
VIM1, a methylcytosine-binding
protein required for centromeric
heterochromatinization
Hye Ryun Woo, Olga Pontes, Craig S. Pikaard, and Eric J. Richards
1
Department of Biology, Washington University, St. Louis, Missouri 63130, USA
Epigenetic regulation in eukaryotes is executed by a complex set of signaling interactions among small RNA
species and chromatin marks, including histone modification and DNA methylation. We identified vim1
(VARIANT IN METHYLATION 1), an Arabidopsis mutation causing cytosine hypomethylation and
decondensation of centromeres in interphase. VIM1 is a member of a small gene family, encoding proteins
containing PHD, RING, and SRA (SET- and RING-associated) domains, which are found together in
mammalian proteins implicated in regulation of chromatin modification, transcription, and the cell cycle.
VIM1 is an unconventional methylcytosine-binding protein that interacts in vitro with 5mCpG- and
5mCpHpG-modified DNA (via its SRA domain), as well as recombinant histones (H2B, H3, H4, and HTR12)
in plant extracts. VIM1 associates with methylated genomic loci in vivo and is enriched in chromocenters.
Our findings suggest that VIM1 acts at the DNA methylation–histone interface to maintain centromeric
heterochromatin.
[Keywords: Cytosine methylation; methylcytosine-binding protein; centromere; epigenetic; SRA domain;
heterochromatin]
Supplemental material is available at http://www.genesdev.org.
Received November 13, 2006; revised version accepted December 14, 2006.
Large regions of higher eukaryotic genomes, particularly
the domains surrounding and encompassing centro-
meres, are composed of repetitive elements, which are
preferential targets for heterochromatin assembly. Het-
erochromatic regions of the genome are characterized by
increased chromatin condensation and decreased or dif-
ferential accessibility to regulatory proteins (Craig 2005;
Huisinga et al. 2006). Most forms of heterochromatin are
stably inherited and contain one or more epigenetic
marks that direct its maintenance during cell division.
Specification and maintenance of heterochromatin and
other functionally distinct chromatin domains rely on
complex interactions among cytosine methylation, his-
tone modification, and RNA interference (RNAi) path-
ways (Tamaru and Selker 2001; Jaenisch and Bird 2003;
Chan et al. 2005; Matzke and Birchler 2005).
Cytosine methylation is an important mechanism for
establishing stable heritable epigenetic marks, thereby
linking primary nucleotide sequence to chromatin orga-
nization (Jaenisch and Bird 2003). The cytosine methyl-
ation reaction is catalyzed by enzymes known as cy-
tosine-DNA-methyltransferases (DNMTs) (Goll and
Bestor 2005). Notably, mammalian cytosine methylation
is mostly restricted to symmetrical CpG sequences, al-
though plant cytosine methylation occurs at CpG,
CpHpG, and CpHpH sequences (where H = A, C, T).
Once established, cytosine methylation can be inherited
through mitosis, and sometimes through meiosis, pro-
viding a stable epigenetic mark (Genereux et al. 2005;
Richards 2006). Cytosine methylation cooperates with
histone modification to generate a self-reinforcing cycle
of epigenetic events that lead to long-term transcrip-
tional repression (Jenuwein and Allis 2001; Nightingale
et al. 2006). The regulatory information encoded by cy-
tosine methylation is deciphered by a family of proteins
that, through a conserved methyl-CpG-binding domain
(MBD), selectively binds methylated CpG dinucleotides
regardless of the sequence context (Klose and Bird 2006).
Some MBD proteins can recruit histone deacetylase
(HDAC) or histone methyltransferase (HMT) complexes
as well as chromatin remodeling factors (Klose and Bird
2006). Chromatin modification can also influence cyto-
sine methylation. For example, methylation at histone
H3K9 creates binding sites for adaptor molecules, such
as HP1 (Heterochromatin Protein 1) and related chromo-
domain proteins, that ultimately recruit DNMTs (Fuks
et al. 2003; Lehnertz et al. 2003). Through these types of
interactions, covalent modifications of DNA and core
1
Corresponding author.
Article published online ahead of print. Article and publication date are
online at http://www.genesdev.org/cgi/doi/10.1101/gad.1512007.
GENES & DEVELOPMENT 21:000–000 © 2007 by Cold Spring Harbor Laboratory Press ISSN 0890-9369/07; www.genesdev.org 1
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
histones play important roles in heterochromatin forma-
tion in mammals and plants.
The last several years have witnessed tremendous ad-
vances in the genetic analysis of eukaryotic DNA meth-
ylation, resulting in the identification of proteins and
pathways necessary for specification and maintenance of
genomic cytosine methylation patterns (Jaenisch and
Bird 2003; Rangwala and Richards 2004; Chan et al.
2005). Two major protein categories important for geno-
mic cytosine methylation, identified through both for-
ward and reverse genetic approaches, are DNMTs and
chromatin modification enzymes (e.g., HDACs, HMTs,
nucleosome remodelers). Post-transcriptional silencing/
RNAi pathways mutants also have cytosine hypometh-
ylation phenotypes, highlighting the importance of
RNA-directed DNA methylation mechanisms guided by
small RNA species. However, only a handful of these
mutations affect centromere DNA methylation, despite
the fact that the centromere is a principle target for cy-
tosine methylation and heterochromatin formation in
the cell. In an effort to identify novel alleles or new loci
affecting centromere DNA methylation, we chose an al-
ternative approach of screening natural strains of the
flowering plant Arabidopsis thaliana for changes in ge-
nomic methylation patterns. Through this approach, we
identified a novel genetic locus, VIM1 (VARIANT IN
METHYLATION 1), encoding a non-MBD class methyl-
cytosine-binding protein that is required for full centro-
mere DNA methylation. In addition, loss of VIM1 func-
tion leads to decondensation of the centromere repeat
sequence in interphase and an alteration in the localiza-
tion pattern of the centromere-specific histone H3 vari-
ant, HTR12. Our results suggest that VIM1 acts at the
interface between DNA methylation and chromatin to
maintain centromere heterochromatin.
Results
Identification of a natural variant with
hypomethylated centromere DNA
In a screen for natural variation in cytosine methylation
among 89 different strains of A. thaliana, we identified
one strain that contained hypomethylated centromeric
180-base-pair (bp) repeats (CEN). Centromere repeats
from strain Borky-4 (Bor-4) showed an increased level of
digestion with the restriction endonuclease HpaII (5⬘-
CCGG-3⬘), which is inhibited by either 5mCpG or
5mCpHpG, relative to that seen for repeats in wild-type
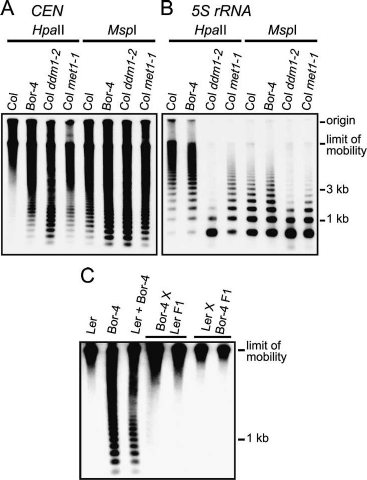
Columbia (Col) plants (Fig. 1A). A similar enhanced di-
gestion was seen when the centromere repeats were cut
with MspI, which cleaves the same site as HpaII but is
inhibited only by 5mCpHpG. The effect of two previ-
ously characterized DNA hypomethylation mutations in
strain Col are shown for comparison: ddm1, which re-
duces cytosine methylation in all sequence contexts
(Vongs et al. 1993); and met1, which predominantly re-
duces 5mCpG (Kankel et al. 2003; Saze et al. 2003).
These observations suggest that Bor-4 contains centro-
meric repeats that are hypomethylated at both CpG and
CpHpG sites.
To determine whether reduced cytosine methylation
in Bor-4 occurs at other genomic locations, we examined
the extent of cytosine methylation at three additional
loci that are heavily methylated in wild-type strains: the
FWA gene (Soppe et al. 2000) and the pericentromeric
tandemly repeated 5S rRNA genes (Campell et al. 1992)
and Athila retrotransposable elements (Pelissier et al.
1995). Compared with wild-type Col, we did not find a
significant change in Bor-4 cytosine methylation at the
5S rRNA genes (Fig. 1B), in the Athlia elements or at the
FWA locus (data not shown). In agreement with previous
reports, we observed a strong hypomethylation of all
these loci in Col plants containing either ddm1-2 or
met1-1 alleles. These results indicate that DNA hypo-
methylation in wild strain Bor-4 preferentially affects
the 180-bp centromere repeats.
Altered centromere organization in Bor-4 interphase
nuclei
To investigate whether centromere heterochromatin
was affected in Bor-4, we used fluorescence in situ hy-
bridization (FISH) to detect the 180-bp centromere re-
Figure 1. Reduced cytosine methylation of centromeric re-
peats in Bor-4. (A) Genomic DNA samples from the indicated
genotypes were digested with the isoschizomers HpaII or MspI,
and DNA blot hybridization with a 180-bp centromere repeat
probe (CEN) was performed. (B) The filter shown in A was re-
hybridized with a 5S rRNA probe (5S rRNA). (C) A DNA blot
hybridization pattern with the CEN probe after HpaII digestion
demonstrates that centromeric repeat arrays hypomethylated in
Bor-4 were fully remethylated in F1 hybrids resulting from re-
ciprocal crosses between strains Bor-4 and Ler. The lane labeled
Ler + Bor-4 contains a 1:1 mixture of Ler and Bor-4 HpaII-di-
gested genomic DNA and shows the hybridization pattern ex-
pected if no remethylation occurred in the F1 hybrids.
Woo et al.
2 GENES & DEVELOPMENT
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
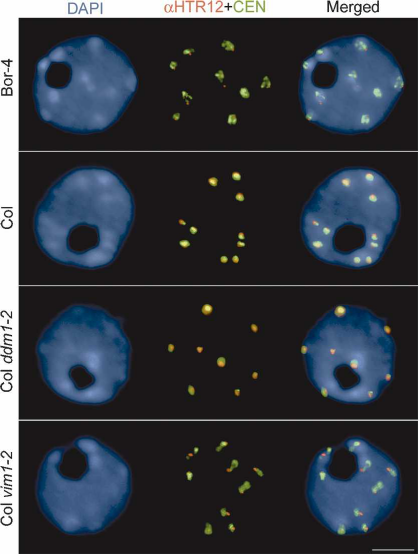
peats in interphase nuclei. The spatial distribution of
hybridization signal indicated that the centromere re-
peats in Bor-4 were decondensed relative to the repeats
in Col nuclei (Fig. 2; Supplementary Table 1). We also
stained fixed interphase nuclei with anti-HTR12 anti-
body (Talbert et al. 2002) to obtain a more comprehen-
sive understanding of centromere heterochromatin in
Bor-4. HTR12 is the Arabidopsis homolog of human
CENP-A, Drosophila Cid, and yeast Cnp1, which are his-
tone H3 variants that define the specialized chromatin
structure associated with kinetochore assembly (Heni-
koff and Dalal 2005). We found that the area of the
HTR12 immunostained signal was decreased in Bor-4
nuclei compared with the pattern observed in Col nuclei.
We noted that neither the distribution of 180-bp repeats,
nor the pattern of HTR12 staining is affected by the
ddm1 mutation in the Col background (Fig. 2). Thus,
Bor-4 exhibits not only abnormal centromere DNA
methylation, but a unique centromere organization phe-
notype not exhibited by a well-characterized mutation
with severe centromere DNA hypomethylation defects.
A trans-acting mutation causes centromere DNA
hypomethylation in Bor-4
To distinguish whether the Bor-4 centromere pheno-
types result from the action of a trans-acting mutation or
epigenetic inheritance of stochastic DNA hypomethyl-
ation, we outcrossed Bor-4 to standard laboratory strains
Col and Ler and followed the segregation of the DNA
hypomethylation phenotype in subsequent generations.
In the F1 generation, regardless of the direction of the
cross, the hypomethylated 180-bp repeat arrays originat-
ing from Bor-4 were fully remethylated, demonstrating
that centromere DNA hypomethylation is reset effi-
ciently (Fig. 1C; data not shown). This result contrasts
with the persistence of hypomethylated genomes in F1
hybrids resulting from outcrossing ddm1 or met1 mu-
tants (Vongs et al. 1993; Kakutani et al. 1999; Kankel et
al. 2003), and argues against the possibility that Bor-4
centromere repeat hypomethylation is due strictly to
epigenetic inheritance of a hypomethylated state. The
hypomethylated centromere trait segregated as a mono-
genic recessive trait in two independent Ler X Bor-4 F2
families (normal:hypomethylated⬋112:36). Recombina-
tional mapping in Ler X Bor-4 F2 families indicated that
the hypomethylated centromere phenotype of Bor-4 is
caused by variation at a single trans-acting locus, VIM1,
at a map position distinct from known Arabidopsis mu-
tations affecting DNA methylation or chromatin modi-
fication (Supplementary Fig. 1).
Identification of VIM1
Using a Ler X Bor-4 F2 mapping population, we nar-
rowed the region containing the vim1-1 allele from Bor-4
to an interval corresponding to a 113-kb region on the
lower arm of chromosome 1. In Col, this window con-
tains 30 annotated genes (Supplementary Fig. 1). The
Bor-4 allele of one gene in this interval, At1g57820, car-
ries a large deletion. The missing segment spans a 3.2-kb
region from just downstream from the first intron donor
site to the middle of the intergenic region between
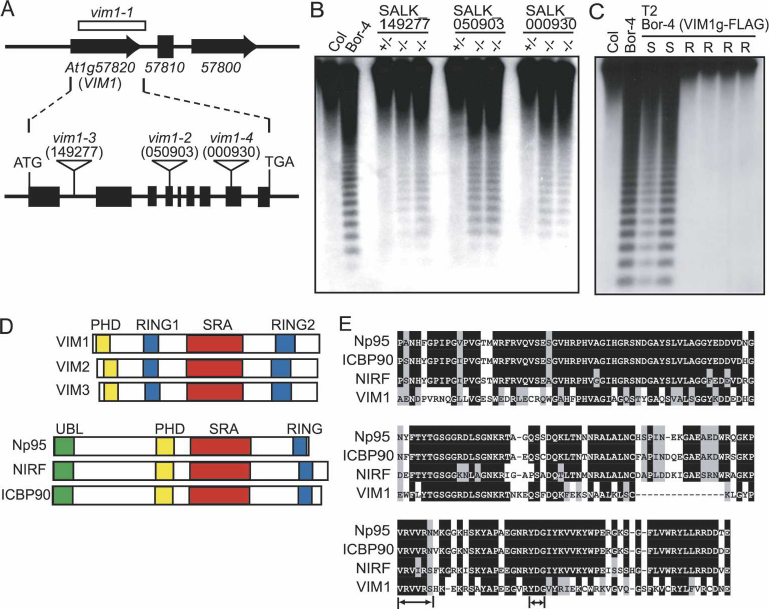
At1g57820 and At1g57810 (Fig. 3A; Supplementary Fig.
2). We could not detect At1g57820 transcript in Bor-4
using RT–PCR analysis; however, the predicted tran-
script was observed in Col and Ler (data not shown). To
test whether mutation of At1g57820 causes hypometh-
ylation of the 180-bp centromere repeats, we analyzed
the effect of Agrobacterium T-DNA insertion alleles in
this gene in the Col background. The T-DNA inserts in
At1g57820 disrupt the first intron, the fourth exon, and
the eighth exon, respectively (Fig. 3A). We could not de-
tect full-length transcripts from any Col At1g57820 ho-
mozygous T-DNA mutant (data not shown). All three
T-DNA insertion mutants in Col showed increased
HpaII cleavage of the 180-bp centromere repeats (Fig. 3B),
although the diagnostic ladder-like hybridization pattern
was weaker than that seen in Bor-4 (Fig. 1A). The muted
effect of the T-DNA insertion alleles in At1g57820 may
be due to the action of strain-specific modifiers that par-
tially cover for At1g57820 loss of function in Col. How-
ever, the vim1-2 allele (SALK_050903) in the Col strain
caused distinct centromere defects similar to those seen
in Bor-4 (Fig. 2), arguing that disruption of VIM1 function
is responsible for both the altered centromere heterochro-
matin and centromere DNA hypomethylation phenotypes.
Figure 2. Centromeric heterochromatin is altered in Bor-4.
One-hundred-eighty-base-pair centromeric repeats (CEN) were
detected by FISH, and HTR12 protein was immunolocalized in
interphase nuclei from root tip cells of Bor-4, Col, Col ddm1-2,
and Col vim1-2 (SALK_050903) plants. The DNA was counter-
stained with DAPI; chromocenters are more intensely stained.
Bar, 5 µm.
5mC-binding protein required for DNA methylation
GENES & DEVELOPMENT 3
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
We confirmed that At1g57820 corresponds to the
VIM1 locus by two independent genetic tests. Crosses
between Bor-4 and Col plants homozygous for an
At1g57820 T-DNA insertion allele demonstrated a lack
of complementation for the centromere DNA methyl-
ation phenotype in the resulting F1 individuals (Supple-
mentary Fig. 3). Furthermore, a 5.5-kb genomic fragment
containing the Col sequence of At1g57820 was sufficient
to complement the hypomethylation phenotype of Bor-4
(Fig. 3C). The VIM1 locus encodes a 645-amino-acid
protein with a PHD domain, two RING finger domains,
and an SRA (SET- and RING-associated) domain (Fig. 3D;
Supplementary Fig. 4A). Four additional genes encod-
ing related proteins (ⱖ68% amino acid identity to
VIM1) with a similar domain organization exist in the
Arabidopsis genome: At1g57800, At1g66040, At1g66050,
and At5g39550 (Fig. 3D; Supplementary Fig. 4B). Only
three of these genes (At1g57820 [VIM1], At1g66050
[VIM2], and At5g39550 [VIM3]) are expressed at an ap-
preciable level in strain Bor-4, Col and Ler based on our
RT–PCR analysis (data not shown) and cDNA resources
available through public databases (e.g., http://www.
arabidopsis.org, http://mpss.udel.edu/at). Neither the
vim2 nor the vim3 T-DNA insertion allele examined in
Col caused a centromere repeat hypomethylation pheno-
type (Supplementary Fig. 5).
Centromeric small interfering RNA (siRNA)
populations are not significantly changed in Col
vim1 plants
Several characterized Arabidopsis DNA hypomethyl-
ation mutations primarily affect small RNA accumula-
tion, which reduces RNA-directed DNA methylation
Figure 3. Identification of the VIM1 gene. (A) Diagram of the intron–exon structure of the VIM1 gene and positions of T-DNA
insertions in At1g57820 with allele designations. At1g57810 (reverse transcriptase pseudogene) is located between At1g57800 and
At1g57820. The deletion in Bor-4 is shown as an open box (vim1-1 allele). Filled boxes represent exons and thin lines represent introns.
The positions of the start and stop codon are indicated, as are the T-DNA insertion sites in At1g57820.(B) DNA blot hybridization
(HpaII digest, CEN probe) analysis of T-DNA insertion mutants in At1g57820. Genomic DNA was isolated from independent plants
in segregation populations. (+/+) Homozygous wild-type allele; (+/−) heterozygous for mutant allele; (−/−) homozygous for mutant
allele. (C) DNA blot hybridization (HpaII digest, CEN probe) analysis demonstrating complementation of the hypomethylation
phenotype of Bor-4 with a transgene expressing At1g57820 (from Col strain, coding sequence fused to a C-terminal Flag epitope tag).
Analysis of six individuals in the T2 generation segregating the transgene (carrying a Basta herbicide resistance marker: [R] resistant;
[S] sensitive) is shown. (D) Schematic representation of possible homologs of VIM1. The RING (Pfam PF00097), PHD (Pfam PF000628),
and SRA (Pfam PF02182) domains are labeled. The mammalian proteins contain an N-terminal ubiquitin-like domain (UBL), which
is absent in the VIM1-like Arabidopsis proteins, and only one RING domain. (E) Sequence alignment of SRA domains from VIM1 and
three related mammalian proteins. Black shading shows identical residues and gray shading shows similar residues. Arrows denote the
conserved VRV(I/V)RG and YDG motifs in SRA domains.
Woo et al.
4 GENES & DEVELOPMENT
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
(Chan et al. 2005). Consequently, we compared the abun-
dance of siRNAs corresponding to the 180-bp centro-
mere repeats in Col and Col vim1-2 plants using RNA
gel blot analysis (Supplementary Fig. 6). We did not find
a significant change in the abundance of centromeric
small RNA populations associated with the VIM1 geno-
type, suggesting that the vim1-2 mutation reduces cen-
tromere DNA methylation without affecting siRNA ac-
cumulation.
VIM1 is a methylcytosine-binding protein
The similarity shared among the SRA domains of VIM1
and three mammalian proteins, human ICBP90 and
NIRF, and murine Np95 (Fig. 3D,E; Fujimori et al. 1998;
Mori et al. 2002) prompted us to investigate whether
VIM1 is a methylcytosine-binding protein because the
SRA domain in these mammalian proteins was shown to
bind methyl-CpG-modified double-stranded DNA
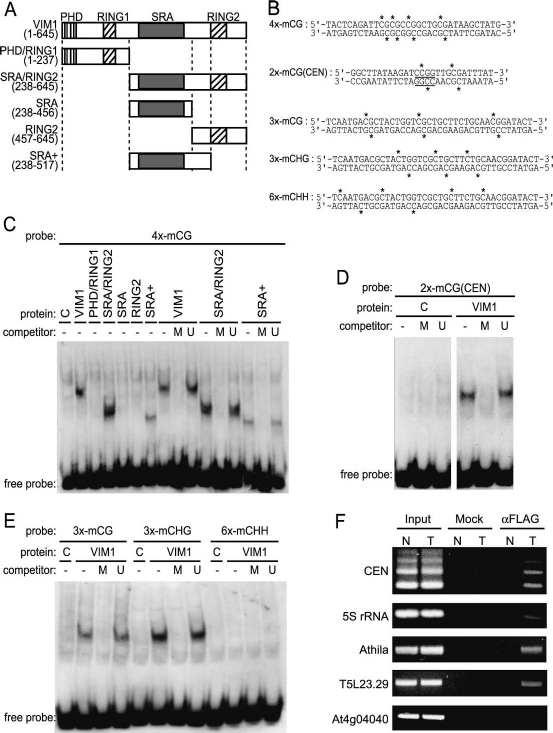
(Unoki et al. 2004). Full-length VIM1 protein and various
VIM1 subdomains (Fig. 4) expressed in vitro were tested
for their ability to bind methylated double-stranded oli-
gonucleotides by electrophoretic mobility shift assay
(EMSA). We demonstrated that full-length VIM1 bound
to a double-stranded oligonucleotide (4x-mCG) contain-
ing four symmetrical 5mCpG sites in a methylation-de-
pendent manner (Fig. 4A–C). VIM1 bound the same oli-
gonucleotide sequence containing only a single 5mCpG
site, but did so more weakly (data not shown). Because
vim1 mutations lead to a reduction in cytosine methyl-
ation in the 180-bp centromere repeats, we tested and
confirmed VIM1’s ability to interact specifically with an
oligonucleotide corresponding to the region of the 180-
bp repeat sequence containing the 5⬘-CCGG-3⬘ site as-
sayed (Fig. 4B,D). Next, we delimited the minimal region
of VIM1 that interacted in a methylation-dependent
manner with the 4x-mCG probe to 279 amino acids com-
prising the entire SRA domain and the interval between
the SRA and second RING domain (Fig. 4A,C, lanes
marked SRA+). While the SRA domain is required for
methylcytosine-binding activity, this domain is not suf-
ficient (Fig. 4A,C, lane marked SRA).
Plant genomes contain cytosine methylation in three
different sequence contexts: two symmetrical sites, CpG
and CpHpG; and the asymmetric site, CpHpH (Chan et
al. 2005). We assayed the specificity of VIM1’s methyl-
cytosine-binding activity using a set of double-stranded
oligonucleotides of identical primary sequence that con-
Figure 4. VIM1 is a methylcytosine-binding protein.
(A) Schematic of the VIM1 protein and derivatives
tested for methylcytosine-binding activity; amino acid
coordinates are shown in the parentheses. Domain des-
ignations include PHD domain, SRA domain, and
RING domains. (B) Oligonucleotide probes used in the
methylcytosine-binding assays shown in C–E. Aster-
isks indicate the positions of 5mC residues. The under-
lined 5⬘-CCGG-3⬘ site corresponds to the HpaII assayed
in Figure 1. (C) VIM1 protein derivatives were expressed
in wheat germ transcription/translation extracts (C,
vector-only control) and incubated with the digoxi-
genin-endlabeled 4x-mCG probe in the presence of 62-
fold excess unlabeled competitor oligonucleotides. (U)
Unmethylated; (M) methylated; (−) no competitor.
EMSA of incubation products is shown; the position of
the unbound free probe is indicated at the bottom of the
image. The same abbreviations and symbols are used in
C–E.(D) EMSA demonstrating the ability of full-length
VIM1 to bind a methylated oligonucleotide probe based
on the 180-bp centromere repeat sequence. (E) EMSA of
interaction of full-length VIM1 with identical probes
methylated in different cytosine contexts. (F) ChIP was
performed on nuclei prepared from Col (N, nontrans-
genic) and transgenic (T) plants expressing Flag-VIM1.
After immunoprecipitation, samples were used as tem-
plates for amplification of different genomic sequences,
which are indicated at the left of the images of the re-
sulting ethidium-stained gels. (Input) 10% of input;
(Mock) precipitation in the absence of antibody; (␣Flag)
precipitation with anti-Flag antibody.
5mC-binding protein required for DNA methylation
GENES & DEVELOPMENT 5
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
tained cytosine methylation in only one of the three pos-
sible sequence contexts (Fig. 4B,E). As expected, VIM1
bound the oligonucleotide modified at 5mCpG sites.
VIM1 was unable to bind the 5mCpHpH-containing oli-
gonucleotide, but interacted in a methylation-dependent
manner with the 5mCpHpG-containing oligonucleotide.
Therefore, VIM1 recognizes cytosine methylation in two
different sequence contexts that occur in plant genomes,
including centromere repeat arrays.
After demonstrating that VIM1 has methylcytosine-
binding activity in vitro, we next determined whether
VIM1 associates with methylated DNA in vivo. We em-
ployed a chromatin immunoprecipitation (ChIP) assay
on nuclei prepared from wild-type Col and transgenic
Arabidopsis plants expressing Flag-VIM1. Cross-linked
chromatin complexes were precipitated with or without
anti-Flag antibody, and the precipitated DNA samples
were used as templates to amplify centromeric 180-bp
repeats. As seen in Figure 4F, the 180-bp centromere re-
peat arrays were associated with the VIM1 protein. The
unmethylated locus At4g04040 was not associated with
VIM1 (Gendrel et al. 2002; Lippman et al. 2004). We also
investigated three other methylated genomic loci, in-
cluding the tandemly repeated 5S rRNA genes (5S
rRNA), repetitive Athila retrotransposable elements
(Athila), and a Cinful-like retrotransposon (T5L23.29)
(Gendrel et al. 2002). The anti-Flag antibody precipitated
all methylated genomic sequences tested. Our results
indicate that VIM1 associates with methylated DNA
both in vitro and in vivo. The specificity of vim1 DNA
hypomethylation effects may stem from a heightened
sensitivity of centromeric chromatin to loss of VIM1
function.
VIM1 is concentrated at chromocenters
We next studied the subnuclear localization of VIM1
relative to interphase chromatin organization. The VIM1
genomic coding sequence, expressed under control of its
native promoter, was engineered to express full-length
VIM1 fused to the Flag epitope at the C terminus
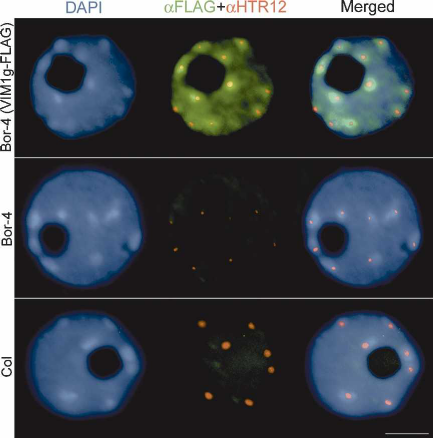
(VIM1g-Flag) in transgenic plants. Fixed nuclei from Col,
Bor-4, and transgenic Bor-4 expressing VIM1g-Flag were
immunolabeled with anti-Flag and anti-HTR12 antibod-
ies. Introduction of the VIM1g-Flag construct comple-
mented three centromere-related phenotypes in Bor-4:
decondensation of 180-bp centromere repeats (data not
shown), altered HTR12 localization (Fig. 5, middle col-
umn; Supplementary Table 2), and centromere DNA hy-
pomethylation (Fig. 3C). These results demonstrate that
the epitope-tagged VIM1 is functional. Epitope-tagged
VIM1 protein was localized to the nucleus, where it
showed a broad distribution (excluding the nucleolus)
but enrichment in the heterochromatic chromocenters
in the majority of the cells examined (Fig. 5, top row;
Supplementary Table 3). VIM1 localization is consistent
with our ChIP results (Fig. 4F) and the known distribu-
tion of methylated DNA sequences, which tend to be
concentrated in the chromocenters.
VIM1–histone interaction
The centromere phenotypes of vim1 mutants, including
the apparent changes in HTR12 localization, coupled
with the observation that the SRA domain of murine
Np95 interacts differentially with core histones (Citterio
et al. 2004), led us to test whether VIM1 physically in-
teracts with HTR12. An epitope-tagged HTR12 (N-ter-
minal c-myc tag) was expressed in Escherichia coli and
partially purified using an anti-myc conjugated resin. In
parallel, we expressed and partially purified recombinant
versions of other Arabidopsis histones, including myc-
H2A (HTA2), myc-H2B (HTB2), myc-H3 (HTR3), and
myc-H4 (HFO4). Purified recombinant histones were in-
cubated with extracts from transgenic plants overex-
pressing Flag-VIM1. Proteins associated with the immo-
bilized recombinant histones were subject to Western
blot analysis to monitor the recovery of the Flag-VIM1
protein. As shown in Figure 6A, Flag-VIM1 interacted
with three of the core histones (with the exception of
H2A [HTA2]), as well as the centromere-specific histone
H3, HTR12.
Discussion
We describe the identification and isolation of the Ara-
bidopsis VIM1 gene encoding a methylcytosine-binding
protein required for maintenance of centromere DNA
methylation and proper interphase centromere organiza-
tion. The VIM1 gene was originally identified by a spon-
taneous loss-of-function mutation in a wild accession
that did not exhibit a striking morphological phenotype.
Figure 5. Subnuclear localization of VIM1. Immunolocaliza-
tion of the epitope-tagged VIM1 (␣Flag) and HTR12 (␣HTR12)
was performed on nuclei derived from nontransgenic Bor-4 and
Col plants, as well as transgenic Bor-4 plants expressing VIM1-
Flag under control of the native VIM1 promoter (VIM1g-Flag).
DAPI was used as a DNA counterstain. Bar, 5 µm.
Woo et al.
6 GENES & DEVELOPMENT
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
Plants are remarkably tolerant of dramatic alterations in
DNA methylation relative to mammals. However, sev-
eral Arabidopsis mutants with altered cytosine methyl-
ation suffer developmental abnormalities and reduced
fertility under optimal laboratory conditions, suggesting
that these mutants would compete poorly in natural set-
tings. The persistence of the vim1-1 mutation in natural
populations indicates that the alteration in centromere
organization and DNA methylation caused by loss of
VIM1 can be tolerated in the wild, and highlights the
viability of mining natural variation for novel alleles
that affect epigenetic regulation.
Loss of VIM1 function causes CpG and CpHpG hypo-
methylation at the centromere, but has no significant
effect on DNA methylation of the FWA gene or pericen-
tromeric sequences (5S rRNA genes and Athila retro-
transposable elements). The apparent specificity of vim1
effects contrasts with the rather broad spectrum of ge-
nomic targets affected by mutations in other Arabidop-
sis genes involved in cytosine methylation (Rangwala
and Richards 2004; Chan et al. 2005). Another interest-
ing feature of the vim1-induced hypomethylation is that
the centromeric repeats are completely remethylated in
F1 hybrids created by outcrossing with wild-type plants.
This behavior contrasts with the inheritance of the hy-
pomethylated state of the centromere induced by ddm1
or met1 in crosses with wild-type plants. The efficient
re-establishment of DNA methylation on the centro-
mere repeats may reflect the presence in vim1 mutants
of centromeric siRNA populations, which may target
these repeats for de novo methylation (Lippman et al.
2003). The altered interphase organization of the centro-
mere observed in vim1 mutants is also unique. Both
ddm1 and met1 mutations cause a decondensation of
pericentromeric sequences, which become dispersed
from chromocenters while the 180-bp centromere re-
peats remain condensed within the chromocenters
(Soppe et al. 2002). In contrast, vim1 mutants display a
decondensation of the 180-bp centromere repeats, sug-
gesting that VIM1 plays a role in centromere organiza-
tion that is distinct from its function in DNA methyl-
ation.
VIM1 is a methylcytosine-binding protein that
interacts with 5mCpG and 5mCpHpG
Methylcytosine-binding proteins bind 5mCpG dinucleo-
tides regardless of sequence context. These proteins re-
cruit a variety of HDAC complexes and chromatin re-
modeling factors, leading to chromatin compaction, and,
consequently, to transcriptional repression (Klose and
Bird 2006). In mammals, MeCP1 and MeCP2 (methyl-
CpG-binding proteins 1 and 2) were the first protein or
complexes identified with high affinity for methylated
DNA (Meehan et al. 1989). Other mammalian methyl-
CpG-binding proteins have been identified subse-
quently, on the basis of the ability to bind methylated
sequences in vitro and/or the possession of a conserved
MBD motif, including MBD2, MBD4, and Kaiso (which
lacks an MBD) (Hendrich and Bird 1998; Prokhortchouk
et al. 2001). The biological importance of MBD proteins
is demonstrated by the wide range of severe phenotypes
that result when genes encoding these proteins are mu-
tated (Amir et al. 1999; Hendrich et al. 2001). In contrast
to the considerable data on the role of mammalian meth-
ylcytosine-binding proteins, knowledge of plant methyl-
cytosine-binding proteins is very limited. Arabidopsis
contains 13 genes encoding MBD motifs. Three Arabi-
dopsis MBD proteins (MBD5, MBD6, and MBD7) have
been shown to bind symmetrically methylated CpG
sites in vitro and two reports indicate that Arabidopsis
MBD5 can also bind 5mCpHpH (Ito et al. 2003; Scebba et
al. 2003; Zemach and Grafi 2003). The role of these MBD
proteins in plant epigenetic regulation has not yet been
characterized.
VIM1 has several interesting features as a methylcy-
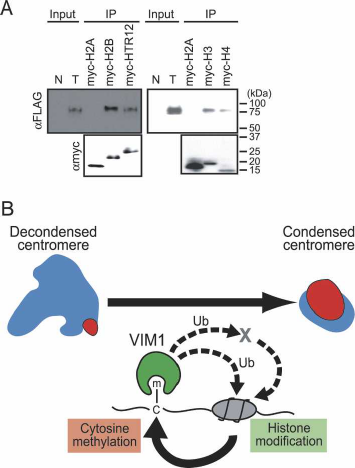
Figure 6. VIM1 at the DNA methylation–chromatin interface.
(A) VIM1 interacts with histones. (Top panel) Immobilized myc-
tagged histones were incubated with protein extracts from
transgenic plants expressing Flag-VIM1, and the associated pro-
teins were collected and detected by Western blot analysis using
an anti-Flag antibody to detect Flag-VIM1. Input lanes corre-
spond to total protein from either nontransgenic (N) or trans-
genic (T) plants (1/240×). The bottom panel shows detection of
myc-tagged histones in the samples using an anti-myc antibody.
(B) A model for the role of VIM1 in the maintenance of centro-
meric heterochromatin. The centromere decondensation phe-
notype is illustrated on the top left. (Blue) 180-bp centromere
repeat; (red) HTR12. The VIM1 biochemical activities illus-
trated at the bottom of the figure are necessary for centromere
condensation (top right). Demonstrated VIM1 activities include
direct binding of methylcytosine (m–C) and histone interaction
(direct or indirect). VIM1 is proposed to act as a ubiquitin (Ub)
ligase to modify centromere chromatin structure through direct
modification of nucleosomal histones or through modification
of one or more chromatin effector proteins (X). An alteration in
centromeric chromatin structure leads to DNA hypomethyl-
ation and decondensation of the centromere repeats.
5mC-binding protein required for DNA methylation
GENES & DEVELOPMENT 7
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
tosine-binding protein. First, VIM1 lacks an MBD do-
main and binds methylcytosine through a region encom-
passing the SRA domain. We note that the SRA domain
in murine Np95 and related mammalian proteins is suf-
ficient for 5mCpG binding (Unoki et al. 2004), but the
region in VIM1 required for methylcytosine-binding in-
cludes the SRA domain and an additional segment (up to
61 amino acids) adjacent to this domain. Second, VIM1 is
able to bind oligonucleotides that contain a single
5mCpG site. Third, VIM1 interacts with both 5mCpG
and 5mCpHpG. This is the first report of a methylcyto-
sine-binding protein able to bind 5mCpHpG, suggesting
that VIM1 may play an important role in interpreting
one of the non-CpG methylation systems found in
plants.
Differences between VIM1 and mammalian proteins
sharing structural similarity
VIM1 is a methylcytosine-binding protein that has a
PHD domain, two RING finger domains, and an SRA
domain. Proteins showing structural and amino acid
similarity to VIM1 are found in animals as well as plants.
The mammalian proteins most closely related to VIM1,
human ICBP90 and NIRF, along with murine Np95, ap-
pear to function as transcriptional regulators tied into
both cell cycle control and DNA repair (Hopfner et al.
2000; Bonapace et al. 2002; Muto et al. 2002; Arima et al.
2004). Despite their similarities, VIM1 shows several
distinct differences from these mammalian proteins.
First, the mammalian proteins have only one RING do-
main and an extra ubiquitin-like domain, and amino acid
similarity between VIM1 and the mammalian proteins is
restricted to the SRA domain. Second, VIM1 is associ-
ated with a range of heavily methylated genomic se-
quences, suggesting that VIM1 could play a role in main-
taining heterochromatin throughout the genome. In con-
trast, ICBP90 regulates the transcription of specific
genes, such as topoisomerase-II␣ and Retinoblastoma
protein 1, as well as participates in the maintenance of
silencing of tumor suppressor genes when they become
ectopically methylated (Hopfner et al. 2000; Unoki et al.
2004; Jeanblanc et al. 2005). Third, VIM1 is enriched at
chromocenters, but Np95 and ICBP90 are broadly local-
ized in the nucleus during interphase; Np95 is globally
associated with chromatin (Miura et al. 2001), whereas
ICBP90 is distributed throughout the entire nucleoplasm
(Arima et al. 2004; Jeanblanc et al. 2005). Fourth, loss of
Np95 or ICBP90 function has not yet been reported to
affect DNA methylation of their target sequences.
Model for VIM1 action
Our working model is that VIM1 acts as a bridge be-
tween cytosine methylation and histone modification in
a mutually reinforcing heterochromatin signaling loop
that is required for proper centromere organization and
centromere DNA methylation (Fig. 6B). VIM1 interacts
with core histones and is able to bind 5mC-modified
DNA, placing it physically at the interface between
DNA methylation and chromatin packaging. The possi-
bility that VIM1 is involved in histone modification is
suggested by the demonstration that the SRA domain of
ICBP90, Np95, and NIRF can recruit mammalian
HDAC1 (Unoki et al. 2004). In addition, Np95, via its
RING domain, possesses histone monoubiquitin ligase
activity (Citterio et al. 2004). Although histone ubiqui-
tylation is often associated with transcriptional activa-
tion, there are examples where this modification corre-
lates with transcriptionally silent heterochromatin
(Emre and Berger 2004; Wang et al. 2004). We note the
recent discovery that a cullin 4-dependent E3 ubiquitin
ligase, complexed with Rik1 and the HMT Clr4, plays a
crucial role in formation of centromeric heterochroma-
tin in fission yeast (Horn et al. 2005; Jia et al. 2005; Li et
al. 2005; Thon et al. 2005). The in vivo substrates of the
Rik1-associated E3 ubiquitin ligase complex are not
known, but histone H2B copurified with the Rik1 com-
plex, and this complex exhibited H2B polyubiquitylation
activity in vitro (Horn et al. 2005).
Based on these considerations, we propose that VIM1
binds to methylated centromeric DNA and alters his-
tone modification in the centromeric chromatin by re-
cruiting HDACs and/or by ubiquitylation of core his-
tones. The demonstrated interaction with VIM1 and his-
tones H2B, H3, H4, and HTR12 makes these core
histones good candidates for substrates of RING-domain
mediated ubiquitylation. Alternatively, VIM1 might
ubiquitylate a chromatin effector (or effectors), leading
to an alteration of centromeric heterochromatin. Alter-
ations in centromeric chromatin are postulated to lead to
a reduction in centromeric DNA methylation due to the
inefficient recruitment of DNMTs, such as CMT3 (re-
cruited to heterochromatic histone methylation marks)
(Lindroth et al. 2004) and DRM2 (which contains ubiq-
uitin-binding domains) (Cao et al. 2000). Our results sug-
gest that this new class of methylcytosine-binding pro-
teins may play a different role in epigenetic regulation
compared with MBD-class methylcytosine-binding pro-
teins, which have not been demonstrated to participate
in maintenance of genomic cytosine methylation pat-
terns or centromeric heterochromatin.
Materials and methods
Plant material
Seeds of natural Arabidopsis strains and T-DNA insertion mu-
tants (Alonso et al. 2003) were obtained from the Arabidopsis
Biological Resources Center. The natural strains examined were
part of the 96-member collection assembled by Nordborg et al.
(2005). We also examined the original collection of Bor acces-
sions made by J. Relichová (Masaryk University, Brno, Czech
Republic) to confirm the presence to the vim1-1 allele in natural
isolates. Plants were grown in a controlled environmental
chamber at 22°C under long day conditions (16 h light per day).
DNA blot hybridization
Genomic DNA was digested with HpaII, MspI, or HhaI accord-
ing to the manufacturer’s (New England Biolabs) instructions.
Woo et al.
8 GENES & DEVELOPMENT
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
Radiolabeled probes were generated by random priming, and
blots were prepared and hybridized using standard methods.
The following probes were generated from purified cloned in-
serts: 180-bp repeat (CEN) clone, pARR20-1 (Vongs et al. 1993),
and 5S rRNA gene clone pCT4.1 (Campell et al. 1992). The
Athila LTR and FWA probes were derived by PCR from genomic
DNA (Soppe et al. 2000; Lindroth et al. 2001).
Construction of plant expression vectors and generation
of transgenic plants
The 5.5-kb Col genomic DNA fragment containing 2.0 kb of the
promoter region and the whole predicted ORF of VIM1 was
PCR-amplified using two primers: 5⬘-CACCACGACCTTCG
GAAGATGCAAGAA-3⬘ and 5⬘-CCTGATGGTCGCAGAAA
CTGTTGC-3⬘. The fragment was cloned into pENTR-D TOPO
(Invitrogen) and the resulting VIM1 insert was recombined into
pEarlyGate302 (Earley et al. 2006). This construct was trans-
formed into Agrobacterium tumefaciens (LBA4404) and was in-
troduced into Bor-4 plants by in planta transformation (Bechtold
and Pelletier 1998). A full-length VIM1 cDNA clone in
pENTR-D TOPO was recombined into pEarlyGate202 (Earley et
al. 2006) to add an N-terminal Flag tag for the ChIP and histone-
binding assays. The resulting constructs were introduced into
Col plants by standard infiltration protocols.
EMSA
Full-length VIM1 cDNA PCR product (amino acids 1–645) or
deletion mutants—designated PHD/RING1 (amino acids
1–237), SRA/RING2 (amino acids 238–645), SRA (amino acids
238–456), RING2 (amino acids 457–645), and SRA+ (amino ac-
ids 238–517)—were cloned into the pDEST17 vector (Invitro-
gen). We synthesized VIM1 or derivatives in vitro using a T7
RNA polymerase-coupled wheat germ extract system (Promega
T
N
T
R
). For binding assays, double-stranded probe DNA was
end-labeled using the DIG gel shift kit (Roche) according to the
manufacturer’s instructions. Labeled DNA (62 fmol) were incu-
bated with in vitro translated VIM1 or its derivatives in the
presence or absence of competitors (3.85 pmol). Binding reac-
tions were carried out in 25 mM HEPES (pH 7.6), 50 mM KCl,
0.1 mM EDTA (pH 8.0), 12.5 mM MgCl
2
, 1 mM DTT, and 5%
(w/v) glycerol; 1.5 µg poly-dIdC was added as a nonspecific com-
petitor. After incubation for 30 min at room temperature, the
reaction mixture was subjected to electrophoresis.
ChIP
ChIP was performed as described in Grendel et al. (2002) on
nuclei prepared from wild-type and Arabidopsis plants express-
ing Flag-VIM1 using anti-Flag (Sigma). Precipitated DNAs were
subjected to PCR using the following primers (see Supplemen-
tary Table 2): CEN-F + CEN-R; 5S rRNA-F + 5S rRNA-R;
Athila-F + Athila-R; T5L23.29-F + T5L23.29-R; At4g04040-
F + At4g04040-R. Amplification of CEN, 5S rRNA, Athila,or
At4g04040 was performed for 26, 22, 30, or 32 cycles, respec-
tively.
Immunolocalization/DNA-FISH
Root meristems from 14-d-old plants were excised and nuclei
were extracted as described previously (Lawrence et al. 2004).
After post-fixation in 4% formaldehyde/PBS (phosphate-buff-
ered saline), washes in PBS, and blocking at 37°C, slides were
exposed overnight to primary antisera mouse anti-Flag (1:100,
Sigma) and rabbit anti-HTR12 (Talbert et al. 2002) in PBS and
0.5% blocking reagent (Roche). After washes in PBS, slides were
incubated at 37°C with anti-mouse-FITC diluted 1:100 (Sigma)
or goat anti-rabbit-TRITC (1:300, Sigma). For combined protein/
DNA-FISH localization experiments, slides were first subjected
to immunolocalization, and then post-fixed in 4% formalde-
hyde/PBS followed by in situ hybridization. FISH using the CEN
repeat probe (pARR20-1) labeled with biotin-dUTP and DIG-
dUTP was performed as described (Pontes et al. 2004). CEN
repeats were detected using either goat anti-biotin conjugated
with avidin (1:200, Vector Laboratories) followed by streptavi-
din-Alexa 543 (Molecular Probes) or anti-digoxeninin (1:200,
Roche) and rabbit anti-mouse Alexa 488 (Molecular Probes).
DNA was counterstained with DAPI (1 µg/mL) in Vectashield
(Vector Laboratories). Nuclei were examined using a Nikon
Eclipse E800i epifluorescence microscope, with images col-
lected using a Photometrics Coolsnap ES Mono digital camera.
The images were pseudocolored, merged, and processed using
Adobe Photoshop (Adobe Systems).
VIM1–histone interaction
Epitope-tagged (myc) recombinant histones were expressed in E.
coli BL21(AI) (Invitrogen) and cells were lysed by sonication in
a buffer containing 500 mM NaCl and 20 mM Tris-HCl (pH 7.5).
For purification, anti-c-Myc agarose conjugate (Sigma) was
added to cleared lysates and incubated for2hat4°C. Beads were
washed in buffer containing 150 mM NaCl, 20 mM Tris-HCl
(pH 7.5), 5 mM MgCl
2
, and 0.5% NP-40. Total proteins were
extracted from transgenic plants overexpressing Flag-VIM1 by
grinding tissue in an extraction buffer containing 150 mM
NaCl, 20 mM Tris-HCl (pH 7.5), and 5 mM MgCl
2
. Bead-bound
myc-tagged histones were incubated with extracts from trans-
genic plants overexpressing Flag-VIM1 overnight at 4°C. Beads
were washed as described above. Proteins were eluted by adding
SDS-PAGE sample buffer and were analyzed by SDS-PAGE elec-
trophoresis followed by Western blotting using anti-myc or
anti-Flag antibodies (Sigma).
Acknowledgments
We thank T. Dittmer, H.-f. Kuo, S. Rangwala, T.R. Smith, and
H. Yi for comments on the manuscript. We are grateful to J.
Relichová for providing Bor natural accessions, and J. Haag (his-
tone constructs) and S. Henikoff (anti-HTR12 antibody) for pro-
viding reagents. This work was supported by grants from the
National Science Foundation to E.J.R (MCB-0321990) and the
Monsanto Company. DNA-FISH and immunolocalization was
performed by O.P., who was supported by fellowship SFRH/
BPD/17508/2004 from the FundaçãoparaaCiência e Tecnolo-
gia (Portugal) and by NIH grants R01GM60380 and
R01GM077590 to C.S.P. We thank the Arabidopsis Biological
Resource Center at The Ohio State University for providing
Arabidopsis strains and SALK T-DNA insertion mutants.
References
Alonso, J.M., Stepanova, A.N., Leisse, T.J., Kim, C.J., Chen, H.,
Shinn, P., Stevenson, D.K., Zimmerman, J., Barajas, P.,
Cheuk, R., et al. 2003. Genome-wide insertional mutagen-
esis of Arabidopsis thaliana. Science 301: 653–657.
Amir, R.E., Van den Veyver, I.B., Wan, M., Tran, C.Q., Francke,
U., and Zoghbi, H.Y. 1999. Rett syndrome is caused by mu-
tations in X-linked MECP2, encoding methyl-CpG-binding
protein 2. Nat. Genet. 23: 185–188.
5mC-binding protein required for DNA methylation
GENES & DEVELOPMENT 9
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
Arima, Y., Hirota, T., Bronner, C., Mousli, M., Fujiwara, T.,
Niwa, S., Ishikawa, H., and Saya, H. 2004. Down-regulation
of nuclear protein ICBP90 by p53/p21Cip1/WAF1-dependent
DNA-damage checkpoint signals contributes to cell cycle
arrest at G1/S transition. Genes Cells 9: 131–142.
Bechtold, N. and Pelletier, G. 1998. In planta Agrobacterium-
mediated transformation of adult Arabidopsis thaliana
plants by vacuum infiltration. Methods Mol. Biol. 82: 259–
266.
Bonapace, I.M., Latella, L., Papait, R., Nicassio, F., Sacco, A.,
Muto, M., Crescenzi, M., and Di Fiore, P.P. 2002. Np95 is
regulated by E1A during mitotic reactivation of terminally
differentiated cells and is essential for S phase entry. J. Cell
Biol. 157: 909–914.
Campell, B.R., Song, Y., Posch, T.E., Cullis, C.A., and Town,
C.D. 1992. Sequence and organization of 5S ribosomal RNA-
encoding genes of Arabidopsis thaliana. Gene 112: 225–228.
Cao, X., Springer, N.M., Muszynski, M.G., Phillips, R.L., Kaepp-
ler, S., and Jacobsen, S.E. 2000. Conserved plant genes with
similarity to mammalian de novo DNA methyltransferases.
Proc. Natl. Acad. Sci. 97: 4979–4984.
Chan, S.W., Henderson, I.R., and Jacobsen, S.E. 2005. Gardening
the genome: DNA methylation in Arabidopsis thaliana.
Nat. Rev. Genet. 6: 351–360.
Citterio, E., Papait, R., Nicassio, F., Vecchi, M., Gomiero, P.,
Mantovani, R., Di Fiore, P.P., and Bonapace, I.M. 2004. Np95
is a histone-binding protein endowed with ubiquitin ligase
activity. Mol. Cell. Biol. 24: 2526–2535.
Craig, J.M. 2005. Heterochromatin—Many flavours, common
themes. Bioessays 27: 17–28.
Earley, K.W., Haag, J.R., Pontes, O., Opper, K., Juehne, T., Song,
K., and Pikaard, C.S. 2006. Gateway-compatible vectors for
plant functional genomics and proteomics. Plant J. 45: 616–
629.
Emre, N.C. and Berger, S.L. 2004. Histone H2B ubiquitylation
and deubiquitylation in genomic regulation. Cold Spring
Harbor Symp. Quant. Biol. 69: 289–299.
Fujimori, A., Matsuda, Y., Takemoto, Y., Hashimoto, Y., Kubo,
E., Araki, R., Fukumura, R., Mita, K., Tatsumi, K., and Muto,
M. 1998. Cloning and mapping of Np95 gene which encodes
a novel nuclear protein associated with cell proliferation.
Mamm. Genome 9: 1032–1035.
Fuks, F., Hurd, P.J., Deplus, R., and Kouzarides, T. 2003. The
DNA methyltransferases associate with HP1 and the
SUV39H1 histone methyltransferase. Nucleic Acids Res. 31:
2305–2312.
Gendrel, A.V., Lippman, Z., Yordan, C., Colot, V., and Martiens-
sen, R.A. 2002. Dependence of heterochromatic histone H3
methylation patterns on the Arabidopsis gene DDM1. Sci-
ence 297: 1871–1873.
Genereux, D.P., Miner, B.E., Bergstrom, C.T., and Laird, C.D.
2005. A population-epigenetic model to infer site-specific
methylation rates from double-stranded DNA methylation
patterns. Proc. Natl. Acad. Sci. 102: 5802–5807.
Goll, M.G. and Bestor, T.H. 2005. Eukaryotic cytosine methyl-
transferases. Annu. Rev. Biochem. 74: 481–514.
Hendrich, B. and Bird, A. 1998. Identification and characteriza-
tion of a family of mammalian methyl-CpG binding pro-
teins. Mol. Cell. Biol. 18: 6538–6547.
Hendrich, B., Guy, J., Ramsahoye, B., Wilson, V.A., and Bird, A.
2001. Closely related proteins MBD2 and MBD3 play dis-
tinctive but interacting roles in mouse development. Genes
& Dev. 15: 710–723.
Henikoff, S. and Dalal, Y. 2005. Centromeric chromatin: What
makes it unique? Curr. Opin. Genet. Dev. 15: 177–184.
Hopfner, R., Mousli, M., Jeltsch, J.M., Voulgaris, A., Lutz, Y.,
Marin, C., Bellocq, J.P., Oudet, P., and Bronner, C. 2000.
ICBP90, a novel human CCAAT binding protein, involved in
the regulation of topoisomerase II␣ expression. Cancer Res.
60: 121–128.
Horn, P.J., Bastie, J.N., and Peterson, C.L. 2005. A Rik1-associ-
ated, cullin-dependent E3 ubiquitin ligase is essential for
heterochromatin formation. Genes & Dev. 19: 1705–1714.
Huisinga, K.L., Brower-Toland, B., and Elgin, S.C. 2006. The
contradictory definitions of heterochromatin: Transcription
and silencing. Chromosoma 115: 110–122.
Ito, M., Koike, A., Koizumi, N., and Sano, H. 2003. Methylated
DNA-binding proteins from Arabidopsis. Plant Physiol. 133:
1747–1754.
Jaenisch, R. and Bird, A. 2003. Epigenetic regulation of gene
expression: How the genome integrates intrinsic and envi-
ronmental signals. Nat. Genet. 33 (Suppl.): 245–254.
Jeanblanc, M., Mousli, M., Hopfner, R., Bathami, K., Martinet,
N., Abbady, A.Q., Siffert, J.C., Mathieu, E., Muller, C.D., and
Bronner, C. 2005. The retinoblastoma gene and its product
are targeted by ICBP90: A key mechanism in the G1/S tran-
sition during the cell cycle. Oncogene 24: 7337–7345.
Jenuwein, T. and Allis, C.D. 2001. Translating the histone code.
Science 293: 1074–1080.
Jia, S., Kobayashi, R., and Grewal, S.I. 2005. Ubiquitin ligase
component Cul4 associates with Clr4 histone methyltrans-
ferase to assemble heterochromatin. Nat. Cell Biol. 7: 1007–
1013.
Kakutani, T., Munakata, K., Richards, E.J., and Hirochika, H.
1999. Meiotically and mitotically stable inheritance of DNA
hypomethylation induced by ddm1 mutation of Arabidopsis
thaliana. Genetics 151: 831–838.
Kankel, M.W., Ramsey, D.E., Stokes, T.L., Flowers, S.K., Haag,
J.R., Jeddeloh, J.A., Riddle, N.C., Verbsky, M.L., and Rich-
ards, E.J. 2003. Arabidopsis MET1 cytosine methyltransfer-
ase mutants. Genetics 163: 1109–1122.
Klose, R.J. and Bird, A.P. 2006. Genomic DNA methylation:
The mark and its mediators. Trends Biochem. Sci. 31: 89–97.
Lawrence, R.J., Earley, K., Pontes, O., Silva, M., Chen, Z.J.,
Neves, N., Viegas, W., and Pikaard, C.S. 2004. A concerted
DNA methylation/histone methylation switch regulates
rRNA gene dosage control and nucleolar dominance. Mol.
Cell 13: 599–609.
Lehnertz, B., Ueda, Y., Derijck, A.A., Braunschweig, U., Perez-
Burgos, L., Kubicek, S., Chen, T., Li, E., Jenuwein, T., and
Peters, A.H. 2003. Suv39h-mediated histone H3 lysine 9
methylation directs DNA methylation to major satellite re-
peats at pericentric heterochromatin. Curr. Biol. 13: 1192–
1200.
Li, F., Goto, D.B., Zaratiegui, M., Tang, X., Martienssen, R., and
Cande, W.Z. 2005. Two novel proteins, dos1 and dos2, in-
teract with rik1 to regulate heterochromatic RNA interfer-
ence and histone modification. Curr. Biol. 15: 1448–1457.
Lindroth, A.M., Cao, X., Jackson, J.P., Zilberman, D., McCal-
lum, C.M., Henikoff, S., and Jacobsen, S.E. 2001. Require-
ment of CHROMOMETHYLASE3 for maintenance of
CpXpG methylation. Science 292: 2077–2080.
Lindroth, A.M., Shultis, D., Jasencakova, Z., Fuchs, J., Johnson,
L., Schubert, D., Patnaik, D., Pradhan, S., Goodrich, J., Sch-
ubert, I., et al. 2004. Dual histone H3 methylation marks at
lysines 9 and 27 required for interaction with CHROMO-
METHYLASE3. EMBO J. 23: 4286–4296.
Lippman, Z., May, B., Yordan, C., Singer, T., and Martienssen,
R. 2003. Distinct mechanisms determine transposon inher-
itance and methylation via small interfering RNA and his-
tone modification. PLoS Biol. 1: E67.
Lippman, Z., Gendrel, A.V., Black, M., Vaughn, M.W., Dedhia,
Woo et al.
10 GENES & DEVELOPMENT
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
N., McCombie, W.R., Lavine, K., Mittal, V., May, B., Kas-
schau, K.D., et al. 2004. Role of transposable elements in
heterochromatin and epigenetic control. Nature 430: 471–
476.
Matzke, M.A. and Birchler, J.A. 2005. RNAi-mediated pathways
in the nucleus. Nat. Rev. Genet. 6: 24–35.
Meehan, R.R., Lewis, J.D., McKay, S., Kleiner, E.L., and Bird,
A.P. 1989. Identification of a mammalian protein that binds
specifically to DNA containing methylated CpGs. Cell 58:
499–507.
Miura, M., Watanabe, H., Sasaki, T., Tatsumi, K., and Muto, M.
2001. Dynamic changes in subnuclear NP95 location during
the cell cycle and its spatial relationship with DNA replica-
tion foci. Exp. Cell Res. 263: 202–208.
Mori, T., Li, Y., Hata, H., Ono, K., and Kochi, H. 2002. NIRF, a
novel RING finger protein, is involved in cell-cycle regula-
tion. Biochem. Biophys. Res. Commun. 296: 530–536.
Muto, M., Kanari, Y., Kubo, E., Takabe, T., Kurihara, T., Fuji-
mori, A., and Tatsumi, K. 2002. Targeted disruption of Np95
gene renders murine embryonic stem cells hypersensitive to
DNA damaging agents and DNA replication blocks. J. Biol.
Chem. 277: 34549–34555.
Nightingale, K.P., O’Neill, L.P., and Turner, B.M. 2006. Histone
modifications: Signalling receptors and potential elements
of a heritable epigenetic code. Curr. Opin. Genet. Dev. 16:
125–136.
Nordborg, M., Hu, T.T., Ishino, Y., Jhaveri, J., Toomajian, C.,
Zheng, H., Bakker, E., Calabrese, P., Gladstone, J., Goyal, R.,
et al. 2005. The pattern of polymorphism in Arabidopsis
thaliana. PLoS Biol. 3: e196.
Pelissier, T., Tutois, S., Deragon, J.M., Tourmente, S., Genes-
tier, S., and Picard, G. 1995. Athila, a new retroelement from
Arabidopsis thaliana. Plant Mol. Biol. 29: 441–452.
Pontes, O., Neves, N., Silva, M., Lewis, M.S., Madlung, A., Co-
mai, L., Viegas, W., and Pikaard, C.S. 2004. Chromosomal
locus rearrangements are a rapid response to formation of the
allotetraploid Arabidopsis suecica genome. Proc. Natl.
Acad. Sci. 101: 18240–18245.
Prokhortchouk, A., Hendrich, B., Jorgensen, H., Ruzov, A.,
Wilm, M., Georgiev, G., Bird, A., and Prokhortchouk, E.
2001. The p120 catenin partner Kaiso is a DNA methylation-
dependent transcriptional repressor. Genes & Dev. 15: 1613–
1618.
Rangwala, S.H. and Richards, E.J. 2004. The value-added ge-
nome: Building and maintaining genomic cytosine methyl-
ation landscapes. Curr. Opin. Genet. Dev. 14: 686–691.
Richards, E.J. 2006. Inherited epigenetic variation—Revisiting
soft inheritance. Nat. Rev. Genet. 7: 395–401.
Saze, H., Scheid, O.M., and Paszkowski, J. 2003. Maintenance of
CpG methylation is essential for epigenetic inheritance dur-
ing plant gametogenesis. Nat. Genet. 34: 65–69.
Scebba, F., Bernacchia, G., De Bastiani, M., Evangelista, M.,
Cantoni, R.M., Cella, R., Locci, M.T., and Pitto, L. 2003.
Arabidopsis MBD proteins show different binding specifici-
ties and nuclear localization. Plant Mol. Biol. 53: 715–731.
Soppe, W.J., Jacobsen, S.E., Alonso-Blanco, C., Jackson, J.P.,
Kakutani, T., Koornneef, M., and Peeters, A.J. 2000. The late
flowering phenotype of fwa mutants is caused by gain-of-
function epigenetic alleles of a homeodomain gene. Mol.
Cell 6: 791–802.
Soppe, W.J., Jasencakova, Z., Houben, A., Kakutani, T., Meister,
A., Huang, M.S., Jacobsen, S.E., Schubert, I., and Fransz, P.F.
2002. DNA methylation controls histone H3 lysine 9 meth-
ylation and heterochromatin assembly in Arabidopsis.
EMBO J. 21: 6549–6559.
Talbert, P.B., Masuelli, R., Tyagi, A.P., Comai, L., and Henikoff,
S. 2002. Centromeric localization and adaptive evolution of
an Arabidopsis histone H3 variant. Plant Cell 14:
1053–1066.
Tamaru, H. and Selker, E.U. 2001. A histone H3 methyltrans-
ferase controls DNA methylation in Neurospora crassa. Na-
ture 414: 277–283.
Thon, G., Hansen, K.R., Altes, S.P., Sidhu, D., Singh, G., Ver-
hein-Hansen, J., Bonaduce, M.J., and Klar, A.J. 2005. The
Clr7 and Clr8 directionality factors and the Pcu4 cullin me-
diate heterochromatin formation in the fission yeast Schizo-
saccharomyces pombe. Genetics 171: 1583–1595.
Unoki, M., Nishidate, T., and Nakamura, Y. 2004. ICBP90, an
E2F-1 target, recruits HDAC1 and binds to methyl-CpG
through its SRA domain. Oncogene 23: 7601–7610.
Vongs, A., Kakutani, T., Martienssen, R.A., and Richards, E.J.
1993. Arabidopsis thaliana DNA methylation mutants. Sci-
ence 260: 1926–1928.
Wang, H., Wang, L., Erdjument-Bromage, H., Vidal, M., Tempst,
P., Jones, R.S., and Zhang, Y. 2004. Role of histone H2A
ubiquitination in Polycomb silencing. Nature 431: 873–878.
Zemach, A. and Grafi, G. 2003. Characterization of Arabidopsis
thaliana methyl-CpG-binding domain (MBD) proteins.
Plant J. 34: 565–572.
5mC-binding protein required for DNA methylation
GENES & DEVELOPMENT 11
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from

10.1101/gad.1512007Access the most recent version at doi:
published online January 22, 2007Genes Dev.
Hye Ryun Woo, Olga Pontes, Craig S. Pikaard, et al.
heterochromatinization
VIM1, a methylcytosine-binding protein required for centromeric
Material
Supplemental
https://genesdev.cshlp.org/content/suppl/2007/01/22/gad.1512007.DC1
Published online January 22, 2007 in advance of the full issue.
License
Service
Email Alerting
click here.right corner of the article or
Receive free email alerts when new articles cite this article - sign up in the box at the top
Published by Copyright © 2007, Cold Spring Harbor Laboratory Press
Cold Spring Harbor Laboratory Presson September 17, 2024 - Published by Downloaded from
