Full Terms & Conditions of access and use can be found at
https://www.tandfonline.com/action/journalInformation?journalCode=iann20
Annals of Medicine
ISSN: (Print) (Online) Journal homepage: https://www.tandfonline.com/loi/iann20
Gastrointestinal microbiome and gluten in celiac
disease
Xingxing Wu, Lin Qian, Kexin Liu, Jing Wu & Zhaowei Shan
To cite this article: Xingxing Wu, Lin Qian, Kexin Liu, Jing Wu & Zhaowei Shan (2021)
Gastrointestinal microbiome and gluten in celiac disease, Annals of Medicine, 53:1, 1797-1805,
DOI: 10.1080/07853890.2021.1990392
To link to this article: https://doi.org/10.1080/07853890.2021.1990392
© 2021 The Author(s). Published by Informa
UK Limited, trading as Taylor & Francis
Group.
Published online: 14 Oct 2021.
Submit your article to this journal
Article views: 6941
View related articles
View Crossmark data
Citing articles: 8 View citing articles
REVIEW ARTICLE
Gastrointestinal microbiome and gluten in celiac disease
Xingxing Wu
a
, Lin Qian
a
, Kexin Liu
a
, Jing Wu
b
and Zhaowei Shan
a
a
Affiliated Hospital of Nanjing University of Chinese Medicine, Nanjing, China;
b
Institute of Chinese Medicine, Nanjing Drum Tower
Hospital, Nanjing University, Drum Tower Clinical Medicine College of Nanjing University of Chinese Medicine, Nanjing, China
ABSTRACT
Coeliac disease (CD), also known as gluten sensitive enteropathy, is an autoimmune intestinal
disease induced by gluten in genetically susceptible individuals. Gluten is a common ingredient
in daily diet and is one of the main environmental factors to induce coeliac disease. Adhering
to gluten free diet (GFD) is an effective method for treating CD. Microbiota plays an extremely
important role in maintaining human health, and diet is the main factor to regulate the com-
position and function of gut microbiota. Recent studies have shown that gluten metabolism is
closely related to gastrointestinal tract (GIT) microbiota. With the increasing prevalence of coel-
iac disease, there is a need for alternative treatments to GFD. In this review, biological medica-
tion of gluten, relationship between gluten and gut microflora, effect of GFD on GIT microflora,
and effect of probiotics on CD were reviewed. By analysing the research progress on relation-
ship between gluten and gastrointestinal microbiome in coeliac disease, this review tried to
explore the prospective and potential mechanism of microecological agents in treating coel-
iac disease.
ARTICLE HISTORY
Received 19 May 2021
Revised 28 June 2021
Accepted 30 September 2021
KEYWORDS
Gluten; gastrointestinal
microbiome; coeliac disease;
gluten free diet; probiotics
1. Introduction
Coeliac disease is an autoimmune disorder that occurs
in genetically predisposed individuals, including adults
and children, who develop an immune reaction to glu-
ten. Even though this disease primarily affects the
small intestine, its clinical manifestations are broad,
with both intestinal and extra-intestinal symptoms.
There are multi-factors might affect this disease, such
as environmental, genetic factors and immune imbal-
ance [1]. The main clinical presentations include intes-
tinal symptoms, such as diarrhoea, abdominal
distension, abdominal pain; and extra-intestinal symp-
toms, such as anaemia, dermatitis herpetiformis, osteo-
penia and peripheral neuropathy. CD patients carry
specific susceptibility genes (HLA-DQ2, HLA-DQ8), but
their existence is not enough to cause the occurrence
of CD, which requires the participation of environmen-
tal factors-gluten [2]. As the consumption of gluten-
containing food increases, the incidence of related
autoimmune diseases has gradually increased (such as
CD, wheat allergic diseases) [3]. In genetically suscep-
tible individuals, gluten is one of the necessary condi-
tions for inducing CD. Gluten is a kind of protein
mainly existing in wheat, barley and rye, accounting
for 80%–85% of the total protein in wheat [4]. It is a
protein mixture composed of hundreds of monomers,
oligomers and polymers, mainly including gliadin and
glutenin. Among them, the main antigen protein caus-
ing CD is gliadin which is rich in glutamine and pro-
line, and cannot be digested by human digestive
enzymes and brush border peptidase [5–6]. Proline-
rich peptides are protected from proteolysis by gastric,
pancreatic and intestinal brush border membrane
enzymes, so they have an opportunity to build up to
high concentrations in the small intestine. However,
oral bacteria that secrete gliadin (gluten) degrading
enzymes had been identified. Their most active glia-
din-cleaving enzymes had also been identified and
purified [7–8]. Apart from interesting biological find-
ings, these bacteria and enzymes may lead to novel
and effective strategies to detoxify immunogenic glu-
ten peptides prior to their reaching the proximal small
intestine. Part of the gluten is hydrolysed by oral
microbial proteases in the oral cavity, thereby reduc-
ing its immunotoxicity. However, most gluten is hydro-
lysed by pepsin into high molecular weight peptides
CONTACT Jing Wu [email protected] Institute of Chinese Medicine, Nanjing Drum Tower Hospital, Nanjing University, Drum Tower Clinical
Medicine College of Nanjing University of Chinese Medicine, 321 Zhongshan Road, Nanjing 210008, China; Zhaowei Shan
Affiliated Hospital of Nanjing University of Chinese Medicine, 155 Han zhong Road, Nanjing 210029, China
This article has been republished with minor changes. These changes do not impact the academic content of the article.
ß 2021 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License ( http://creativecommons.org/licenses/by/4.0/), which permits
unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
ANNALS OF MEDICINE
2021, VOL. 53, NO. 1, 1797–1805
https://doi.org/10.1080/07853890.2021.1990392

in the stomach. The peptides that enter the small
intestine from the stomach are not easily degraded
due to their rich proline. They stay in the intestine for
a long time and increase the probability of triggering
immune response. A large number of immunogenic
polymer peptides gathered in the intestinal lumen,
mainly including immunodominant peptides (such as
P57-P89 peptide and 33 peptide in a-gliadin) and
non-immune dominant peptides (such as P31P43),
triggered the adaptive immune response mediated by
CD4 þ Th1 cells and the innate immune response
mediated by intraepithelial lymphocytes respectively,
and leaded to infiltration of intestinal epithelial inflam-
matory cells, villus atrophy and crypt hyperplasia [9].
Thus, it will lead to the destruction of intestinal epi-
thelial cells and the increase of intestinal permeability,
resulting in diarrhoea, abdominal distention, abdom-
inal pain, emaciation, dermatitis herpetiformis and
other clinical symptoms. Although the gluten-free diet
can significantly improve the clinical symptoms of
patients with CD, gluten-free diet is expensive and has
very few products. In order to have a good quality of
life, patients have to adhere to the gluten-free diet. It
not only brings a financial burden to society and
patients themselves, but also brings social and psycho-
logical impact to patients [10]. In addition to gluten,
the microbiota dysbiosis of the digestive tract flora
may be another environmental factor that triggers CD.
Studies have confirmed that patients with CD have
disorders of the digestive tract flora. The abundance
and diversity of beneficial commensals have
decreased, while, pathobionts have increased. Coeliac
disease is associated with intestinal dysbiosis charac-
terized by increases in pathobionts virulence features
[11–12]. Research also shown that, diet has a great
influence on the composition and function of intes-
tinal flora, and gluten can affect the stability of intes-
tinal flora. Therefore, this review will mainly focus on
the relationship between gluten and oral flora, intes-
tinal flora in coeliac disease. It will also pay special
attention to analyse perspectives and trends of probi-
otics in coeliac disease treatment.
2. Gluten and oral flora
2.1. Oral flora
So far, it has been found that there are more than
1,000 kinds of bacteria in the oral cavity inhabiting sal-
iva, teeth, gingiva and other different parts. There are
about 10
11
bacteria per gram of dental plaque and
10
8
bacteria per millilitre of saliva, so the oral cavity
becomes the second place where microorganisms are
densely colonized in digestive tract [13–14]. The latest
research shows that the oral symbiotic flora can
increase the immune function of the oral mucosa to
prevent the invasion of pathogenic microorganisms.
For example, oral microbiota genera Veillonella and
Streptococcus promote the production of anti-micro-
bial peptides and the secretion of inflammatory cyto-
kines, increasing the epithelial barrier function and
thickness characteristic of oral mucosa [15]. However,
other studies have found that some pathogenic bac-
teria in oral microflora are not only related to oral dis-
eases such as dental caries, periodontitis, and oral
ulcers [16], but also related to infective endocarditis
[17] and coronary atherosclerosis [18], pneumonia [19],
obesity [20], intestinal diseases, etc. The flora in the
oral cavity of healthy humans can be transported in
large quantities to the distal end of the digestive tract,
yet the translocation of oral species to the intestine is
considered a rare aberrant event, and a hallmark of
disease [21]. Certain oral microorganisms may induce
intestinal diseases in genetically susceptible individu-
als. For example, Atarashi et al. found that oral
Klebsiella colonises the colon and induces Th1 cell dif-
ferentiation to elicit a severe intestinal mucosal inflam-
mation [22].
2.2. Oral flora reduces the immunogenicity
of gliadin
At present time, most studies are confined largely to
explore the relationship between microflora and intes-
tinal diseases. However, the oral cavity is the first
digestive organ that comes into contact with food and
is correlated to digestive system diseases directly.
Therefore, on the basis of duodenal flora and colonic
flora studies, the salivary flora should be further ana-
lysed to improve description of the digestive tract
flora characteristics [23]. Patients with CD have oral
flora dysbiosis. There are microbial flora in the oral
cavity, which are related to the metabolism of gluten
in CD (Table 1). Although the food containing gluten
stays in the oral cavity for a short time, the number
and types of flora in the saliva are significantly greater
than those colonized in the stomach and duodenum.
The effect of oral flora on digesting gluten should not
be ignored. Researchers have found that the initial
metabolism of gliadin in the oral cavity may be related
to the genus of Rothia, Actinomyces, Neisseria, and
Streptococcus that colonized the oral cavity [24].
Compared with healthy people, the saliva of patients
with CD is rich in bacteria that could degrade gluten,
and the degradation rate of gluten is higher. The
1798 X. WU ET AL.

significant increase of Lactobacillus species may be
one of the reasons [23]. The protease-resistant highly
immunogenic 33-mer a-gliadin peptide could be com-
pletely degraded by dental plaque bacteria to reduce
immunogenicity [23,25]. However, there were studies
on the contrary standpoint, reported that oral micro-
bial enzymes degrade part of gluten, which in turn
increases immunogenic small molecule peptides epito-
pes and further induces intestinal inflammation [23].
3. Gluten and intestinal flora
3.1. Intestinal flora
As we mentioned before, the oral cavity is the second
place where microorganisms are densely colonized in
digestive tract. However, the gut is the most densely
colonized place of microflora in digestive tract. A
refined estimate showed that microflora in one human
body were in a ratio of 1.3:1 to human cells. It was
estimated that more than 1,000 kinds of microorgan-
isms live in the gut, the gut microbiome of healthy
people mainly includes Firmicutes, Bacteroides,
Proteobacteria, Actinomycetes . And some researchers
estimated that there were thousands of bacterial spe-
cies in the gastrointestinal tract [26–27]. According to
the interaction with the host, the intestinal flora is div-
ided into three categories: probiotics (such as
Lactobacillus, Bifidobacterium, etc.), pathobionts (such
as Clostridium, Enterococcus faecalis , etc.) and oppor-
tunistic pathogens. The intestinal flora of healthy peo-
ple can protect and maintain the intestinal barrier
function, promote the metabolism and absorption of
nutrients, regulate immunity, anti-aging, prevent can-
cer and suppress cancer, etc [28]. There is a mutually
symbiotic relationship between the flora and the host.
The host provides nutrients and the microenvironment
for the flora. The flora helps to maintain human intes-
tinal homeostasis by participating in a series of physio-
logical functions of the host. A large number of
studies have shown that once the balance between
intestinal microflora and the human body is broken, it
will lead to multiple systemic diseases, such as obesity,
diabetes, atherosclerosis, irritable bowel syndrome,
inflammatory bowel disease, and coeliac disease
through bile acid metabolism, brain-gut axis, intestinal
barrier, and immune system and so on [29].
3.2. Gliadin directly induces intestinal
flora dysbiosis
For the coeliac disease patients, the balance between
intestinal microflora and the human body could be bro-
ken by the gliadin. From the mouth and stomach, large
quantity of undegraded gliadin is being pushed into
the small intestine and large intestine, provides abun-
dant substrates for different bacteria in the intestinal
cavity, thereby promotes the reproduction of gliadin-
degrading bacteria and breaks the steady state of intes-
tinal flora [30–31]. At present time, the composition
and structure of the small intestinal flora are mainly
evaluated by detecting the abundance and diversity of
the duodenal flora. D’Argenio et al. tested the duo-
denal mucosal flora of patients with active CD and
found that the abundance of Proteobacteria increased,
while the abundance of Firmicutes and Actinobacteria
decreased. Compared with GFD patients and healthy
individuals, members of the Neisseria genus
(Betaproteobacteria class) were significantly more abun-
dant in active CD patients. Neisseria flavescens was the
most abundant Neisseria species in active CD duode-
num [32]. Sanchez E et al. found that compared with
children with GFD and healthy children, the duodenal-
mucosal bacteria of children with active CD (normal
gluten-containing food diet) has increased abundance
of Proteobacteria and decreased abundance of
Firmicutes at the phylum level; the abundance of
Enterobacteriaceae and Staphylococcaceae increased,
and Streptococcaceae decreased at family level [33]. In
the CD animal experiment, it was also found that the
intestinal flora was imbalanced. For example, compar-
ing the gluten-sensitive (GS) macaques with healthy
macaques, it was found that the alpha diversity
(Shannon diversity index) and abundance of the faecal
flora of the GS macaques were reduced, and it was
found that, two of the top eight families,
Table 1. The relationship between Oral flora and gluten in CD.
Substrate types Degradability Outcome
Salivary flora [23] Gluten The degradation rate of gluten is
higher than healthy people
unspecified
Rothia, Actinomyces, Neisseria, and
Streptococcus [24]
Gliadin unspecified unspecified
Dental plaque bacteria [23,25] Immunogenic 33-mer
a-gliadin peptide
Complete degradation Reduce immunogenicity
Oral microbial enzymes [23] Gluten Partial degradation Induce immunogenicity
ANNALS OF MEDICINE 1799

Streptococcaceae and Lactobacillaceae, were enriched in
GS macaques [34].
3.3. Intestinal flora promotes the hydrolysis
of gliadin
The human body lacks proteases, which are able to
completely digest gluten. The role of intestinal flora in
the process of such protein metabolism cannot be
ignored. The undegraded gliadin is transported from
the small intestine to the large intestine. Once it
enters the large intestine, it is in close contact with a
large number of microorganisms in the gut. Due to
the diversity of bacterial genes in large intestine and
their different biochemical pathways from the human
body, it makes certain intestinal microorganisms have
the ability to metabolize gliadin [35]. Researchers have
found that there are flora related to the metabolism
of gliadin in the human intestine (such as the genera
Lactobacillus, Streptococcus, Staphylococcus, Clostridium,
Bifidobacterium)[36]. These microorganisms not only
exist in the large intestine, but also in the small intes-
tine to metabolize gluten. Camino et al. showed that,
compared with the healthy group, the duodenal
mucosal flora of CD mice on a gluten-containing diet
had a higher proteolytic activity against gluten
(“glutenasic” activity), and it is related to the abun-
dance of Proteobacteria (including Pseudomonas)[37].
Herr
an et al. studied the small intestinal flora that
decomposes gliadin in healthy people and patients
with CD, and isolated 114 bacterial strains belonging
to 32 different species from the duodenal mucosa, of
which, 85 strains were able to grow in a medium con-
taining gluten as the sole nitrogen source. 31 strains
showed extracellular proteolytic activity against gluten
protein and 27 strains showed peptidolytic activity
towards the 33-mer peptide, an immunogenic peptide
for coeliac disease patients [38].
3.4. Gliadin combined with intestinal flora induces
intestinal inflammation
Obviously, researchers cannot determine that intestinal
flora dysbiosis is the result of coeliac disease or an
environmental factor for CD, or both. Conventionally,
gliadin was considered to activate innate immunity
and adaptive immunity, and activate intestinal inflam-
mation by inducing the production of cytokines and
chemokines. In particular, the gliadin is deamidated by
tissue transglutaminase in the lamina propria of the
small intestine, and binds to HLA class II DQ2/8 mole-
cules of antigen-presenting cells, activates T cells,
macrophages and dendritic cells, and secretes inflam-
matory cytokines. It follows the activation of the adap-
tive immune response through the production of anti-
endomysium, antigliadin, and anti-transglutaminase
antibodies by B cells that increase intestinal perme-
ability [39]. In addition to gliadin, intestinal microflora
also play an indispensable role in inducing inflamma-
tion in the intestinal mucosa of patients with CD. As
we all know, immune factors are one of the causes for
CD, and adaptive immunity plays an important role in
the pathogenesis of CD. Studies have found that the
intestinal flora is closely related to adaptive immunity,
and intestinal microflora have important regulatory
effects on the two embranchments of host adaptive
immunity, B cells and T cells. The intestinal flora can
promote the production of IgA in the intestine by reg-
ulating the B cells response; it can also maintain the
balance between intestinal inflammation and immune
tolerance by inducing the differentiation of intestinal
Th17 and Treg cells [40].
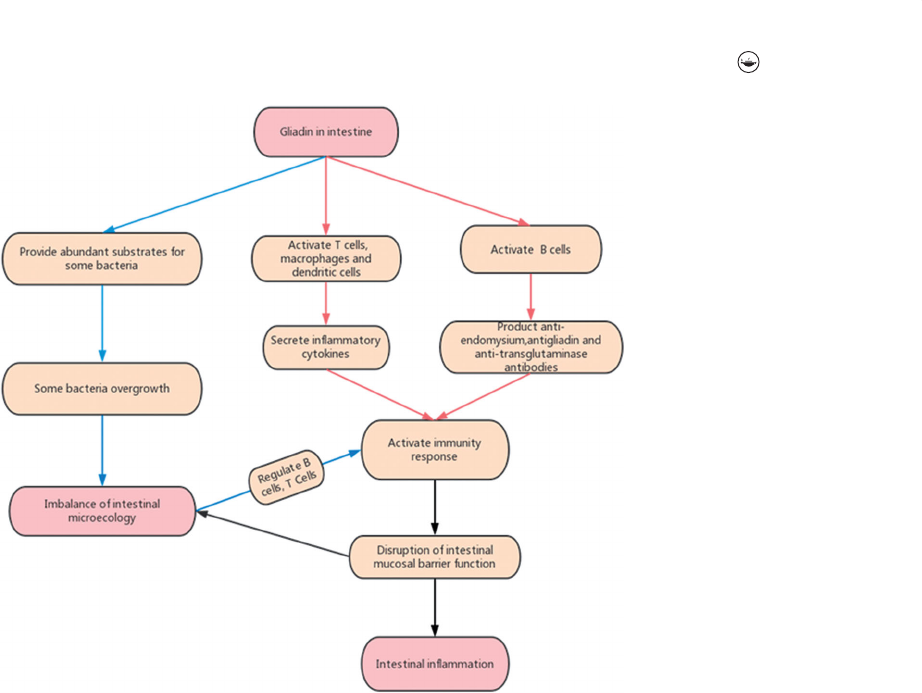
Gliadin evokes intestinal barrier dysfunction, which
leads to the excessive growth and translocation of intes-
tinal pathogenic bacteria, resulting in intestinal micro-
ecological imbalance; The microecological imbalance
activates the immune inflammatory response by regulat-
ing the B cell and T cell. Inflammatory factors can fur-
ther increase the permeability of the intestinal mucosa
by destroying the intestinal epithel ial cells and aggra-
vate coeliac disease [41]. The CD intestinal mucosal
immune response may directly destroy the biological
barrier, thereby affecting the microbial homeostasis. The
imbalanceoftheflora,ordysbiosisactsasapathogenic
factor to be counteractive at CD, thus forming a vicious
circle and continuing inflammation.
In CD, intestinal flora and gluten have a complex rela-
tionship (Figure 1). There are two essential different sit-
uations, one is the imbalance of intestinal microecology
caused by coeliac disease, the other is abnormal intes-
tinal flora, which is a co-factor of gliadin in inducing
coeliac disease. In CD patients, the abundance of
Firmicute s and Acti nobacte ria decreased , while the abun-
dance of Proteobacteria increased. The intestinal micro-
flora might be sometimes the cause and sometimes the
result, which needs analysis case by case. In the future,
reasonable research methods can be designed to con-
firm it. With the development of research, the role of lac-
tic acid bacteria can be further defined.
4. Gluten-free diet and digestive tract flora
CD patients have disorders of the digestive tract flora.
However, has the digestive tract flora of CD patients
1800 X. WU ET AL.
improved significantly after GFD treatment? Studies
have found that the digestive tract flora of CD
patients who are treated with GFD is still in an imbal-
anced state. A study confirmed that, compared with
the same number of healthy children, Bacteroidetes in
saliva of children with GFD (n ¼ 13) increased,
Actinobacteria and Streptococcus thermophilus
decreased [42]. Di Cagno et al. found that the duo-
denal mucosal flora of CD patients had not fully recov-
ered after two years of GFD treatment. Although the
abundance of the pathogenic bacteria declined, the
abundance of the beneficial bacteria was still low
[39,43]. Research by De Palma et al. showed that after
healthy adults persisted in GFD, Bifidobacterium,
Lactobacillus, and Bifidobacterium longum counts
decreased, while the Enterobacteriaceae and
Escherichia coli increased [44]. A similar study in CD
children outlined differences in the microbiota com-
position before and after GFD treatment; mean Dice
similarity index between coeliac individuals before and
after GFD treatment was 63.9% ± 15.8%. There’s a loss
of 36.1% of inter-individual similarity. This study also
found that, Bacteroides vulgatus and Escherichia coli
were detected more often in CD patients than in con-
trols (Functional Dyspepsia), and a significant higher
biodiversity in CD paediatric patients ’ duodenal
mucosa was shown [45]. GFD is an important factor
affecting the composition of the intestinal flora. GFD
not only fails to fully restore the digestive tract flora
of CD patients, but also affects the homeostasis of the
flora in healthy people (Table 2). A GFD diet clearly
influences the abundance of several species, in par-
ticular those involved specifically in carbohydrate and
starch metabolism such as family Veillonellaceae (class
Clostridia)[46]. Diet is an important factor affecting
the abundance, diversity and function of the flora.
Under physiological conditions, dietary patterns and
nutritional status have certain effects on the intestinal
flora [47–48]. Among different dietary components,
fibre has a positive effect on gut microflora and their
related metabolites. Compared with standard diet,
GFD contains less fibre [49–51]. Therefore, it can be
preliminarily inferred that, GFD, which contains a small
amount of fibre, is a factor leading to the dysbiosis of
intestinal flora.
5. The effect of probiotics on CD
There were not much studies about probiotics in
affecting CD (Table 3). Olivares et al. found that B. lon-
gum CECT 7347 could help improve the health status
of CD patients who tend to show alterations in gut
microbiota composition and a biased immune
response even on a GFD [52]. A strict diet without
Figure 1. The relationship between Intestinal flora and Gluten in CD.
ANNALS OF MEDICINE 1801

gluten is the only effective way to treat CD for present
time. Although long-term GFD can improve the symp-
toms of CD patients, there are still existing intestinal
flora dysbiosis. At present, there are few studies on
using probiotics as an intervention for CD patients on
the basis of GFD. However, these limited research
results still show that probiotics combined with GFD
can restore the intestinal flora of CD patients. Studies
have confirmed that the ratio of Firmicutes to
Bacteroides and the abundance of Actinobacteria
decrease in children with CD when compared with
healthy controls. In children with GFD, oral probiotics
containing two Bifidobacterium strains (B632 and
BR03) were taken for 3 months. Compared with chil-
dren with GFD who were not supplemented with pro-
biotics, ratio of Firmicutes to Bacteroides and the
abundance of Actinobacteria increased more than
before. And it is basically similar to the faecal flora
composition of healthy children [53]. Probiotic admin-
istration has clearly revealed a negative relationship
between Firmicutes and pro-inflammatory TNF-a.
Moreover, probiotic effect has exposed some new
phyla, particularly Synergistetes, which negatively cor-
related to acetic acid and total SCFAs, suggesting a
potential role in microbiome restoration [54]. A 6-
week probiotic treatment is effective in improving the
severity of IBS-type symptoms in CD patients on strict
GFD, and is associated with a modification of gut
microbiota, characterized by an increase of
Bifidobacteria [55].
There also was study showed that, the probiotic for-
mula when taken orally over the 12-week period did not
significantly alter the microbiota of CD patients who
were strictly adhere to GFD. The probiotic bacteria con-
tained 450 billion viable lyophilized bacteria Streptococcus
thermophilus, Bifidobacterium breve, Bifidoba cter ium lon-
gum, Bifidobacterium infantis, Lactobacillus acidophilus,
Lactobacillus plantarum, Lactobacillus paracasei,and
Lactobacillus delbrueckii subsp. bulgaricus.[56].
6. Summary and outlook
In summary, coeliac disease, gluten, and digestive
tract microflora have complex interactions (Table 4).
CD patients not only have intestinal flora dysbiosis,
but also accompanied by oral microbial dysbiosis. The
current research has not yet determined the exact
microbial model of microflora and CD, and the causal
relationship between the imbalance of the digestive
tract flora and CD is still inconclusive. Gliadin and
digestive tract flora are the environmental factors that
induce CD, and there is a close relationship between
the two factors. In the process of CD intestinal muco-
sal immune response, gliadin and flora play a synergis-
tic effect, so that the intestinal immune response is
continuously activated, causing clinical symptoms of
CD. In addition, gluten-containing food provides abun-
dant material energy for the digestive tract flora,
which further leads to the imbalance of the flora.
Some specific bacteria or some bacterial metabolites
in the digestive tract can degrade gliadin even though
mammals lack proteases to digest gliadin. On one
hand, there is a causal relationship between gliadin
and the imbalance of the flora. On the other hand,
Table 2. The influence of GFD on gastrointestinal flora.
CD with GFD Healthy people with GFD
Francavilla et al. [42] Decreased: Bifidobacterium, Lactobacillus, and
Bifidobacterium longum
Increased: Actinobacteria and Streptococcus
thermophilus
–
Di Cagno et al. [43] Decreased: pathogenic bacteria declined –
De Palma et al. [44] – Decreased: Bifidobacterium, Lactobacillus, and
Bifidobacterium longum
Increased: Enterobacteriaceae and
Escherichia coli
Schippa et al. [45] Increased: Bacteroides vulgatus and Escherichia
coli; biodiversity of flora
–
Bonder et al. [46] – Decreased: Veillonellaceae
Table 3. The effect of probiotics on CD with GFD.
Positive result Negative result
GFD combined B. longum CECT 7347 [52] Improve the health status of CD patients –
GFD combined probiotics (containing two
Bifidobacterium strains) [53]
Restore intestinal flora basically –
GFD combined probiotics [55] Increase of Bifidobacteria; improve severity of
IBS–type symptoms of CD patients
–
GFD combined probiotics [56] – No effect on intestinal flora
1802 X. WU ET AL.

there is a synergistic relationship in the process of
inducing CD intestinal immune response. In the future,
we should conduct intensive research to clarify the
common role of intestinal microecology and gluten in
the pathogenesis of CD. Nevertheless, alterations of
microbiota in CD subjects may not be considered
exclusively as a consequence of the disease itself, but
rather as a part of a complex relationship between
many causative factors, including those of diet and
psychological nature.
Persistence of symptoms in patients with CD who
adhere to a GFD is common. Probiotics (especially
Bifidobacterium and Lactobacillus related to gliadin
metabolism) are expected to become an adjuvant
preparation for CD patients and minimize the related
adverse reactions caused by strict GFD. Probiotics may
help to control symptoms in patients with CD adher-
ing to a GFD, however, the data are limited and this
could not be an absolute prediction. After all, previous
research had shown that Lactobacillaceae were
enriched in gluten sensitive animal models, so it can-
not be ruled out the possibility that lactobacilli could
not act as probiotics at some time. Future research
studies involving high-quality clinical trials are needed
to improve the quality of the evidence and to estab-
lish the optimal species, timing, and dosage of probi-
otics that may benefit patients with CD [57].
Author contributions
Conceptualisation, WU Xing xing, QIAN Lin and LIU
Kexin,writing- original draft preparation, WU Jing and
SHAN Zhao wei, editing review. All authors have read
and agreed to the published version of
the manuscript.
Disclosure statement
No potential conflict of interest was reported by
the author(s).
Funding
This review was funded by National Nature Science
Foundation of China. No.81873160.
References
[1] Lebwohl B, Sanders DS, Green PHR. Coeliac disease.
Lancet. 2018;391(10115):70–81.
[2] Bai JC, Ciacci C. World gastroenterology organisation
global guidelines: Celiac Disease February 2017. J Clin
Gastroenterol. 2017;51(9):755–768.
Table 4. Summary of changes in the digestive tract flora of patients with coeliac disease and healthy individuals with GFD.
Increase Decrease
Site A- CD T-CD H-GFD Active CD, T-CD H-GFD
Oral Lactobacillus species [23] (A-CD VS HC)Bacteroidetes
[42] (T-CD
1
VS HC)
––Actinobacteria,
Streptoco-ccus thermophilus
[42]
(T-CD
1
VS HC)
–
Small intestine
(Duodenal biopsy)
Proteobacteria [32,33],
Neisseria genus
[32],
Enterobacteriaceae and
Staphylococcus [33]
CD VS T-CD
1
, HC)
Bacteroides vulgatus and
Escherichia coli [45] (A-CD VS FD)
––Firmicutes [32,33],
Actinomycetes
[32]
Streptococcaccae
[33]
(A-CD VS T-CD
1
, HC)
––
Large intestine
(faecal)
– the ratio of Firmicutes to Bacteroides,
Actinomycetes [53] (T-CD
3
VS T-CD
2
),
Bifidobactea [55] (T-CD
3
VS T-CD
2
)
Enterobacteriaceae
and Escherichia coli [44]
(H-GFD VS HC)
the ratio of Firmicutes
to Bacteroides,
Actinomycetes [53]
(A-CD VS HC)
– Bifidobacterium,
Lactobacillus
and Bifidobacterium longum [44]
(H-GFD VS HC)
A-CD: Active CD; T-CD: Treated Coeliac Disease (1: CD with GFD; 2:CD with GFD combined probiotics; 3: CD with GFD combined placebo); H-GFD: Healthy individuals with GFD; HC: Healthy controls; FD:
Functional Dyspepsia.
ANNALS OF MEDICINE 1803

[3] Lerner A, Shoenfeld Y, Matthias T. Adverse effects of
gluten ingestion and advantages of gluten with-
drawal in nonceliac autoimmune disease. Nutr Rev.
2017;75(12):1046–1058.
[4] Cong-Yang YAN, Li ZHOU. Research progresses on
wheat gluten-related diseases. J Food SafQual. 2019;7:
1776–1781.
[5] Stepniak D, Koning F. Celiac disease-sandwiched
between innate and adaptive immunity. Hum
Immunol. 2006;67(6):460–468.
[6] Gianfrani C, Auricchio S, Troncone R. Adaptive and
innate immune responses in celiac disease. Immunol
Lett. 2005;99(2):141–145.
[7] Helmerhorst EJ, Zamakhchari M, Schuppan D, et al.
Discovery of a novel and rich source of gluten-
degrading microbial enzymes in the oral cavity. PLoS
One. 2010;5(10):e13264– 9.
[8] Shan L, Qiao SW, Arentz-Hansen H, et al.
Identification and analysis of multivalent proteolytic-
ally resistant peptides from gluten: implications for
celiac sprue. J Proteome Res. 2005;4(5):1732–1741.
[9] Juan-Li YUAN, Xu J, Shuai HU, et al. Recent advances
in celiac disease. J Food SafQual. 2015;11:4510–4515.
[10] Allen B, Orfila C. The availability and nutritional
adequacy of gluten-free bread and pasta. Nutrients.
2018;10(10):1370.
[11] Cenit MC, Olivares M, Codo
~
ner-Franch P, et al.
Intestinal microbiota and celiac disease: cause, conse-
quence or Co-Evolution? Nutrients. 2015;7(8):
6900–6923.
[12] Sanz Y. Microbiome and gluten. Ann Nutr Metab.
2015;67(Suppl. 2):27–41.
[13] Benn A, Heng N, Broadbent JM, et al. Studying the
human oral microbiome: challenges and the evolution
of solutions. Aust Dent J. 2018;63(1):14–24.
[14] Maukonen J, M
€
att
€
o J, Suihko M-L, et al. Intra-individ-
ual diversity and similarity of salivary and faecal
microbiota. J Med Microbiol. 2008;57(Pt 12):
1560–1568.
[15] Shang L, Deng D, Buskermolen JK, et al. Multi-species
oral biofilm promotes reconstructed human gingiva
epithelial barrier function. Sci Rep. 2018;8(1):16061.
[16] Ling Z, Kong J, Jia P, et al. Analysis of oral microbiota
in children with dental caries by PCR-DGGE and bar-
coded pyrosequencing. Microb Ecol. 2010;60(3):
677–690.
[17] Lockhart PB, Durack DT. Oral microflora as a cause of
endocarditis and other distant site infections. Infect
Dis Clin North Am. 1999;13(4):833–850.
[18] Beck JD, Eke P, Heiss G, et al. Periodontal disease and
coronary heart disease: a reappraisal of the exposure.
Circulation. 2005;112(1):19–24.
[19] Brown JS. Oral biofilms,periodontitis and pulmonary
infections. Oral Dis. 2007;13(6):508–512.
[20] Piombino P, Genovese A, Esposito S, et al. Saliva from
obese individuals suppresses the release of aroma
compounds from wine. PLoS One. 2014;9(1):e85611.
[21] Schmidt TS, Hayward MR, Coelho LP, et al. Extensive
transmission of microbes along the gastrointestinal
tract. Elife. 2019;8:42693.
[22] Atarashi K, Suda W, Luo C, et al. Ectopic colonization
of oral bacteria in the intestine drives TH1 cell
induction and inflammation. Science. 2017;358(6361):
359–365.
[23] Tian N, Faller L, Leffler DA, et al. Salivary gluten deg-
radation and oral microbial profiles in healthy individ-
uals and celiac disease patients. Appl Environ
Microbiol. 2017;83(6):03330–03316.
[24] Fernandez-Feo M, Wei G, Blumenkranz G, et al. The
cultivable human oral gluten degrading microbiome
and its potential implications in coeliac disease and
gluten sensitivity. Clin Microbiol Infect. 2013;19:
386–394.
[25] Di Cagno R, De Angelis M, Lavermicocca P, et al.
Function and diversity of the healthy human micro-
biome. Nature. 2012;486(7402):207–214.
[26] Gilbert JA, Blaser MJ, Caporaso JG, et al. Current
understanding of the human microbiome. Nat Med.
2018;24(4):392–400. 400.
[27] Tap J, Mondot S, Levenez F, et al. Towards the human
intestinal microbiota phylogenetic core. Environ
Microbiol. 2009;11(10):2574–2584.
[28] Sommer F, B
€
ackhed F. The gut microbiota-masters of
host development and physiology. Nat Rev Microbiol.
2013;11(4):227–238.
[29] Zhang L, Xian-Peng Z, Hai-Sheng X, et al. Research
progress in mechanism of intestinal microorganisms
in human diseases. Acta Pharmaceutica Sinica. 2016;6:
843–852.
[30] Davila AM, Blachier F, Gotteland M, et al. Re-print
“intestinal luminal nitrogen metabolism: role of the
gut microbiota and consequences for the host”.
Pharmacol Res. 2013;69(1):114–126.
[31] Bernardo D, Garrote JA, Nadal I, et al. Is it true that
coeliacs do not digest gliadin? Degradation pattern of
gliadin in coeliac disease small intestinal mucosa. Gut.
2009;58(6):886–887.
[32] D’Argenio V, Casaburi G, Precone V, et al.
Metagenomics reveals dysbiosis and a potentially
pathogenic N. flavescens Strain in Duodenum of
Adult Celiac Patients. Am J Gastroenterol. 2016;111(6):
879–890.
[33] Sanchez E, Donat E, Ribes-Koninckx C, et al.
Duodenal-mucosal bacteria associated with celiac dis-
ease in children. Appl Environ Microbiol. 2013;79(18):
5472–5479.
[34] Mohan M, Chow CT, Ryan CN, et al. Dietary gluten-
induced gut dysbiosis is accompanied by selective
upregulation of microRNAs with intestinal tight junc-
tion and bacteria-binding motifs in rhesus macaque
model of celiac disease. Nutrients. 2016;8(11):684.
[35] Macfarlane GT, Allison C, Gibson SAW, et al.
Contribution of the microflora to proteolysis in the
human large intestine. J Appl Bacteriol. 1988;64(1):
37–46.
[36] Caminero A, Herran AR, Nistal E, et al. Diversity of the
cultivable human gut microbiome involved in gluten
metabolism: isolation of microorganisms with poten-
tial interest for coeliac disease. FEMS Microbiol Ecol.
2014;88(2):309–319.
[37] Caminero A, McCarville JL, Galipeau HJ, et al.
Duodenal bacterial proteolytic activity determines
sensitivity to dietary antigen through protease-acti-
vated receptor-2. Nat Commun. 2019;10(1):1198.
1804 X. WU ET AL.

[38] Herr
an AR, P
erez-Andr
es J, Caminero A, et al. Gluten-
degrading bacteria are present in the human small
intestine of healthy volunteers and celiac patients.
Res Microbiol. 2017;168(7):673–684.
[39] Reddel S, Putignani L, Del Chierico F. The impact of
Low-FODMAPs, gluten-free, and ketogenic diets on
gut microbiota modulation. in pathological condi-
tions. Nutrients. 2019;11(2):373.
[40] Hao-Nan YU, Zhi-Hua LIU. Recent progress in intes-
tinal microbiota and mucosal immunity. Chin J
Immun. 2019;16:1921–1930.
[41] Yue ZHOU, Shan DU, Bin CHEN. The theory of an
intestinal microecological imbalance. J Pathogen Biol.
2019;17(7):867–870.
[42] Francavilla R, Ercolini D, Piccolo M, et al. Salivary
microbiota and metabolome associated with celiac
disease. Appl Environ Microbiol. 2014;80(11):1–27.
[43] Di Cagno R, Angelis M, Pasquale I, et al. Duodenal
and faecal microbiota of celiac children: molecular,
phenotype and metabolome characterization. BMC
Microbiol. 2011;11:219.
[44] De Palma G, Nadal I, Collado MC, et al. Effects of a
gluten-free diet on gut microbiota and immune func-
tion in healthy adult human subjects. Br J Nutr. 2009;
102(8):1154–1160.
[45] Schippa S, Iebba V, Barbato M, et al. A distinctive
‘microbial signature’ in celiac pediatric patients. BMC
Microbiol. 2010;10(1):175.
[46] Bonder MJ, Tigchelaar EF, Cai X, et al. The influence
of a short-term gluten-free diet on the human gut
microbiome. Genome Med. 2016;8(1):45.
[47] Albenberg LG, Wu GD. Diet and the intestinal micro-
biome: associations, functions, and implications for
health and disease. Gastroenterology. 2014;146(6):
1564–1572.
[48] Sheflin AM, Melby CL, Carbonero F, et al. Linking diet-
ary patterns with gut microbial composition and func-
tion. Gut Microbes. 2017;8(2):113–129.
[49] De Angelis M, Montemurno E, Vannini L, et al. Effect
of whole-grain barley on the Human Fecal Microbiota
and Metabolome. Appl Environ Microbiol. 2015;81(22):
7945–7956.
[50] Sonnenburg ED, Smits SA, Tikhono M, et al. Diet-
induced extinctions in the gut microbiota compound
over generations. Nature. 2016;529(7585):212–215.
[51] Vitaglione P, Mennella I, Ferracane R, et al. Whole-
grain wheat consumption reduces inflammation in a
randomized controlled trial on overweight and obese
subjects with unhealthy dietary and lifestyle behav-
iors: role of polyphenols bound to cereal dietary fiber.
Am J Clin Nutr. 2015;101(2):251–261.
[52] Olivares M, Castillejo G, Varea V, et al. Double-blind,
randomised, placebo-controlled intervention trial to
evaluate the effects of Bifidobacterium longum CECT
7347 in children with newly diagnosed coeliac dis-
ease. Br J Nutr. 2014;112(1):30–40.
[53] Quagliariello A, Aloisio I, Bozzi Cionci N, et al. Effect
of Bifidobacterium breve on the intestinal microbiota
of coeliac children on a gluten free diet: a pilot study.
Nutrients. 2016;8(10):660.
[54] Ma
sa P, Martina K, Diana DG, et al. Clinical interven-
tion using bifidobacterium strains in celiac disease
children reveals novel microbial modulators of TNF-a
and short-chain fatty acids. Clinical Nutrition. 2019;
38(3):931.
[55] Francavilla R, Piccolo M, Francavilla A, et al. Clinical
and microbiological effect of a multispecies probiotic
supplementation in celiac patients with persistent
IBS-type symptoms: a randomized, Double-Blind, pla-
cebo-controlled, multicenter trial. J Clin Gastroenterol.
2019;53(3):e117–e25.
[56] Seiler C, Kiflen M, Stefanolo J, et al. Probiotics for
celiac disease: a systematic review and Meta-Analysis
of randomized controlled trials. Am J Gastroenterol.
2020;115(10):1584–1595.
[57] Marasco G, Cirota G, Rossini B, et al. Probiotics, prebi-
otics and other dietary supplements for gut micro-
biota modulation in celiac disease patients. Nutrients.
2020;12(9):2674.
ANNALS OF MEDICINE 1805
