-dependent nonpromoter DNA methylation facilitates
transcription of neurogenic genes. Science 329: 444–448.
Yap DBJ, Chu T, Berg M, Schapira SW, Cheng A, Moradian RD, Morin AJ,
Mungall B, Meissner M, Boyle et al. 2010. Somatic mutations at EZH2 Y641
act dominantly through a mechanism of selectively altered PRC2 catalytic
activity, to increase H3K27 trimethylation. Blood 117: 2451–2459.
Yuan BZ, Miller MJ, Keck CL, Zimonjic DB, Thorgeirsson SS, Popescu NC.
1998. Cloning, characterization, and chromosomal localization of
a gene frequently deleted in human liver cancer (DLC-1) homologous to
rat RhoGAP. Cancer Res 58: 2196–2199.
Received May 4, 2011; accepted in revised form September 26, 2011.
Hon et al.
258 Genome Research
www.genome.org
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from

10.1101/gr.125872.111Access the most recent version at doi:
2012 22: 246-258 originally published online December 7, 2011Genome Res.
Gary C. Hon, R. David Hawkins, Otavia L. Caballero, et al.
domain formation and gene silencing in breast cancer
Global DNA hypomethylation coupled to repressive chromatin
Material
Supplemental
http://genome.cshlp.org/content/suppl/2011/10/06/gr.125872.111.DC1
References
http://genome.cshlp.org/content/22/2/246.full.html#ref-list-1
This article cites 103 articles, 26 of which can be accessed free at:
License
Service
Email Alerting
click here.top right corner of the article or
Receive free email alerts when new articles cite this article - sign up in the box at the
https://genome.cshlp.org/subscriptions
go to: Genome Research To subscribe to
Copyright © 2012 by Cold Spring Harbor Laboratory Press
Cold Spring Harbor Laboratory Press on September 17, 2024 - Published by genome.cshlp.orgDownloaded from
